Figure 9.
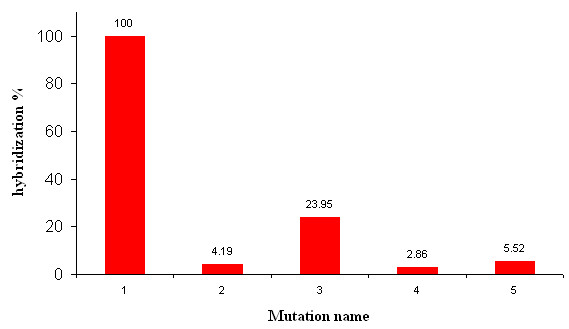
Mutation detection. Comparison of the hybridization signal intensity of the target molecules on 5 different printed patterns differing by only single or double mutations. "Mutation" 1 corresponds to the exact match of the target molecule and serves as a reference. Five 20-mer oligonucleotides probes were printed at 10 μM in Na-Pi buffer 0.3 M, pH 9.0 on a dendrislide. These oligonucleotides were part of the yeast HSP12 sequence, and varied from each other by a single or two mutations proximal to the 5' end or 3' end or in the middle of the sequence. The 20-mer sequences from HSP12 are noted as follows: 1: NH2 5'-AATATGTTTCCGGTCGTGTC-3'; 2: NH2 5'-AATATGTTTCAGGTCGTGTC-3'; 3: NH2 5'-AATATGTTTCCGGTCGTGTA-3'; 4: NH2 5'-AATATGATTCCGGACGTGTC-3'; 5: NH2 5'-AATAAGTTTCCGGTCGTGTC-3'; Hybridisation was carried out with Cy5-labelled oligonucleotide (Cy5 5'-GACACGACCGGAAACATATT 3'). Values of fluorescence intensity were measured at 635 nm with the GenePix 4000B from axon at 600 PMT and correspond to an average of 4 experiments. Statistics errors are less than 0.4% for the 4 experiments.
