Abstract
Objectives
Kidney renal clear cell carcinoma (KIRC) is one of the most common lethal cancers in the human urogenital system. As members of the Homeobox (HOX) family, Homeobox-A (HOXA) cluster genes have been reported to be involved in the development of many cancer types. However, the expression and clinical significance of HOXA genes in KIRC remain largely unknown.
Materials and methods
In this study, we comprehensively analyzed the mRNA expression and prognostic values of HOXA genes in KIRC using The Cancer Genome Atlas (TCGA) analysis databases online. Colony formation assay, flow cytometry and Western blot were used to detect cell proliferation, apoptosis, cell cycle, and protein level of the indicated gene.
Results
We found that the HOXA genes were differentially expressed in KIRC tissues when compared with normal tissues. The expression of HOXA4 and HOXA13 were significantly up-regulated, while HOXA7 and HOXA11 were down-regulated in KIRC. High mRNA levels of HOXA2, HOXA3 and HOXA13, and low level of HOXA7 predicted poor overall survival (OS) of KIRC patients. High mRNA level of HOXA13 further indicated a poor disease-free survival (DFS) of KIRC patients. Functionally, knockdown of HOXA13 significantly suppressed cell proliferation of KIRC in vitro, increased the protein level of p53 and decreased the protein level of cyclin D1 in KIRC cells. Over-expression of HOXA13 had the opposite effects on KIRC cells.
Conclusion
Collectively, our findings suggest that HOXA13 functions as a novel oncogene in KIRC and may be a potential biomarker for this malignancy.
Electronic supplementary material
The online version of this article (10.1007/s00432-020-03259-x) contains supplementary material, which is available to authorized users.
Keywords: Kidney renal clear cell carcinoma, Homeobox-A cluster genes, HOXA13, Oncogene, Prognosis
Introduction
Kidney renal clear cell carcinoma (KIRC), a common genitourinary malignancy, is the main subtype and accounts for about 80% of all renal cell carcinoma (RCC) cases. Patients with KIRC have higher rates of tumor recurrence and metastasis when compared to patients with other subtypes of RCC (Dong et al. 2019; Siegel et al. 2017). According to the Cancer Statistics, approximately 140,000 cases per year of KIRC are estimated to die worldwide (Siegel et al. 2017). Although several treatment strategies including chemoradiotherapy, INF-α therapy and targeted therapy (sorafenib, sunitinib) have been implemented in clinical practice, the prognosis of KIRC patients remains unsatisfactory (Siegel et al. 2017; Huang et al. 2019). Therefore, exploring novel effective biomarkers will contribute to improving the treatment and prognosis of KIRC.
Homeobox (HOX) genes, a large group of evolutionary conserved family, are crucial transcriptional regulators that play diverse roles from embryogenesis to tumorigenesis. In humans, the HOX family contains four gene clusters termed HOXA, HOXB, HOXC and HOXD, which are located on different chromosomes (7p15, 17q21, 12q13 and 2q31) (Bhatlekar et al. 2018). HOXA cluster genes include 11 members named as HOXA1, HOXA2, HOXA3, HOXA4, HOXA5, HOXA6, HOXA7, HOXA9, HOXA10, HOXA11 and HOXA13. Recent evidence has shown that HOXA genes participated in the initiation and development of various human malignancies (Bhatlekar et al. 2014,2018; Wang et al. 2016). For example, HOXA1 is up-regulated in breast cancer tissues and is associated with patient prognosis (Liu et al. 2019). HOXA3 promotes colon cancer progression by regulating EGFR/Ras/Raf/MEK/ERK pathway (Zhang et al. 2018). HOXA5 can cooperate with p53 to inhibit cell invasion and serve as a possible prognostic factor in non-small cell lung cancer (Chang et al. 2017). Up-regulated HOXA13 promotes gastric cancer progression while high HOXA10 exerts tumor suppressive role in prostate cancer (Qin et al. 2019; Hatanaka et al. 2019). These findings imply that HOXA genes may contribute to carcinogenesis and have potential clinical application values. However, to date, little is known about the expression profile and clinical significance of HOXA genes in KIRC.
As far as we know, comprehensive bioinformatics analysis has not been reported to explore the expression and clinical significance of HOXA genes in KIRC. In this study, several members of the HOXA genes were found to have substantial clinical values in KIRC. The findings of this study highlight the crucial role of HOXA13 in the progression and prognosis of KIRC.
Materials and methods
The Cancer Genome Atlas (TCGA) data analysis
Four online TCGA analysis databases (GEPIA, starBase v3.0, UALCAN and Kaplan–Meier Plotter) were used to observe the expression and prognostic value of 11 members of HOXA genes in KIRC. GEPIA (Tang et al. 2017) was used to observe the expression and prognosis (OS and DFS) of HOXA genes in KIRC. The starBase v3.0 (Li et al. 2014) tool was utilized to examine the expression and prognosis (OS) of HOXA genes in KIRC. UALCAN (Chandrashekar et al. 2017) was used to observe the expression of HOXA genes in KIRC. Kaplan–Meier Plotter (Nagy et al. 2018) was utilized to determine the relationship between HOXA genes expression and patient prognosis (OS and DFS) in KIRC. The co-expressed genes of HOXA13 in KIRC were obtained from the TCGA database LinkedOmics (Vasaikar et al. 2018). Gene Set Enrichment Analysis (GSEA) of HOXA13-related genes in KIRC based on the Kyoto Encyclopedia of Genes and Genomes (KEGG) was analyzed by the WebGestalt (Liao et al. 2019).
Cell culture and siRNA transfection
Three human KIRC cell lines (786-O, Caki-1 and A498) and one normal renal tubular epithelial cell line (HK-2) were previously obtained from Procell Life Science & Technology (China). 786-O, Caki-1 and A498 were cultured in RPMI medium supplemented with 10% FBS (Gibco, USA) (37 ℃, 5% CO2). HK-2 was maintained in DMEM medium supplemented with 10% FBS (Gibco, USA) (37 ℃, 5% CO2). The INTERFERin reagent (Polyplus Transfection, France) was used to transfect siRNAs or plasmids into Caki-1 cells.
Quantitative real-time PCR (qRT-PCR)The qRT-PCR analysis was utilized to examine the transcriptional level of HOXA13 in KIRC tissues and cells. In brief, the Trizol reagent (Invitrogen, USA) and cDNA synthesis kit (DBI, Germany) were used for RNA isolation and reverse transcription, respectively. Then, qRT-PCR analysis was performed using the SYBRGreen qRT-PCR kit (DBI, Germany) and the Fast 7500 system (ABI, USA). The primers are shown as follows:
HOXA13-forward: 5′-CGCTTCAGAACTCGTTGCTTTGC-3′;
reverse: 5′-CGGAAGAACTGGCAGTCTTTACCT3′.
GAPDH-forward: 5′-TCAGAGGACGGCATGAGACTTA-3′;
reverse: 5′-AGCAGGACCCAGGTGTCATT3′.
Colony formation assay
Cell proliferative ability of Caki-1 after transfected with siRNAs or pcDNA-HOXA13 plasmids was examined by the colony formation assay. Briefly, transfected cells were seeded into 6-well plates (1000 cells per well) and cultured in a cell incubator for about 7 days with medium replacement. Then, cell colonies were fixed in methanol, stained with crystal violet solution and calculated by the Image J software.Flow cytometry
Cell apoptosis and cell cycle of Caki-1 transfected with siRNAs or pcDNA-HOXA13 plasmids were detected by flow cytometry. Briefly, cells were seeded into 6-well plates. Forty-eight hours after transfection, cells were subjected to apoptosis and cell cycle detection by using the Annexin V-FITC/PI kit (7sea biotech, China) and PI kit (7sea biotech, China), respectively. The procedures were following the user’s instructions.
Western blot analysis
Total protein of each sample was obtained by using RIPA lysis buffer. After denaturation, an equal amount of protein from each sample was used to carry out SDS-PAGE and PVDF membrane transfer. Then, the membranes were blocked in 5% skim milk and incubated with diluted primary antibodies (GAPDH, Abcam, USA; p53, Santa, USA; Cyclin D1, Santa, USA) and HRP-conjugated second antibody. Finally, the immunoreactive protein bands were detected by ECL reagent and the imaging system (Bio-Rad, USA).
Statistical analysis
For the expression of HOXA genes in cancer, fold change greater than 2 and p value less than 0.05 were considered as significant difference. For the prognostic value of HOXA genes in cancer, patients in each database were divided into two groups (low and high) according to the median level of each HOXA gene in KIRC tissues, and log-rank p value less than 0.05 was considered to be statistically significant. Data of functional experiments are presented as mean ± SD. SPSS 19.0 was used for statistical analysis (Student’s t-test). A p value less than 0.05 was considered to be statistically significant.
Results
The transcriptional level of each member of HOXA genes in KIRC
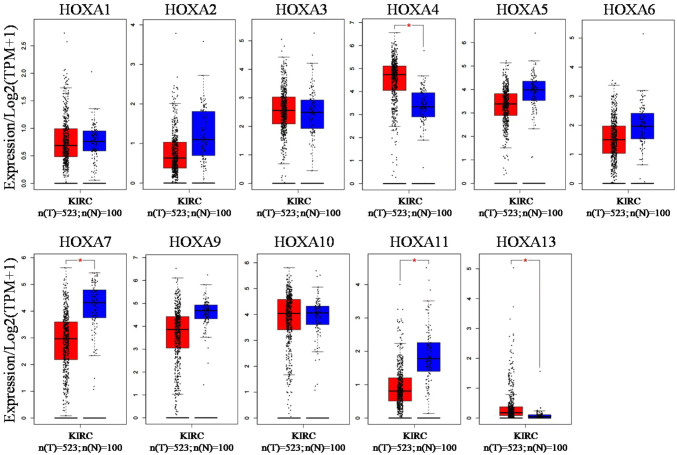
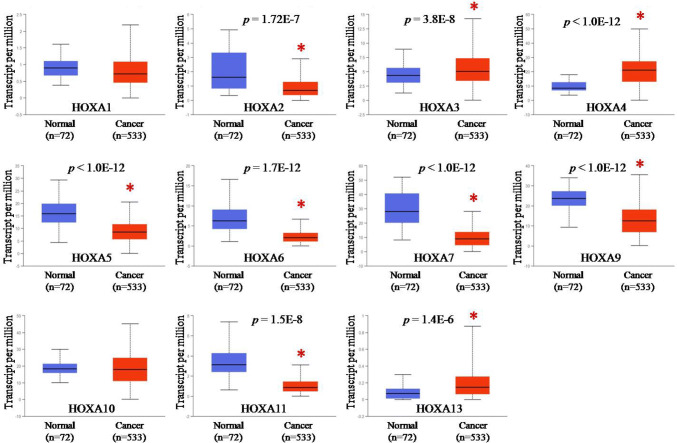
Three online TCGA analysis tools (GEPIA, UALCAN and starBase v3.0) were used to analyze the transcriptional levels of HOXA genes in patients with KIRC. Data from GEPIA showed that the mRNA levels of HOXA7 (p < 0.05) and HOXA11 (p < 0.05) were significantly down-regulated, while HOXA4 (p < 0.05) and HOXA13 (p < 0.05) were obviously up-regulated in KIRC tissues (n = 523) as compared with the adjacent normal tissues (n = 100) (Fig. 1). There were no significant changes in mRNA levels of HOXA1, HOXA2, HOXA3, HOXA5, HOXA6, HOXA9 and HOXA10 between KIRC tissues and normal tissues. Data from UALCAN displayed that the mRNA levels of HOXA2 (p = 1.72E-7), HOXA5 (p < 1.0E-12), HOXA6 (p = 1.0E-12), HOXA7 (p < 1.0E-12), HOXA9 (p < 1.0E-12) and HOXA11 (p = 1.5E-8) were evidently down-regulated, while HOXA3 (p = 3.8E-8), HOXA4 (p < 1.0E-12) and HOXA13 (p = 1.4E-6) were obviously up-regulated in KIRC tissues (n = 533) as compared with the adjacent normal tissues (n = 72) (Fig. 2). There were no significant changes in mRNA levels of HOXA1 and HOXA10 between KIRC tissues and normal tissues. Data from starBase v3.0 (Fig. 3) showed the same trend of HOXA genes expression between KIRC (n = 535) and normal (n = 72) tissues with the results from UALCAN. A comparison of these three datasets revealed that HOXA4 and HOXA13 were significantly increased, while HOXA7 and HOXA11 were dramatically decreased in KIRC tissues (Fig. 9a).
Fig. 1.
The transcriptional levels of HOXA genes in KIRC were analyzed by GEPIA
Fig. 2.
The transcriptional levels of HOXA genes in KIRC were analyzed by UALCAN
Fig. 3.
The transcriptional levels of HOXA genes in KIRC were analyzed by starBase v3.0
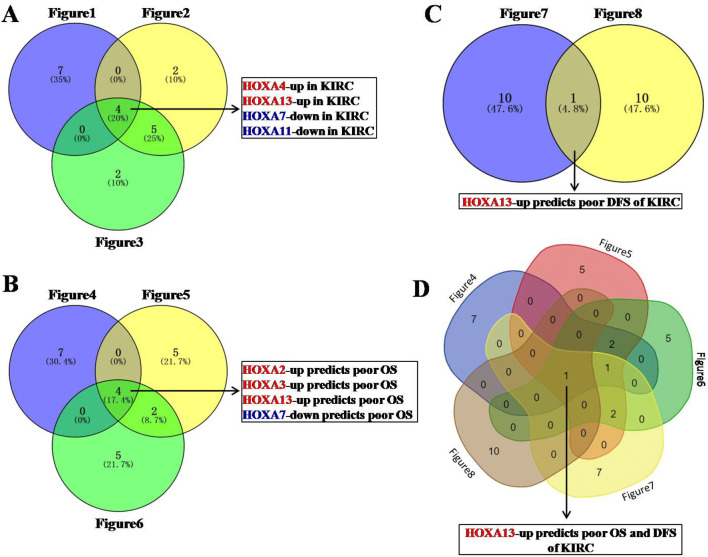
Fig. 9.
Data comparison was shown as Venn diagram (a) Data comparison of Fig. 1, Fig. 2 and Fig. 3 revealed four dysregulated HOXA members (HOXA4, HOXA13, HOXA7 and HOXA11) in KIRC. (b) Data comparison of Fig. 4, Fig. 5 and Fig. 6 revealed that four HOXA members (HOXA2, HOXA3, HOXA13 and HOXA7) were closely related to patient OS in KIRC. (c) Data comparison of Fig. 7 and Fig. 8 revealed that HOXA13 was closely related to patient DFS in KIRC. (d) Data comparison of Fig. 4, Fig. 5, Fig. 6, Fig. 7 and Fig. 8 revealed that HOXA13 was closely related to both OS and DFS of KIRC patients
In this database, HOXA4 and HOXA13 were significantly up-regulated, while HOXA7 and HOXA11 were down-regulated in KIRC as compared to the normal tissues. The red bar graph refers to tumor tissue (n = 523) and the blue bar graph refers to normal tissue (n = 100), *p < 0.05.
In this database, HOXA3, HOXA4 and HOXA13 were significantly up-regulated, while HOXA2, HOXA5, HOXA6, HOXA7 and HOXA11 were down-regulated in KIRC as compared to the normal tissues. The red bar graph refers to tumor tissue (n = 533) and the blue bar graph refers to normal tissue (n = 72), *p < 0.05.
In this database, HOXA3, HOXA4 and HOXA13 were significantly up-regulated, while HOXA2, HOXA5, HOXA6, HOXA7 and HOXA11 were down-regulated in KIRC as compared to the normal tissues. The orange bar graph refers to tumor tissue (n = 535) and the purple bar graph refers to normal tissue (n = 72), *p < 0.05.
The prognostic values of HOXA genes for OS in KIRC patients
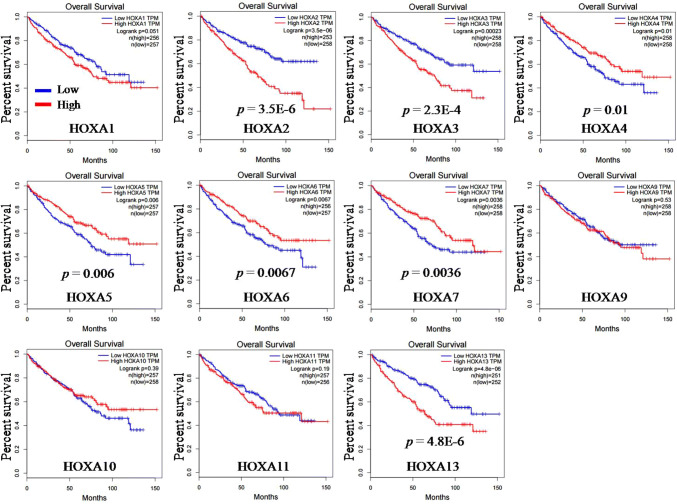
Three online TCGA analysis tools (GEPIA, starBase v3.0 and Kaplan–Meier Plotter) were used to analyze the relationship between each HOXA gene expression and patient OS in KIRC. Data from GEPIA showed that high expression of HOXA2 (p = 3.5E-6), HOXA3 (p = 2.3E-4) and HOXA13 (p = 4.8E-6), while low expression of HOXA4 (p = 0.01), HOXA5 (p = 0.006), HOXA6 (p = 0.0067) and HOXA7 (p = 0.0036) independently predicted poor OS of patients with KIRC (Fig. 4). Data from starBase v3.0 displayed that high expression of HOXA1 (p = 0.0011), HOXA2 (p = 2.7E-8), HOXA3 (p = 1.6E-4), HOXA10 (p = 0.035), HOXA11 (p = 0.0097) and HOXA13 (p = 8.5E-7), while low expression of HOXA7 (p = 0.041) independently predicted poor OS of patients with KIRC (Fig. 5). Data from Kaplan–Meier Plotter indicated that high expression of HOXA1 (p = 4.1E-4), HOXA2 (p = 2.3E-8), HOXA3 (p = 5.4E-5), HOXA11 (p = 0.013) and HOXA13 (p = 3.7E-6), while low expression of HOXA7 (p = 0.025) independently predicted poor OS of patients with KIRC (Fig. 6). Comparison of these three datasets revealed that high expression of HOXA2, HOXA3 and HOXA13, while low expression of HOXA7 independently predicted poor OS of patients with KIRC (Fig. 9b).
Fig. 4.
The OS prognostic values of HOXA genes in KIRC were analyzed by GEPIA
Fig. 5.
The OS prognostic values of HOXA genes in KIRC were analyzed by starBase v3.0
Fig. 6.
The OS prognostic values of HOXA genes in KIRC were analyzed by Kaplan–Meier Plotter
Seven members (HOXA2, HOXA3, HOXA4, HOXA5, HOXA6, HOXA7 and HOXA13) were found to be significantly associated with patient OS in KIRC. Up-regulation of HOXA2, HOXA3 and HOXA13 was significantly associated with poor patient OS, while down-regulation of HOXA4, HOXA5, HOXA6 and HOXA7 was correlated with poor patient OS in KIRC. The red line refers to high gene expression and the blue line refers to low gene expression.
Seven members (HOXA1, HOXA2, HOXA3, HOXA7, HOXA10, HOXA11 and HOXA13) were found to be significantly associated with patient OS in KIRC. Up-regulation of HOXA1, HOXA2, HOXA3, HOXA10, HOXA11 and HOXA13 was significantly associated with poor patient OS, while down-regulation of HOXA7 was correlated with poor patient OS in KIRC. The green line refers to high gene expression and the brown line refers to low gene expression.
Six members (HOXA1, HOXA2, HOXA3, HOXA7, HOXA11 and HOXA13) were found to be significantly associated with patient OS in KIRC. Up-regulation of HOXA1, HOXA2, HOXA3, HOXA11 and HOXA13 was significantly associated with poor patient OS, while down-regulation of HOXA7 was correlated with poor patient OS in KIRC. The red line refers to high gene expression and the black line refers to low gene expression.
The prognostic values of HOXA genes for DFS in KIRC patients
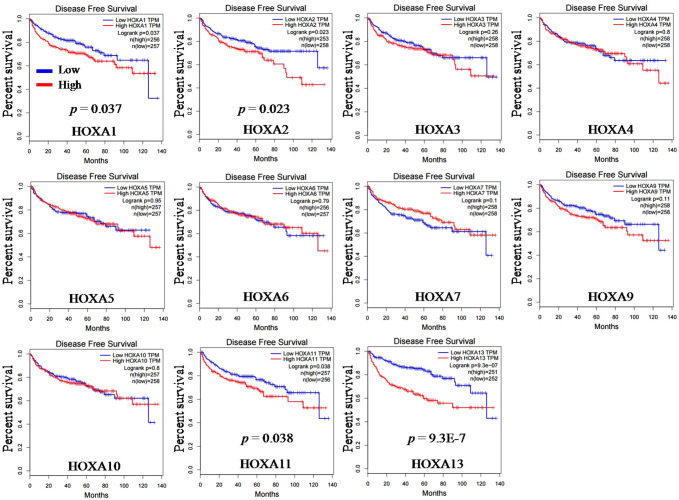
Two online TCGA analysis tools (GEPIA and Kaplan–Meier Plotter) were used to analyze the relationship between each HOXA gene expression and patient DFS in KIRC. Data from GEPIA showed that high expression of HOXA1 (p = 0.037), HOXA2 (p = 0.023), HOXA11 (p = 0.038) and HOXA13 (p = 9.3E-7) independently predicted poor DFS of patients with KIRC (Fig. 7). Data from Kaplan–Meier Plotter indicated that high expression of HOXA5 (p = 0.046) and HOXA13 (p = 0.021) independently predicted poor DFS of patients with KIRC (Fig. 8). A comparison of these two datasets revealed that high expression of HOXA13 predicted poor DFS of patients with KIRC (Fig. 9c). Next, we compared the survival data from five datasets (Figs. 4, 5, 6, 7 and 8), and found that only HOXA13 was closely associated with the OS and DFS of KIRC patients. High mRNA level of HOXA13 predicts unfavorable OS and DFS for patients with KIRC (Fig. 9d).
Fig. 7.
The DFS prognostic values of HOXA genes in KIRC were analyzed by GEPIA
Fig. 8.
The DFS prognostic values of HOXA genes in KIRC were analyzed by Kaplan–Meier Plotter
Four members (HOXA1, HOXA2, HOXA11 and HOXA13) were found to be significantly associated with patient DFS in KIRC. Up-regulation of HOXA1, HOXA2, HOXA11 and HOXA13 was significantly associated with poor patient DFS in KIRC. The red line refers to high gene expression and the blue line refers to low gene expression.
Two members (HOXA5 and HOXA13) were found to be significantly associated with patient DFS in KIRC. Up-regulation of HOXA5 and HOXA13 was significantly associated with poor patient DFS in KIRC. The red line refers to high gene expression and the black line refers to low gene expression.
HOXA13 functions as an oncogene and promotes KIRC proliferation
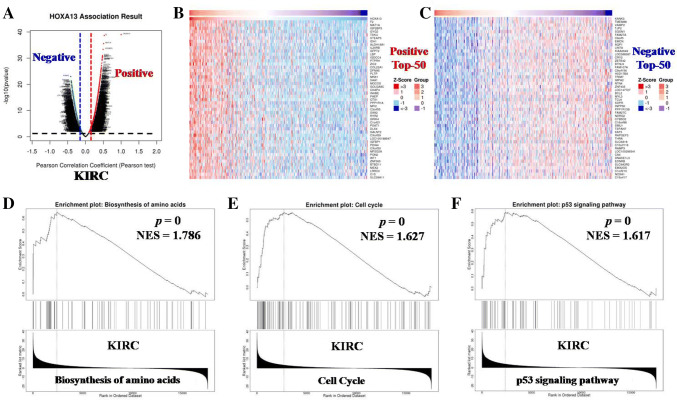
Given that HOXA13 was significantly up-regulated in KIRC and closely correlated with patient prognosis including both OS and DFS, we speculated that HOXA13 might act as an oncogene in KIRC. To investigate the possible pathways mediated by HOXA13 in KIRC, we obtained the co-expressed genes of HOXA13 in KIRC from the LinkedOmics. The overall closely co-expressed genes of HOXA13 in KIRC were shown as a volcano plot (Fig. 10a). The top-50 positively and negatively co-expressed genes of HOXA13 in KIRC were shown as heat maps (Fig. 10b–c), respectively. GSEA based on the KEGG database was used to explore the potential pathways mediated by HOXA13 in KIRC. The results displayed that HOXA13 was primarily implicated in the regulation of biosynthesis of amino acids (p = 0, NES = 1.786), cell cycle (p = 0, NES = 1.627) and p53 signaling (p = 0, NES = 1.617) in KIRC (Fig. 10d–f). To confirm this, we observed the effects of HOXA13 knockdown or over-expression on KIRC proliferation in vitro. First, we examined the expression of HOXA13 in 8 pairs of KIRC tissues as well as in KIRC cell lines, and found that HOXA13 was highly expressed in KIRC tissues (p < 0.05) (Suppl. Fig. 1a) and cell lines (p < 0.05) (Suppl. Fig. 1b) as compared to the counterparts. Then, our results of functional experiments displayed that knockdown of HOXA13 significantly suppressed cell proliferation (p < 0.05) (Fig. 11a) might be through arresting cell cycle at the G0/G1 phase (p < 0.05) (Fig. 11b), rather than promoting apoptosis (Suppl. Fig. 1c). Results of Western blot indicated that down-regulation of HOXA13 increased the protein level of p53 and decreased the level of cell cycle-related protein cyclin D1 in Caki-1 cells (Fig. 11c). Over-expression of HOXA13 had the opposite effects on KIRC cells (p < 0.05) (Fig. 11a–c). These results suggest that HOXA13 promotes KIRC proliferation in vitro.
Fig. 10.
GSEA analysis of the HOXA13-related genes in KIRC. a The overall co-expressed genes of HOXA13 in KIRC were displayed as a volcano plot. b–c The positively (b) and negatively (c) co-expressed top-50 genes of HOXA13 in KIRC were shown as heat maps. d–f Three pathways (biosynthesis of amino acids, cell cycle and p53) were closely associated with HOXA13 in KIRC. NES refers to normalized enrichment score
Fig. 11.
The effects of HOXA13 knockdown or over-expression on cell proliferation in Caki-1 cells. a Colony formation assay was used to detect cell proliferation, *p < 0.05. b Flow cytometry was used to examine cell cycle, *p < 0.05. c Protein levels of p53 and cyclin D1 were determined by Western blot
Discussion
Accumulating evidence highlights the role of the HOXA gene family in the occurrence and progression of many human cancers. For instance, HOXA4 and HOXA10 contribute to the growth and metastasis of ovarian cancer (Idaikkadar et al. 2019). High expression of HOXA10 promotes cell proliferation, inhibits apoptosis and indicates worse OS of patients with gastric cancer (Song et al. 2019). Epigenetic silence of three HOXA genes (HOXA2, HOXA5 and HOXA5) may be important in the development of colorectal cancer (Li et al. 2019). However, the roles of all members of the HOXA gene family in KIRC remain unclear. Herein, we for the first time comprehensively analyzed the expression profile and clinical significance of HOXA genes based on the online TCGA analysis databases. We report that HOXA13 is a novel oncogene in KIRC based on bioinformatic analysis and experimental verification.
HOXA4, HOXA7, HOXA11 and HOXA13 have been proven to be involved in cancer-related biological processes. HOXA4 promotes cisplatin and gefitinib resistance in lung cancer (Yu et al. 2018; Tang et al. 2019). HOXA7 potentiates liver cancer metastasis via Snail signaling (Tang et al. 2016). HOXA11 functions as an oncogenic gene and is associated with tumor stage in renal cell carcinoma (Wang et al. 2017a, b). In this study, we explored the expression profile of HOXA genes in KIRC and found that HOXA4 and HOXA13 were significantly up-regulated, while HOXA7 and HOXA11 were dramatically down-regulated in KIRC. This implies that these four members of HOXA genes are potential candidates to be developed as early diagnostic or (and) therapeutic biomarkers for patients with KIRC.
Previous studies have shown that several HOXA genes have certain prognostic values in some cancer types. HOXA2 may serve as a prognostic parameter in non-small cell lung cancer and cervical cancer patients (Heller et al. 2013; Eoh et al. 2017). Up-regulated expression of HOXA3 is closely associated with a low OS rate of colon cancer patients (Zhang et al. 2018). In both prostate carcinoma and bladder cancer, HOXA13 is an unsatisfactory prognostic factor for patient survival (Dong et al. 2017; Hu et al. 2017). In this study, four members (HOXA2, HOXA3, HOXA13 and HOXA7) of the HOXA gene family were found to be closely associated with the OS of KIRC patients. These data indicate that these four members of HOXA genes are possible prognostic biomarkers for KIRC.
Considering that HOXA13 was significantly up-regulated in KIRC and significantly correlated with both OS and DFS in KIRC. We speculated that HOXA13 might be an important tumor driving gene in KIRC. Then, we performed KEGG signaling pathway analysis to explore the possible pathways mediated by HOXA13 in KIRC. Cell cycle and p53 signaling were found to be closely associated with HOXA13. P53, known as a suppressor in human cancer, is also involved in the development of KIRC (Wang et al. 2017a, b). In a previous study, it has been reported that HOXA13 contributed to gastric cancer progression in a p53-dependent way (Han et al. 2018). Thus, we speculated that HOXA13 might participate in KIRC through p53. To test this hypothesis, we performed some preliminary in vitro functional experiments, and found that HOXA13 promoted KIRC proliferation through cell cycle arrest. The protein level of p53 was found to be significantly increased after down-regulation of HOXA13 in KIRC cells.
In summary, this study comprehensively analyzed the expression and prognostic values of the HOXA gene family in KIRC using public online TCGA analysis databases. The study implies that some members of the HOXA genes may serve as clinical biomarkers for KIRC. Our findings highlight the important role of HOXA13 in the progression of KIRC and its crucial clinical significance in KIRC. Furthermore, we provide a preliminary understanding regarding the function and regulatory pathway mediated by HOXA13 in KIRC. Our study suggests that HOXA13 promoting KIRC progression may correlate with the p53 signaling pathway. The specific role of HOXA13 in KIRC needs to be further extensively explored.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Acknowledgements
We thank all participants for this manuscript.
Author contributions
YBC conceived and designed this study and wrote the manuscript. MY, CYZ, JHX, QWZ and SSM participated in functional experiments and data analysis. WC and FXG assisted in the revision of the manuscript and sample collection. All authors read the manuscript and approved it for publication.
Funding
This work was supported by the Key Scientific Research Projects of Institutions of Higher Learning in Henan Province (20A310018), the Provincial Co-construction Project of Medical Science and Technology in Henan (SB201903032), the Key Scientific and Technological Project in Henan Province (182102310328).
Compliance with ethical standards
Conflict of interest
The authors have declared that they have no competing interests.
Ethical approval
This study was approved by the Ethics Committee of the Zhengzhou Central Hospital Affiliated to Zhengzhou University.
Informed consent
Not applicable for this study.
Footnotes
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Yuanbo Cui and Ming Yan contributed equally to this work.
Contributor Information
Yuanbo Cui, Email: cuiyuanbo18@126.com.
Fangxia Guan, Email: fxguan@126.com.
Wei Cao, Email: caoweiyu@hotmail.com.
References
- Bhatlekar S, Fields JZ, Boman BM (2014) HOX genes and their role in the development of human cancers. J Mol Med 92(8):811–823 [DOI] [PubMed] [Google Scholar]
- Bhatlekar S, Fields JZ, Boman BM (2018) Role of HOX genes in stem cell differentiation and cancer. Stem Cells Int 2018:3569493 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chandrashekar DS, Bashel B, Balasubramanya SAH, Creighton CJ, Rodriguez IP, Chakravarthi BVSK, Varambally S (2017) UALCAN: A portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia 19(8):649–658 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chang CJ, Chen YL, Hsieh CH, Liu YJ, Yu SL, Chen JJW, Wang CC (2017) HOXA5 and p53 cooperate to suppress lung cancer cell invasion and serve as good prognostic factors in non-small cell lung cancer. J Cancer 8(6):1071–1081 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dong D, Mu Z, Wei N, Sun M, Wang W, Xin N, Shao Y, Zhao C (2019) Long non-coding RNA ZFAS1 promotes proliferation and metastasis of clear cell renal cell carcinoma via targeting miR-10a/SKA1 pathway. Biomed Pharmacother 111:917–925 [DOI] [PubMed] [Google Scholar]
- Dong Y, Cai Y, Liu B, Jiao X, Li ZT, Guo DY, Li XW, Wang YJ, Yang DK (2017) HOXA13 is associated with unfavorable survival and acts as a novel oncogene in prostate carcinoma. Future Oncol 13(17):1505–1516 [DOI] [PubMed] [Google Scholar]
- Eoh KJ, Kim HJ, Lee JY, Nam EJ, Kim S, Kim SW, Kim YT (2017) Upregulation of homeobox gene is correlated with poor survival outcomes in cervical cancer. Oncotarget 8(48):84396–84402 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Han Y, Song C, Wang J, Tang H, Peng Z, Lu S (2018) HOXA13 contributes to gastric carcinogenesis through DHRS2 interacting with MDM2 and confers 5-FU resistance by a p53-dependent pathway. Mol Carcinog 57(6):722–734 [DOI] [PubMed] [Google Scholar]
- Hatanaka Y, de Velasco MA, Oki T, Shimizu N, Nozawa M, Yoshimura K, Yoshikawa K, Nishio K, Uemura H (2019) HOXA10 expression profiling in prostate cancer. Prostate 79(5):554–563 [DOI] [PubMed] [Google Scholar]
- Heller G, Babinsky VN, Ziegler B, Weinzierl M, Noll C, Altenberger C, Müllauer L, Dekan G, Grin Y, Lang G, End-Pfützenreuter A, Steiner I, Zehetmayer S, Döme B, Arns BM, Fong KM, Wright CM, Yang IA, Klepetko W, Posch M, Zielinski CC, Zöchbauer-Müller S (2013) Genome-wide CpG island methylation analyses in non-small cell lung cancer patients. Carcinogenesis 34(3):513–521 [DOI] [PubMed] [Google Scholar]
- Huang Y, Wang J, Jia P, Li X, Pei G, Wang C, Fang X, Zhao Z, Cai Z, Yi X, Wu S, Zhang B (2019) Clonal architectures predict clinical outcome in clear cell renal cell carcinoma. Nat Commun 10(1):1245 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hu H, Chen Y, Cheng S, Li G, Zhang Z (2017) Dysregulated expression of homeobox gene HOXA13 is correlated with the poor prognosis in bladder cancer. Wien Klin Wochenschr 129(11–12):391–397 [DOI] [PubMed] [Google Scholar]
- Idaikkadar P, Morgan R, Michael A (2019) HOX Genes in High Grade Ovarian Cancer. Cancers (Basel) 11(8):E1107 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liao Y, Wang J, Jaehnig EJ, Shi Z, Zhang B (2019) WebGestalt 2019: gene set analysis toolkit with revamped UIs and APIs. Nucleic Acids Res 47(W1):W199–W205 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li D, Bai Y, Feng Z, Li W, Yang C, Guo Y, Lin C, Zhang Y, He Q, Hu G, Li X (2019) Study of promoter methylation patterns of HOXA2, HOXA5, and HOXA6 and its clinicopathological characteristics in colorectal cancer. Front Oncol 9:394 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Li JH, Liu S, Zhou H, Qu LH, Yang JH (2014) starBase v2.0 decoding miRNA-ceRNA, miRNA-ncRNA and protein-RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Res 42(Database issue):D92–7 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu J, Liu J, Lu X (2019) HOXA1 up-regulation is associated with poor prognosis and tumor progression in breast cancer. Exp Ther Med 17(3):1896–1902 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nagy A, Lánczky A, Menyhárt O, Győrffy B (2018) Validation of miRNA prognostic power in hepatocellular carcinoma using expression data of independent datasets. Sci Rep 8:9227 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Qin Z, Chen Z, Weng J, Li S, Rong Z, Zhou C (2019) Elevated HOXA13 expression promotes the proliferation and metastasis of gastric cancer partly via activating Erk1/2. Onco Targets Ther 12:1803–1813 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Siegel RL, Miller KD, Jemal A (2017) Cancer Statistics, 2017. CA Cancer J Clin 67(1):7–30 [DOI] [PubMed] [Google Scholar]
- Song C, Han Y, Luo H, Qin Z, Chen Z, Liu Y, Lu S, Sun H, Zhou C (2019) HOXA10 induces BCL2 expression, inhibits apoptosis, and promotes cell proliferation in gastric cancer. Cancer Med. 10.1002/cam4.2440 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tang B, Qi G, Sun X, Tang F, Yuan S, Wang Z, Liang X, Li B, Yu S, Liu J, Huang Q, Wei Y, Zhai R, Lei B, Guo X, He S (2016) HOXA7 plays a critical role in metastasis of liver cancer associated with activation of Snail. Mol Cancer 15(1):57 [DOI] [PMC free article] [PubMed] [Google Scholar] [Retracted]
- Tang X, Jiang J, Zhu J, He N, Tan J (2019) HOXA4-regulated miR-138 suppresses proliferation and gefitinib resistance in non-small cell lung cancer. Mol Genet Genomics 294(1):85–93 [DOI] [PubMed] [Google Scholar]
- Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z (2017) GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res 45(W1):W98–W102 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vasaikar SV, Straub P, Wang J, Zhang B (2018) LinkedOmics: analyzing multi-omics data within and across 32 cancer types. Nucleic Acids Res 46(D1):D956–D963 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang L, Cui Y, Sheng J, Yang Y, Kuang G, Fan Y, Jin J, Zhang Q (2017a) Epigenetic inactivation of HOXA11, a novel functional tumor suppressor for renal cell carcinoma, is associated with RCC TNM classification. Oncotarget 8(13):21861–21870 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang X, Cheng Y, Zhu Y, Li H, Ge W, Wu X, Zhao K, Yuan J, Li Z, Jiang S, Han Z, Jiang Q, Wu Q, Liu T, Zhang C, Yu M, Hu Y (2017b) Epigenetic silencing of ASPP1 confers 5-FU resistance in clear cell renal cell carcinoma by preventing p53 activation. Int J Cancer 141(7):1422–1433 [DOI] [PubMed] [Google Scholar]
- Wang Y, Dang Y, Liu J, Ouyang X (2016) The function of homeobox genes and LncRNAs in cancer. Oncol Lett 12(3):1635–1641 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yu S, Ren H, Li Y, Liang X, Ning Q, Chen X, Chen M, Hu T (2018) HOXA4-dependent transcriptional activation of AXL promotes cisplatin- resistance in lung adenocarcinoma cells. Anticancer Agents Med Chem 18(14):2062–2067 [DOI] [PubMed] [Google Scholar]
- Zhang X, Liu G, Ding L, Jiang T, Shao S, Gao Y, Lu Y (2018) HOXA3 promotes tumor growth of human colon cancer through activating EGFR/Ras/Raf/MEK/ERK signaling pathway. J Cell Biochem 119(3):2864–2874 [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.