Abstract
Background
Multiple myeloma (MM) is a malignant disorder originating from plasma cells. Bortezomib (BTZ) resistance has become a huge obstacle to MM treatment. Herein, we elucidated the action of Kruppel-like factor 2 (KLF2), a crucial transcription factor (TF), on BTZ resistance of MM.
Methods
Two BTZ-resistant cell lines (MM1.S/BTZ and NCI-H929/BTZ) were generated and used. KLF2 mRNA was quantified by quantitative PCR, and protein expression was analyzed by immunoblotting. MTT cell cytotoxicity assay was used to test BTZ sensitivity. Cell growth was detected by MTT and EdU assays. Flow cytometry was used for apoptosis and cycle distribution analyses. The USP4/KLF2 relationship was examined by Co-IP and protein stability assays. The KLF2/HMGA2 interplay was confirmed by luciferase and ChIP assays.
Results
Upregulation of KLF2 was observed in MM serum and BTZ-resistant MM cells. Depletion of KLF2 suppressed cell growth and enhanced apoptosis and BTZ sensitivity in MM1.S/BTZ and NCI-H929/BTZ cells. Moreover, USP4 increased the stability of KLF2 protein by deubiquitination and affected cell growth, apoptosis and BTZ sensitivity via KLF2. KLF2 functioned as a regulator of HMGA2 transcription and modulated cell growth, apoptosis and BTZ sensitivity through HMGA2. Additionally, USP4 modulated HMGA2 expression via KLF2 in the two BTZ-resistant cell lines.
Conclusion
Our study demonstrates the crucial role of the USP4/KLF2/HMGA2 cascade in regulating cell growth, apoptosis and BTZ sensitivity in BTZ-resistant MM cells, providing novel targets for improving anti-MM efficacy of BTZ.
Graphical Abstract
Supplementary Information
The online version contains supplementary material available at 10.1186/s13018-025-05537-1.
Keywords: Multiple myeloma (MM), Drug resistance, KLF2, Deubiquitination, Transcription factor
Highlights
USP4 increased the stability of KLF2 protein by deubiquitination.
KLF2 functioned as a regulator of HMGA2 transcription.
The USP4/KLF2/HMGA2 axis regulates cell growth and BTZ sensitivity in BTZ-resistant MM cells.
Supplementary Information
The online version contains supplementary material available at 10.1186/s13018-025-05537-1.
Introduction
Originating from plasma cells, multiple myeloma (MM) is a malignant disorder, which is specifically designated as plasma cell myeloma/plasmacytoma by the WHO [1]. MM often causes anemia, multiple osteolytic lesions, hypercalcemia, cardiac toxicities, and renal impairment, significantly compromising patients’ quality of life. The incidence of MM peaks in individuals over 40 years old, with a higher prevalence in the population above 60 [2]. The pathogenesis of MM involves a complex interplay of factors, including malignant proliferation of plasma cells, immune dysregulation, overexpression of microenvironmental support factors, genetic susceptibility, and epigenetic dysregulation [3–6]. Currently, the treatment of MM is multifaceted, encompassing targeted therapy, chemotherapy, immunotherapy, and stem cell transplantation [2, 7, 8]. Bortezomib (BTZ), a proteasome suppressor, has emerged as a prominent targeted drug in combating MM [9]. However, BTZ resistance frequently occurs and poses a huge challenge for MM effective cure. Thus, further research on the mechanisms of drug resistance is imperative to improve patient outcomes.
As a zinc-finger transcription factor (TF), Kruppel-like factor 2 (KLF2) plays pivotal roles in various cellular processes and has been implicated in the regulation of inflammation, cardiac protection, and atherosclerosis protection [10, 11]. In the context of cancer biology, KLF2 has emerged as a potential anti-tumor factor, with its expression often downregulated in various cancers, including clear cell renal cell carcinoma, colorectal cancer and gastric cancer. This downregulation is believed to contribute to the carcinogenesis of these malignancies by influencing gene expression and signaling pathways [12–14]. Furthermore, KLF2 can function as an essential modulator in plasma cell homing, highlighting its importance for the development of B/plasma cell-associated malignancies [15]. In MM patients, KLF2 expression is remarkably upregulated and tightly linked to enhanced R-ISS stage and worse progression-free survival. When complete remission induced by treatment, KLF2 expression is significantly reduced [16]. Interestingly, increased expression of KLF2 contributes to enhanced proliferation and adhesion of MM cells, while KLF2 downregulation induces MM cell apoptosis [17, 18], suggesting the oncogenic role of KLF2 in MM. Moreover, BTZ therapy in MM can increase KLF2 expression [17], and the thromboprotective efficacy of BTZ in MM is KLF2-dependent [19]. Despite these advancements, the action of KLF2 on the BTZ resistance of MM remains to be fully elucidated.
Ubiquitination and deubiquitination are central post-translational modification processes in proteins, and their dynamic balance is crucial for maintaining cellular homeostasis and regulating various biological processes [20]. USP4 is an enzyme involved in deubiquitination processes that is essential for modulating protein stability and degradation [21]. Studies have indicated that USP4 has the ability to promote MM development [22]. HMGA2, belonging to the non-histone chromosomal architectural proteins, possesses pivotal functions in chromatin remodeling and gene expression regulation. HMGA2 has established a role in driving tumorigenesis by regulating growth and therapy resistance [23]. However, it is largely unknown whether HMGA2 participates in MM BTZ resistance.
In this work, we demonstrate the effects of KLF2 on BTZ sensitivity, growth, and apoptosis of BTZ-resistant MM cells by establishing two BTZ-resistant cell lines (MM1.S/BTZ and NCI-H929/BTZ). Furthermore, we have uncovered that the USP4/KLF2/HMGA2 axis underlies the phenotypes of BTZ-resistant MM cells. Thus, our study elucidates a novel mechanism in BTZ resistance in MM and suggests potential targets for improving anti-MM efficacy of BTZ.
Materials and methods
Human serum samples and KLF2 mRNA analysis
Under the protocols authorized by the Ethics Committee of Zhuhai People’s Hospital, Zhuhai Hospital Affiliated with Jinan University, Jinan University with approval No.20240616, we collected blood specimens from 51 MM sufferers, including 24 patients with primary MM (called sensitive MM (S-MM) and 27 patients with recurrent MM after treatment based on BTZ therapy (R-MM), from Zhuhai People’s Hospital, Zhuhai Hospital Affiliated with Jinan University, Jinan University. Control subjects (n = 31) were age- and sex-matched healthy volunteers with no known illnesses and no medications. Before blood collection, all participants gave informed consent. For KLF2 mRNA analysis, we harvested serum samples by centrifugalization and extracted total RNA with the RNeasy Plus Mini Kit as suggested by the producer (Qiagen, Hombrechtikon, Switzerland). After RNA quantification, cDNA generation was conducted starting from 200 ng of RNA with PrimeScript RT Reagent Kit from TaKaRa (Dalian, China). Using a Rotorgene 6000 (Qiagen), quantitative PCR reactions were done using SYBR Premix ExTaq (TaKaRa) with KLF2-specific primer pairs (5’-GGGGTGAGTTCCCCATTCTG-3’-sense and 5’-CAGAGCAGGCTAACAACCCA-3’-antisense). Following normalization to GAPDH (5’-GACCACAGTCCATGCCATCAC-3’-sense and 5’-ACGCCTGCTTCACCACCTT-3’-antisense), fold-change in KLF2 expression was calculated by the method 2−ΔΔCt.
Cell lines, generation of BTZ-resistant cells, and cell treatment
Human MM1.S MM cells (#IM-H470, Immocell, Xiamen, China), NCI-H929 MM cells (#SNL-557, Sunncell, Wuhan, China) and non-tumor nPCs cells (#ABC-371, Bioiech, Shanghai, China) were used in this study. The RPMI-1640 (Beyotime, Shanghai, China) was used for the culture of MM1.S and NCI-H929 cells, which was also enriched with 1% Penicillin/Streptomycin (Beyotime) and 10% FBS (Nichirei Bioscience, Tokyo, Japan). Human nPCs cells were maintained in standard nPCs growth medium procured from Bioiech.
We generated BTZ-resistant cells MM1.S/BTZ and NCI-H929/BTZ by exposing naive MM1.S and NCI-H929 MM cells to escalating concentrations of BTZ (starting from 5 nM and progressively elevating the concentration up to 80 nM) as described [24]. To analyze the deubiquitination status of KLF2, MM1.S/BTZ and NCI-H929/BTZ cells were subjected to treatment of PR-619 (the suppressor of deubiquitinating enzymes; 50 µM, Selleck, Shanghai, China) or MG132 (a ubiquitin-proteasome inhibitor; 50 µM, InvivoChem, Guangzhou, China) for 24 h.
Plasmids and transient transfection of cell lines
KLF2 Human Pre-designed siRNA Set A (si-KLF2), USP4 Human Pre-designed siRNA Set A (si-USP4), and the corresponding negative control si-NC were procured from MCE (Shanghai, China) and used for siRNA silencing assays. pCMV-USP4(human)-3×FLAG-Neo (OE-USP4), pCMV-KLF2(human)-3×FLAG-Neo (OE-KLF2), pCMV-3×HA-HMGA2(human)-Blast (OE-HMGA2) and the matched control OE-control were obtained from Miaoling Biotech (Wuhan, China) and applied to perform overexpression experiments. We conducted transfection of siRNA, plasmid or siRNA + plasmid into MM1.S/BTZ and NCI-H929/BTZ cells, in accordance with the manufactory guidelines (Life Technologies, Hemel Hempstead, UK), under the application of Lipofectamine 3000. Transfection was allowed for 6–8 h, and the cells were continued to be cultivated for 24–72 h after the medium change. At 48 h after introduction, we examined silencing or overexpression efficiency using the immunoblotting method.
Immunoblotting
For protein lysate preparation, cultivated cells were homogenized using the Protein Extraction Kit as suggested by the supplier (Abcam, Cambridge, UK). After quantification by BCA assay (Life Technologies), protein samples (~ 25 µg) were subjected to SDS-PAGE electrophoresis, and the resulting gels were imaged and electroblotted to PVDF membranes (Servicebio, Wuhan, China). We carried out membrane probing with mouse anti-KLF2 mAb (#MA5-26827, Invitrogen, Bleiswijk, the Netherlands, 1 to 2,000), rabbit anti-HMGA2 mAb (#ab207301, Abcam, 1 to 1,000), rabbit anti-MRP1 mAb (#ab260038, Abcam, 1 to 1,000), rabbit anti-USP4 pAb (#ab236987, Abcam, 1 to 3,000), rabbit anti-ubiquitin pAb (anti-Ub, #10201-2-AP, Proteintech, Wuhan, China, 1 to 6,000), and rabbit anti-GAPDH pAb (#ab9485, Abcam, 1 to 2,500; loading buffer). Chemiluminescence was achieved by the application of the EZ-ECL Kit (Biological Industries, Beit-Haemek, Israel). The signal intensity was analyzed by the iBright 1500 System (Invitrogen).
MTT cell proliferation and cytotoxicity assays
For proliferation analysis, in a 96-multiwell culture plate, MM1.S/BTZ and NCI-H929/BTZ cells subjected to the relevant transfection were monitored for the number of viable cells at 24, 48 and 72 h post-transfection. For in vitro cytotoxicity analysis, MM1.S/BTZ and NCI-H929/BTZ cells at 48 h after transfection were challenged with various concentrations (10, 20, 40, 80, 160, 320, 640, and 1280 nM) of BTZ for 24 h. In both assays, we changed the media with fresh media containing MTT reagent (Beyotime, 0.5 mg/mL) and performed a 4-h incubation at 37 °C. Following the addition of formazan-solving liquid (Beyotime), we gauged the absorbance at 490 nm with a 96-well spectrometer (BMG Labtech, Ortenberg, Germany). We estimated the IC50 value for BTZ using the plot of the percentage of viable cells versus BTZ concentration.
EdU incorporation assay for cell proliferation
At 48 h after transfection, MM1.S/BTZ and NCI-H929/BTZ cells were subjected to EdU incorporation assay under the use of the Yefluor 488 EdU Imaging Kit as per the accompanying instructions (Yeasen, Shanghai, China). Briefly, EdU reagent was added into each well and applied for 1 h, followed by staining with Yefluor 488. After DAPI incubation for nuclear staining, fluorescence images were taken under a TCS SP5 confocal microscope (Leica, Wetzlar, Germany). Using ImageJ software (NIH, Bethesda, MD, USA), we quantified the EdU positive cells (green fluorescence) relative to total cell count (blue fluorescence).
Flow cytometry
For detection of cell apoptosis (at 72 h post transfection) and cycle distribution (at 48 h post transfection), transfected MM1.S/BTZ and NCI-H929/BTZ cells were processed for staining by the Cell Cycle Analysis Kit (Beyotime) and Annexin A-FITC/PI Apoptosis Analysis Kit (Vazyme Biotech, Nanjing, China), respectively, as suggested by the vendors. We performed data analysis on CytoFLEX LX platforms (Beckman Coulter, Fullerton, CA, USA).
Prediction of deubiquitination-related regulators of KLF2 and the binding site between KLF2 and the HMGA2 promoter
We utilized the online algorithm Ubibrowser2.0 to predict the deubiquitinases associated with KLF2 at http://ubibrowser.bio-it.cn/ubibrowser_v3/. The binding motif of KLF2 and its binding site to the HMGA2 promoter were predicted by JASPAR version 2024 at https://jaspar.elixir.no/.
Analysis of KLF2 protein stability
OE-control- or OE-USP4-transfected MM1.S/BTZ and NCI-H929/BTZ cells were subjected to cycloheximide (CHX, Selleck) treatment at a final concentration of 20 ng/mL. At 0, 5, 10, 20 and 25 h post-treating, the remaining KLF2 protein was assessed by immunoblotting.
Co-IP and IP assays
We utilized an IP Assay Kit to perform these assays as recommended by the supplier (Beyotime). Total extractions of MM1.S/BTZ, NCI-H929/BTZ and si-USP4- or si-NC-transfected MM1.S/BTZ cells were prepared. The complex of Protein A/G beads with rabbit anti-USP4 pAb (#A300-830 A, Bethyl Laboratories, Hemel Hempstead, UK, 5 µg/mg lysate), rabbit anti-KLF2 pAb (#23384-1-AP, Proteintech), or rabbit anti-IgG pAb (#30000-0-AP, Proteintech) was prepared and added into cell lysates. After a 6-h incubation at 4 °C, the immunoprecipitates were harvested, washed, and subjected to protein isolation for quantification of USP4, KLF2, and deubiquitinating KLF2 enrichment levels by immunoblotting.
Luciferase reporter assay
Using the pGL3 based vector (Invitrogen) as templates, HMGA2 luciferase reporters were produced by inserting the HMGA2 promoter segment harboring the predicted binding site (WT-HMGA2, CACCACCCCCT) or its mutations (MUT-HMGA2, AGTTGAATTTC) into the vector. MM1.S/BTZ and NCI-H929/BTZ cells were subjected to transfection with WT-HMGA2 + si-NC, WT-HMGA2 + si-KLF2, MUT-HMGA2 + si-NC, or MUT-HMGA2 + si-KLF2. Meantime, the pRL-TK Renilla vector (Promega, Paris, France) was co-transfected and used as the control for normalization. 48 h later, the Dual-Luciferase Assay Kit (Beyotime) was used for the measurement of luciferase activity (firefly/Renilla).
ChIP experiment
To confirm the interplay of KLF2 with the HMGA2 promoter, we employed the ChIP Assay Kit provided by Gene Create (Wuhan, China). MM1.S/BTZ and NCI-H929/BTZ cells were cross-linked using 1% formaldehyde. Subsequently, the cells were treated with glycine buffer and sonicated to generate chromatin fragments. These fragments were then incubated with the complex of Protein A/G beads with rabbit anti-KLF2 pAb (#PA5-40591, Invitrogen) or anti-IgG isotype antibody (#30000-0-AP, Proteintech). DNA fragments associated with KLF2 were isolated and subjected to quantitative PCR to assess the enrichment abundance of the HMGA2 promoter.
Data analysis and bioinformatics
We conducted data analysis using one- or two-way ANOVA, with statistical significance defined as P values < 0.05. In figures, data were shown as mean ± SD. The GSE55667 dataset (https://www.ncbi.nlm.nih.gov/search/all/?term=GSE55667%20) was used for the analysis of gene expression profiles of KLF2 depletion in myeloma RPMI8226 cells.
Results
Upregulation of KLF2 in MM serum and BTZ-resistant MM cells
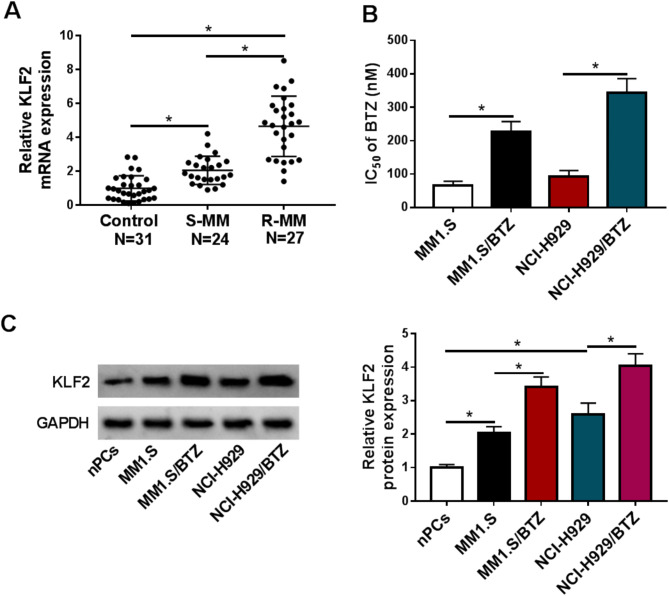
We first examined the expression of KLF2 in serum samples of patients with primary MM (called sensitive MM (S-MM) or recurrent MM after treatment based on BTZ therapy (R-MM). Through quantitative PCR, we discovered that S-MM serum samples had higher levels of KLF2 transcript than serum controls, and R-MM serum specimens exhibited elevated expression of KLF2 mRNA compared with S-MM controls (Fig. 1A), implying the potential implication of KLF2 dysregulation in MM pathogenesis and BTZ resistance. To determine the role of KLF2 in BTZ resistance of MM, we generated two BTZ-resistant MM cell lines (MM1.S/BTZ and NCI-H929/BTZ). MTT cell cytotoxicity assay confirmed the successful generation of BTZ-resistant MM cell lines, as presented by the elevated IC50 value for BTZ in MM1.S/BTZ and NCI-H929/BTZ cells compared with that in their parental cells (Fig. 1B). Similarly, KLF2 protein levels were strongly increased in MM1.S and NCI-H929 cells compared with non-tumor nPCs cells (Fig. 1C). Interestingly, MM1.S/BTZ and NCI-H929/BTZ cells exhibited higher levels of KLF2 protein than their parental cells (Fig. 1C). In addition, increased expression of KLF2 by OE-KLF2 introduction significantly elevated the IC50 value of MM1.S/BTZ and NCI-H929/BTZ cells (Supplementary Fig. 1), suggesting the significant association between KLF2 upregulation and BTZ resistance in MM cells.
Fig. 1.
KLF2 expression is upregulated in MM serum and BTZ-resistant MM cells. (A) Relative KLF2 mRNA expression (quantitative PCR) in serum samples of patients with primary MM (called sensitive MM (S-MM), n = 24) or recurrent MM after treatment based on BTZ therapy (R-MM, n = 27). (B) The IC50 value for BTZ (MTT cell cytotoxicity assay) in MM1.S, NCI-H929, MM1.S/BTZ and NCI-H929/BTZ cells (n = 3). (C) Relative expression of KLF2 protein (immunoblotting) in MM1.S, NCI-H929, MM1.S/BTZ and NCI-H929/BTZ cells (n = 3). *P < 0.05
Depletion of KLF2 suppresses cell growth and enhances apoptosis and BTZ sensitivity in MM1.S/BTZ and NCI-H929/BTZ cells
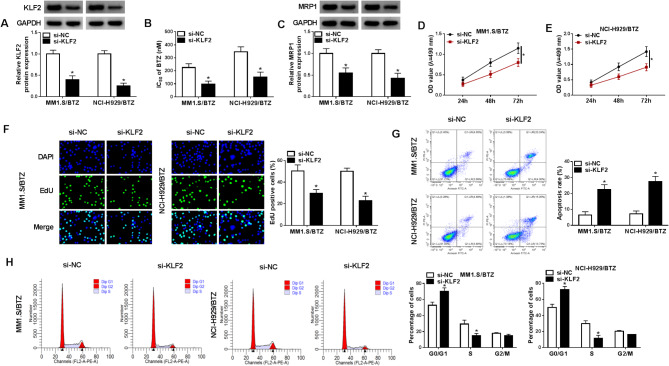
We then evaluated the precise functions of KLF2 in MM1.S/BTZ and NCI-H929/BTZ cells. To this end, we conducted siRNA silencing experiments using a KLF2-siRNA (si-KLF2) pool. The reduced effect of si-KLF2 on KLF2 expression was significant in MM1.S/BTZ and NCI-H929/BTZ cells (Fig. 2A). Following KLF2 decrease, the IC50 value for BTZ of MM1.S/BTZ and NCI-H929/BTZ cells diminished (Fig. 2B), indicating the promoting impact of KLF2 depletion on cell BTZ sensitivity. This result was also verified by detecting multidrug resistance protein 1 (MRP1) expression. In MM1.S/BTZ and NCI-H929/BTZ cells, KLF2 depletion resulted in reduced expression of MRP1 (Fig. 2C). In contrast, reduced expression of KLF2 significantly retarded cell proliferation (Fig. 2D, E and F). Flow cytometry analysis revealed that KLF2-depleted MM1.S/BTZ and NCI-H929/BTZ cells showed a dramatic increase in the apoptotic rate and the G0/G1 phase as compared with si-NC control cells (Fig. 2G and H). All these data indicate that KLF2 inhibition can retard cell growth and promote the anti-cancer efficacy of BTZ in MM1.S/BTZ and NCI-H929/BTZ cells.
Fig. 2.
KLF2 depletion impedes cell growth and enhances apoptosis and BTZ sensitivity in MM1.S/BTZ and NCI-H929/BTZ cells. (A) Relative expression of KLF2 protein (immunoblotting) in si-KLF2- or si-NC-introduced MM1.S/BTZ and NCI-H929/BTZ cells. (B) The IC50 value for BTZ (MTT cell cytotoxicity assay) in si-KLF2- or si-NC-introduced MM1.S/BTZ and NCI-H929/BTZ cells. (C) Relative MRP1 protein expression (immunoblotting) in KLF2-depleted MM1.S/BTZ and NCI-H929/BTZ cells. (D and E) MTT cell proliferation assay with KLF2-depleted MM1.S/BTZ and NCI-H929/BTZ cells and control cells. (F) The percentage of EdU positive cells (EdU incorporation assay) in KLF2-depleted MM1.S/BTZ and NCI-H929/BTZ cells and control cells. (G and H) The apoptotic rate (G) and cell cycle distribution (H) (Flow cytometry) in KLF2-depleted MM1.S/BTZ and NCI-H929/BTZ cells and control cells. n = 3 in A-H. *P < 0.05
USP4 increases the stability of KLF2 protein by deubiquitination and affects cell growth, apoptosis and BTZ sensitivity via KLF2
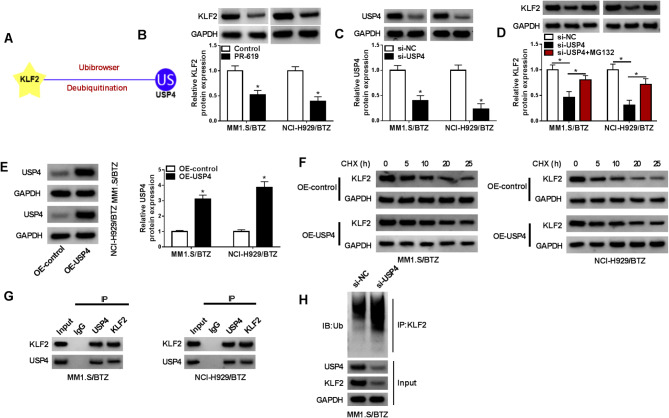
The ubiquitination/deubiquitination system has pivotal regulatory roles in cellular biology and human carcinogenesis by affecting protein stability and degradation [25]. Considering the upregulation of KLF2 in BTZ-resistant MM cells (Fig. 1C), we sought to identify the deubiquitinases that induce KLF2 dysregulation. Using the online algorithm Ubibrowser, we found that the deubiquitinase USP4, an established oncogenic factor in MM progression [22], was a potential regulator of KLF2 protein via deubiquitination (Fig. 3A). To validate the ubiquitination/deubiquitination modification in KLF2 protein, we first used PR-619, the general suppressor of deubiquitinating enzymes, to treat MM1.S/BTZ and NCI-H929/BTZ cells. Treatment of PR-619 remarkably decreased the protein levels of KLF2 in both cell lines (Fig. 3B). Second, we examined the regulation of USP4 in KLF2 protein expression. Reduced USP4 expression by si-USP4, verified by immunoblotting (Fig. 3C), led to a clear downregulation in KLF2 protein level in MM1.S/BTZ and NCI-H929/BTZ cells (Fig. 3D). However, treatment of MG132, a well-known inhibitor of ubiquitin-proteasome [26], reversed si-USP4-mediated KLF2 downregulation (Fig. 3D). We then analyzed the influence of USP4 in the stability of KLF2 protein. Introduction of OE-USP4, but not OE-control, strongly elevated USP4 expression in the two BTZ-resistance MM cell lines (Fig. 3E). Increased expression of USP4 in MM1.S/BTZ and NCI-H929/BTZ cells strikingly enhanced the stability of KLF2 protein in the presence of CHX (Fig. 3F). Through Co-IP experiments with an antibody specific for USP4 or KLF2, we confirmed the interplay between USP4 and KLF2 in MM1.S/BTZ and NCI-H929/BTZ cells (Fig. 3G). Next, the modulation of USP4 in KLF2 protein deubiquitination was tested. By contrast, si-USP4-driven USP4 depletion led to a distinct increase in ubiquitinated KLF2 levels in MM1.S/BTZ cells (Fig. 3H), indicating that USP4 inhibition can promote KLF2 protein ubiquitination and degradation. Taken together, these findings demonstrate that USP4 deubiquitinates and stabilizes KLF2 protein.
Fig. 3.
USP4 deubiquitinates and stabilizes KLF2 protein in MM1.S/BTZ and NCI-H929/BTZ cells. (A) The online algorithm Ubibrowser predicted the potential influence of USP4 in KLF2 protein deubiquitination. (B) KLF2 protein expression (immunoblotting) in MM1.S/BTZ and NCI-H929/BTZ cells treated with or without PR-619 (n = 3). (C) USP4 protein expression (immunoblotting) in MM1.S/BTZ and NCI-H929/BTZ cells transfected with si-NC or si-USP4 (n = 3). (D) Expression of KLF2 protein in transfected MM1.S/BTZ and NCI-H929/BTZ cells treated with or without MG132 (n = 3). (E) Expression of USP4 protein in OE-control- or OE-USP4-transfected MM1.S/BTZ and NCI-H929/BTZ cells (n = 3). (F) Immunoblotting of KLF2 protein in OE-control- or OE-USP4-transfected MM1.S/BTZ and NCI-H929/BTZ cells along with CHX treatment. (G) IP and Co-IP experiments with an antibody specific for USP4 or KLF2 in MM1.S/BTZ and NCI-H929/BTZ cells. (H) IP experiments with lysates of si-NC- or si-USP4-transfected MM1.S/BTZ and NCI-H929/BTZ cells using anti-KLF2 antibody and the subsequent immunblotting (IB) using anti-Ub antibody. *P < 0.05
Given our data that USP4 is responsible for KLF2 expression, we then carried out rescue experiments to investigate whether USP4 participates in MM BTZ resistance by regulating KLF2. Immunoblotting analysis verified that KLF2 increase could be achieved by transfecting OE-KLF2 into MM1.S/BTZ and NCI-H929/BTZ cells (Fig. 4A). Increased expression of KLF2 exerted a counteracting impact on USP4 depletion-mediated IC50 value reduction (Fig. 4B) and MRP1 expression inhibition (Fig. 4C) in both cell lines. Furthermore, KLF2 increase abolished si-USP4-mediated anti-proliferation (Fig. 4D, E and F), pro-apoptosis (Fig. 4G and H), as well as cell cycle arrest (Fig. 4I and J) compared with the control group. These results further support the regulatory effect of USP4 on cell growth, apoptosis and BTZ sensitivity through KLF2.
Fig. 4.
KLF2 increase partially abolishes USP4 depletion-mediated alterations on cell growth, apoptosis and BTZ sensitivity in MM1.S/BTZ and NCI-H929/BTZ cells. (A) KLF2 protein expression (immunoblotting) in OE-control- or OE-KLF2-transfected MM1.S/BTZ and NCI-H929/BTZ cells. (B) The IC50 value for BTZ (MTT cell cytotoxicity assay) in MM1.S/BTZ and NCI-H929/BTZ cells transfected with si-NC, si-USP4, or si-USP4 + OE-KLF2. (C) Relative MRP1 protein expression (immunoblotting) in transfected MM1.S/BTZ and NCI-H929/BTZ cells. (D and E) MTT cell proliferation assay with MM1.S/BTZ and NCI-H929/BTZ cells transfected as indicated. (F) The percentage of EdU positive cells (EdU incorporation assay) in transfected cells. (G-J) The apoptotic rate (G and H) and cell cycle distribution (I and J) (Flow cytometry) in MM1.S/BTZ and NCI-H929/BTZ cells transfected as indicated. n = 3 in A-J. *P < 0.05
KLF2 functions as a regulator of HMGA2 transcription and modulates cell growth, apoptosis and BTZ sensitivity through HMGA2
In view of the function of KLF2 in BTZ-resistant MM cells, we set out to elucidate how KLF2 dysregulation modulates cell phenotypes. The GSE55667 dataset revealed that multiple transcripts were markedly downregulated following KLF2 depletion in myeloma RPMI8226 cells (Fig. 5A). Among these transcripts, HMGA2 has established a role in promoting human carcinogenesis by influencing growth and therapy resistance [23]. Intriguingly, the open-access database JASPAR predicted the putative binding region for KLF2 within the HMGA2 promoter (Fig. 5B). Consistent with KLF2 expression profiling, we observed that HMGA2 protein expression was significantly elevated in MM1.S and NCI-H929 cells compared with non-tumor nPCs cells, and MM1.S/BTZ and NCI-H929/BTZ cells exhibited higher protein levels of HMGA2 than their parental cells (Fig. 5C). We then examined the modulation of KLF2 in HMGA2 expression. In contrast, KLF2 depletion by si-KLF2 led to a striking reduction in HMGA2 protein expression in MM1.S/BTZ and NCI-H929/BTZ cells (Fig. 5D), indicating the positive modulation of KLF2 in HMGA2 expression. Next, we tested the impact of KLF2 on HMGA2 transcription. Luciferase assays were conducted by generating HMGA2 luciferase reporters carrying the predicted binding site (WT-HMGA2, CACCACCCCCT) or its mutations (MUT-HMGA2, AGTTGAATTTC). With KLF2 deficiency, the luciferase activity of WT-HMGA2 significantly diminished in MM1.S/BTZ and NCI-H929/BTZ cells (Fig. 5E and F). However, reduced KLF2 expression did not alter the luciferase activity of MUT-HMGA2 (Fig. 5E and F). In addition, the relationship of KLF2 with the HMGA2 promoter in MM1.S/BTZ and NCI-H929/BTZ cells was validated by ChIP experiments using an antibody recognizing KLF2 (Fig. 5G).
Fig. 5.
KLF2 transcriptionally regulates HMGA2 expression. (A) The GSE55667 dataset revealed that HMGA2 was markedly downregulated following KLF2 depletion in myeloma RPMI8226 cells. (B) JASPAR database predicted the putative binding region for KLF2 within the HMGA2 promoter. (C) Expression of HMGA2 protein (immunoblotting) in nPCs, MM1.S, NCI-H929, MM1.S/BTZ and NCI-H929/BTZ cells (n = 3). (D) Expression of HMGA2 protein in si-NC- or si-KLF2-transfected MM1.S/BTZ and NCI-H929/BTZ cells (n = 3). (E and F) Luciferase assays with MM1.S/BTZ and NCI-H929/BTZ cells transfected with WT-HMGA2 or MUT-HMGA2 along with si-KLF2 or si-NC (n = 3). (G) ChIP experiments using an antibody recognizing KLF2 with lysates of MM1.S/BTZ and NCI-H929/BTZ cells (n = 3). *P < 0.05
To further evaluate the contribution of HMGA2 to the function of KLF2 in BTZ-resistant MM cells, we intended to increase HMGA2 expression in KLF2-depleted cells to observe the consequences. Immunoblot analysis showed that HMGA2 upregulation was achieved by OE-HMGA2 transfection (Fig. 6A). Upregulation of HMGA2 strongly reversed KLF2 depletion-imposed IC50 value reduction (Fig. 6B), MRP1 expression inhibition (Fig. 6C), cell proliferation defect (Fig. 6D, E and F), apoptosis promotion (Fig. 6G and H), as well as cell cycle progression suppression (Fig. 6I and J) in MM1.S/BTZ and NCI-H929/BTZ cells. Collectively, our findings demonstrate that KLF2 transcriptionally regulates HMGA2 expression to affect cell growth, apoptosis and BTZ sensitivity in MM1.S/BTZ and NCI-H929/BTZ cells.
Fig. 6.
KLF2 targets HMGA2 to affect cell growth, apoptosis and BTZ sensitivity in MM1.S/BTZ and NCI-H929/BTZ cells. (A) Expression of HMGA2 protein in MM1.S/BTZ and NCI-H929/BTZ cells transfected with OE-control or OE-HMGA2. (B) The IC50 value for BTZ (MTT cell cytotoxicity assay) in MM1.S/BTZ and NCI-H929/BTZ cells transfected with si-NC, si-KLF2, or si-KLF2 + OE-HMGA2. (C) MRP1 protein expression (immunoblotting) in transfected MM1.S/BTZ and NCI-H929/BTZ cells. (D and E) MTT cell proliferation assay with MM1.S/BTZ and NCI-H929/BTZ cells transfected as indicated. (F) The percentage of EdU positive cells (EdU incorporation assay) in transfected cells. (G-J) The apoptotic rate (G and H) and cell cycle distribution (I and J) (Flow cytometry) in MM1.S/BTZ and NCI-H929/BTZ cells transfected as indicated. n = 3 in A-J. *P < 0.05
USP4 modulates HMGA2 expression via KLF2
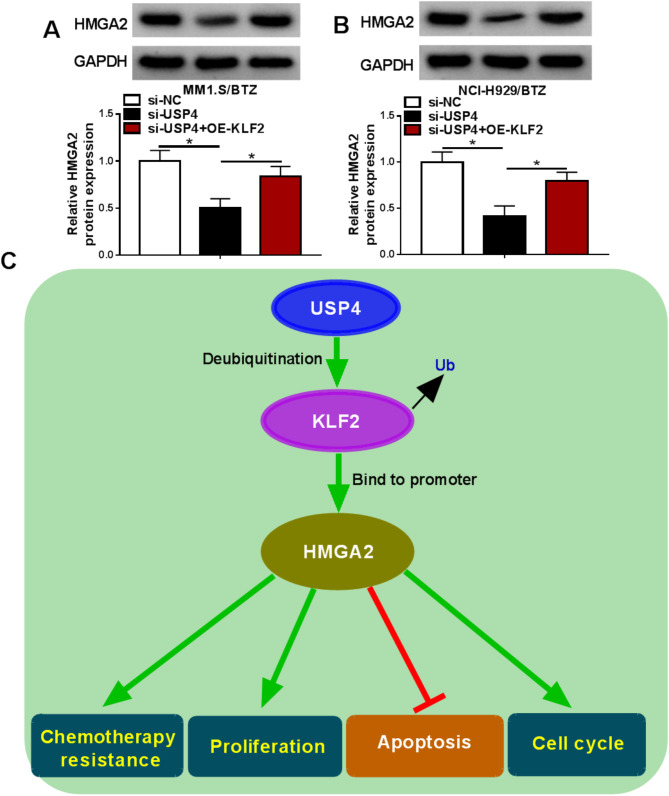
Finally, we examined whether USP4 can control HMGA2 expression through KLF2. In comparison with the si-NC control, si-UPS4-mediated USP4 depletion resulted in decreased expression of HMGA2 protein in MM1.S/BTZ and NCI-H929/BTZ cells (Fig. 7A and B). Furthermore, increased KLF2 expression by OE-KLF2 significantly abrogated USP4 depletion-driven HMGA2 downregulation (Fig. 7A and B). These data establish the novel USP4/KLF2/HMGA2 cascade in BTZ-resistant MM cells.
Fig. 7.
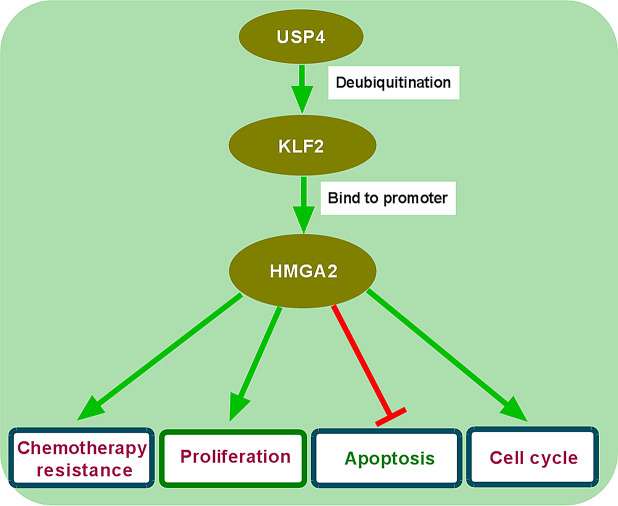
The USP4/KLF2/HMGA2 cascade in BTZ-resistant MM cells. (A and B) Expression of HMGA2 protein (immunoblotting) in MM1.S/BTZ and NCI-H929/BTZ cells transfected with si-NC, si-USP4, or si-USP4 + OE-KLF2 (n = 3). (C) In BTZ-resistant MM cells, USP4 stabilizes KLF2 protein via deubiquitination and thus enhances HMGA2 expression, thereby modulating cell sensitivity to BTZ, growth, and apoptosis. *P < 0.05
Discussion
Drug resistance remains a huge obstacle to the treatment of MM, a malignant disorder originating from plasma cells. Therefore, uncovering the mechanisms of drug resistance is imperative to improve the outcomes of patients with MM. Previous work has illuminated the oncogenic activity of KLF2 in MM by promoting cancer cell growth and hindering apoptosis [17, 18]. Considering the increased impact of BTZ therapy on KLF2 expression in MM [17], we hypothesized that KLF2 is involved in BTZ resistance development in MM. By generating MM1.S/BTZ and NCI-H929/BTZ cells, we have observed that BTZ-resistant MM cells exhibit enhanced expression of KLF2 compared with the corresponding sensitive counterparts. Using siRNA-mediated depletion experiments, we demonstrate that silencing KLF2 can retard cell growth and enhances apoptosis and BTZ sensitivity in MM1.S/BTZ and NCI-H929/BTZ cells. Thus, KLF2 inhibition may be a potential strategy for promoting the anti-cancer efficacy of BTZ in MM.
USP4, a key deubiquitinase, mediates post-translational modification in proteins and controls their stability [21]. Growing evidence has highlighted the multifaceted roles of USP4 in tumor biology [27]. For instance, USP4 functions as a pro-tumorigenic factor in esophageal squamous cell carcinoma by stabilizing the TAK1 protein in a deubiquitination-dependent way [28]. USP4 deubiquitinates and stabilizes β-catenin to drive colon cancer cell migration [29]. Upregulated expression of USP4 has been discovered to forebode a worse prognosis in MM [30]. Moreover, USP4 upregulation contributes to enhanced proliferation in MM cells [22]. Our results show that in MM1.S/BTZ and NCI-H929/BTZ cells, USP4 can increase the stability of KLF2 protein by deubiquitination. Specifically, we have established that USP4 affects cell growth, apoptosis, and sensitivity to BTZ via its modulation of KLF2. These results contribute to a deeper understanding of the complex regulatory networks involving USP4 and KLF2 in MM.
Functioning in chromatin remodeling and gene expression regulation, HMGA2 exerts tumor-promoting activity by influencing cancer cell growth and therapy resistance [23]. As an example, HMGA2 is capable of enhancing colorectal tumorigenesis by targeting the STAT3/CCL2 axis [31]. Additionally, HMGA2 affects drug sensitivity in gastric cancer cells depending on the mTOR/P-gp axis [32]. Through the microarray method, HMGA2 is predicted as a player involved in MM pathogenesis [33]. Nonetheless, no reports showed the action of HMGA2 on MM resistance to BTZ. Here, we unveil a novel regulatory mechanism where KLF2 functions as a key regulator of HMGA2 transcription by binding to the HMGA2 promoter, highlighting a previously unknown layer of complexity in the regulatory network governing HMGA2 expression. Furthermore, we point out, for the first time, that KLF2 modulates cell growth, apoptosis, and sensitivity to BTZ through HMGA2.
Our study further elucidates the intricate regulatory network in BTZ-resistant MM cells by revealing that USP4 modulates HMGA2 expression via KLF2, establishing a previously uncharacterized USP4/KLF2/HMGA2 cascade in MM resistance to BTZ. However, it is important to note that our current understanding of this cascade is primarily based on in vitro studies. There is a lack of in vivo evidence to fully support the role of this regulatory axis in MM BTZ resistance. Therefore, future research should aim to address this gap by conducting comprehensive in vivo studies to validate the findings. Although we show the regulation of USP4 in KLF2 stability via deubiquitination, the specific mechanisms are not defined. Uncovering the ubiquitination site of KLF2 and the deubiquitination functional domain of USP4 in the regulation of KLF2 would provide significant insights and be valuable for studying the role of KLF2 in MM resistance to BTZ. In addition, HMGA2 possesses pivotal functions in chromatin remodeling and gene expression regulation. Future work will be warranted to uncover the downstream effectors underlying the activity of HMGA2.
In summary, our study demonstrates a novel regulatory network where USP4 stabilizes KLF2 protein via deubiquitination and thus enhances HMGA2 expression, with the ability to modulate BTZ sensitivity, growth, and apoptosis of BTZ-resistant MM cells (Fig. 7C). We therefore suggest that targeting this regulatory cascade may be an encouraging strategy to improve the anti-MM efficacy of BTZ.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplementary Material 1: Supplementary Figure 1. The IC50 value for BTZ (MTT cell cytotoxicity assay) in MM1.S/BTZ and NCI-H929/BTZ cells after transfection with OE-control or OE-KLF2 (n=3). *P < 0.05.
Acknowledgements
None.
Author contributions
Conceptualization and Methodology: Xiaoliang Li and Yin He; Formal analysis and Data curation: Fengqun Xie, Ziyi Liu and Yanrong Zhang; Validation and Investigation: Mufang Huang, Yinjingwen Wang and Weijian Zhu; Writing - original draft preparation and Writing - review and editing: Ziyi Liu, Xiaoliang Li and Yin He; Approval of final manuscript: all authors.
Funding
None.
Data availability
No datasets were generated or analysed during the current study.
Declarations
Competing interests
The authors declare no competing interests.
Footnotes
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
References
- 1.Cowan AJ, Green DJ, Kwok M, Lee S, Coffey DG, Holmberg LA, Tuazon S, Gopal AK, Libby EN. Diagnosis and management of multiple myeloma: a review. JAMA. 2022;327(5):464–77. [DOI] [PubMed] [Google Scholar]
- 2.Rajkumar SV. Multiple myeloma: 2024 update on diagnosis, risk-stratification, and management. Am J Hematol. 2024;99(9):1802–24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Rees MJ, Kumar S. High-risk multiple myeloma: redefining genetic, clinical, and functional high-risk disease in the era of molecular medicine and immunotherapy. Am J Hematol. 2024;99(8):1560–75. [DOI] [PubMed] [Google Scholar]
- 4.Long S, Long S, He H, Luo L, Liu M, Ding T. Exosomal miR-182 derived from bone marrow mesenchymal stem cells drives carfilzomib resistance of multiple myeloma cells by targeting SOX6. J Orthop Surg Res. 2023;18(1):937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Zeng P, Wang F, Long X, Cao Y, Wen F, Li J, Luo Z. CPEB2 enhances cell growth and angiogenesis by upregulating ARPC5 mRNA stability in multiple myeloma. J Orthop Surg Res. 2023;18(1):384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Zhang Q, Duan H, Yang W, Liu H, Tao X, Zhang Y. Circ_0005615 restrains the progression of multiple myeloma through modulating mir-331-3p and IGF1R regulatory cascade. J Orthop Surg Res. 2023;18(1):356. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Lucca LE. Multiple myeloma treatment: one bridge closer. Blood. 2023;142(21):1763–4. [DOI] [PubMed] [Google Scholar]
- 8.Abhijit D, Rupa M, Avijit M, Rajesh S, Shikha S. <i > in silico identification of HDAC inhibitors for multiple myeloma: a structure-based virtual screening, drug likeness, ADMET profiling, Molecular Docking, and Molecular Dynamics Simulation Study</i >. Lett Drug Des Discovery. 2024;21(5):961–78. [Google Scholar]
- 9.Sonneveld P, Dimopoulos MA, Boccadoro M, Quach H, Ho PJ, Beksac M, Hulin C, Antonioli E, Leleu X, Mangiacavalli S, et al. Daratumumab, Bortezomib, Lenalidomide, and Dexamethasone for multiple myeloma. N Engl J Med. 2024;390(4):301–13. [DOI] [PubMed] [Google Scholar]
- 10.Tang X, Wang P, Zhang R, Watanabe I, Chang E, Vinayachandran V, Nayak L, Lapping S, Liao S, Madera A, et al. KLF2 regulates neutrophil activation and thrombosis in cardiac hypertrophy and heart failure progression. J Clin Investig. 2022;132(3). [DOI] [PMC free article] [PubMed]
- 11.Tamargo IA, Baek KI, Xu C, Kang DW, Kim Y, Andueza A, Williams D, Demos C, Villa-Roel N, Kumar S, et al. HEG1 protects against atherosclerosis by regulating stable Flow-Induced KLF2/4 expression in endothelial cells. Circulation. 2024;149(15):1183–201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Lu Y, Qin H, Jiang B, Lu W, Hao J, Cao W, Du L, Chen W, Zhao X, Guo H. KLF2 inhibits cancer cell migration and invasion by regulating ferroptosis through GPX4 in clear cell renal cell carcinoma. Cancer Lett. 2021;522:1–13. [DOI] [PubMed] [Google Scholar]
- 13.Li J, Jiang JL, Chen YM, Lu WQ. KLF2 inhibits colorectal cancer progression and metastasis by inducing ferroptosis via the PI3K/AKT signaling pathway. J Pathol Clin Res. 2023;9(5):423–35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Huo FC, Zhu ZM, Zhu WT, Du QY, Liang J, Mou J. METTL3-mediated m(6)a methylation of SPHK2 promotes gastric cancer progression by targeting KLF2. Oncogene. 2021;40(16):2968–81. [DOI] [PubMed] [Google Scholar]
- 15.Wittner J, Schuh W. Krüppel-like factor 2: a central regulator of B cell differentiation and plasma cell homing. Front Immunol. 2023;14:1172641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Li Y, Zhang L, Gong J. Relation among EGFL7, ITGB3, and KLF2 and their clinical implication in multiple myeloma patients: a prospective study. Ir J Med Sci. 2022;191(5):1995–2001. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Salama Y, Heida AH, Yokoyama K, Takahashi S, Hattori K, Heissig B. The EGFL7-ITGB3-KLF2 axis enhances survival of multiple myeloma in preclinical models. Blood Adv. 2020;4(6):1021–37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ohguchi H, Hideshima T, Bhasin MK, Gorgun GT, Santo L, Cea M, Samur MK, Mimura N, Suzuki R, Tai YT, et al. The KDM3A-KLF2-IRF4 axis maintains myeloma cell survival. Nat Commun. 2016;7:10258. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Nayak L, Shi H, Atkins GB, Lin Z, Schmaier AH, Jain MK. The thromboprotective effect of bortezomib is dependent on the transcription factor kruppel-like factor 2 (KLF2). Blood. 2014;123(24):3828–31. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Liu F, Chen J, Li K, Li H, Zhu Y, Zhai Y, Lu B, Fan Y, Liu Z, Chen X, et al. Ubiquitination and deubiquitination in cancer: from mechanisms to novel therapeutic approaches. Mol Cancer. 2024;23(1):148. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Hu B, Zhang D, Zhao K, Wang Y, Pei L, Fu Q, Ma X. Spotlight on USP4: structure, function, and Regulation. Front cell Dev Biology. 2021;9:595159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Chen CJ, Huang JY, Huang JQ, Deng JY, Shangguan XH, Chen AZ, Chen LT, Wu WH. Metformin attenuates multiple myeloma cell proliferation and encourages apoptosis by suppressing METTL3-mediated m6A methylation of THRAP3, RBM25, and USP4. Cell Cycle (Georgetown Tex). 2023;22(8):986–1004. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Hashemi M, Rashidi M, Hushmandi K, Ten Hagen TLM, Salimimoghadam S, Taheriazam A, Entezari M, Falahati M. HMGA2 regulation by miRNAs in cancer: affecting cancer hallmarks and therapy response. Pharmacol Res. 2023;190:106732. [DOI] [PubMed] [Google Scholar]
- 24.Rizzolio S, Cagnoni G, Battistini C, Bonelli S, Isella C, Van Ginderachter JA, Bernards R, Di Nicolantonio F, Giordano S, Tamagnone L. Neuropilin-1 upregulation elicits adaptive resistance to oncogene-targeted therapies. J Clin Investig. 2018;128(9):3976–90. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Cockram PE, Kist M, Prakash S, Chen SH, Wertz IE, Vucic D. Ubiquitination in the regulation of inflammatory cell death and cancer. Cell Death Differ. 2021;28(2):591–605. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Sulkshane P, Duek I, Ram J, Thakur A, Reis N, Ziv T, Glickman MH. Inhibition of proteasome reveals basal mitochondrial ubiquitination. J Proteom. 2020;229:103949. [DOI] [PubMed] [Google Scholar]
- 27.Wang Y, Zhou L, Lu J, Jiang B, Liu C, Guo J. USP4 function and multifaceted roles in cancer: a possible and potential therapeutic target. Cancer Cell Int. 2020;20:298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Zhang H, Han Y, Xiao W, Gao Y, Sui Z, Ren P, Meng F, Tang P, Yu Z. USP4 promotes the proliferation, migration, and invasion of esophageal squamous cell carcinoma by targeting TAK1. Cell Death Dis. 2023;14(11):730. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Yun SI, Kwak C, Lee SY, Shin S, Oh C, Kim JS, Rhee HW, Kim KK. Binding of USP4 to cortactin enhances cell migration in HCT116 human colon cancer cells. FASEB Journal: Official Publication Federation Am Soc Experimental Biology. 2023;37(5):e22900. [DOI] [PubMed] [Google Scholar]
- 30.Kumari R, Majumder MM, Lievonen J, Silvennoinen R, Anttila P, Nupponen NN, Lehmann F, Heckman CA. Prognostic significance of esterase gene expression in multiple myeloma. Br J Cancer. 2021;124(8):1428–36. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Wang X, Wang J, Zhao J, Wang H, Chen J, Wu J. HMGA2 facilitates colorectal cancer progression via STAT3-mediated tumor-associated macrophage recruitment. Theranostics. 2022;12(2):963–75. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Jiang L, Zhang Y, Guo L, Liu C, Wang P, Ren W. Exosomal microRNA-107 reverses chemotherapeutic drug resistance of gastric cancer cells through HMGA2/mTOR/P-gp pathway. BMC Cancer. 2021;21(1):1290. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Yyusnita N, Zakiah I, Chang KM, Purushotaman VS, Zubaidah Z, Jamal R. MicroRNA (miRNA) expression profiling of peripheral blood samples in multiple myeloma patients using microarray. Malays J Pathol. 2012;34(2):133–43. [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Material 1: Supplementary Figure 1. The IC50 value for BTZ (MTT cell cytotoxicity assay) in MM1.S/BTZ and NCI-H929/BTZ cells after transfection with OE-control or OE-KLF2 (n=3). *P < 0.05.
Data Availability Statement
No datasets were generated or analysed during the current study.