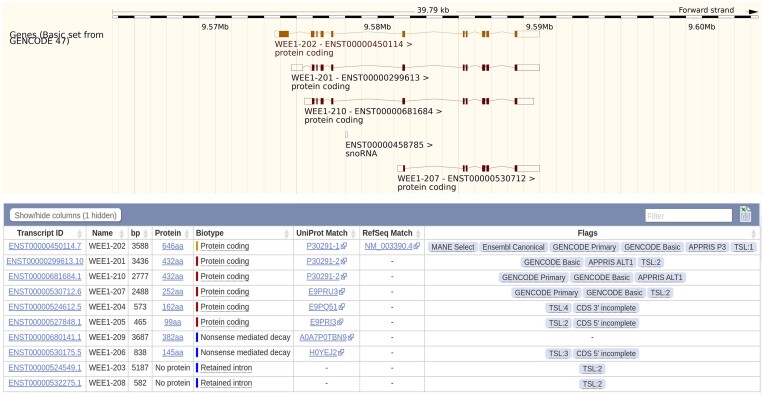
Figure 2.
An Ensembl genome browser view of human protein-coding gene WEE1. Certain sections of the webpage have been omitted for clarity. Here, the Basic set is displayed within the annotation view (top), which thus shows only the four transcripts within the locus that have been annotated with full-length CDS. However, as the transcript table shows (bottom), WEE1 contains ten transcript models, six of which are annotated as protein-coding. WEE1-202 is the MANE Select choice, with its RefSeq match displayed, and as such it is automatically considered as the single Ensembl Canonical model by the Ensembl project. WEE1-202 is also included in the GENCODE Primary set, as are additional models WEE1-210 and WEE1-207. In contrast, the fact that models WEE1-204 and WEE1-205 have partial CDS keeps them out of both the Basic set and GENCODE Primary, while the other four models (WEE1-203, WEE1-206, WEE1-208 and WEE1-209) are annotated as either nonsense mediated decay candidates or else models that contain retained introns (i.e. introns that have not been spliced out). Tags pertaining to APPRIS support are also visible (P for principal, ALT for alternative isoform; see https://apprisws.bioinfo.cnio.es/). Finally, the TSL tag refers to Transcript Support Level, which we consider in the process of being superseded by the functionality of Basic and GENCODE Primary. It highlights the level of support for a transcript model according to information from traditional mRNA and expressed sequence tag evidence sets.