Fig. 5.
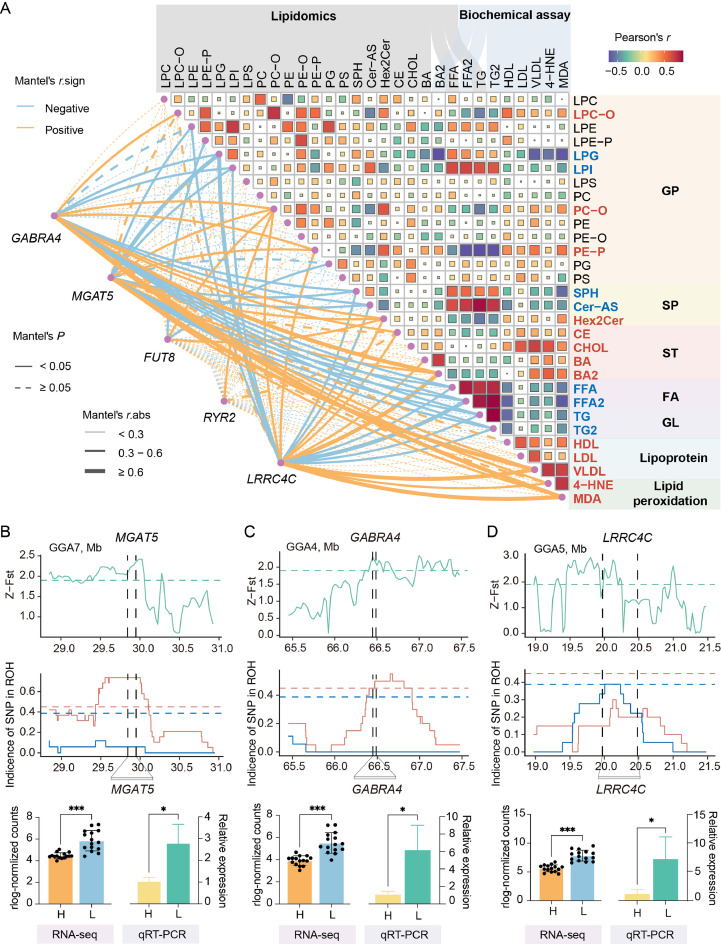
Important candidate genes and lipids associated with RFI. (A) The relationship between lipid profiles and key candidate genes. Lipid data were obtained from lipidomics and biochemical assays. The heatmap on the right displays the Pearson correlation coefficients between lipid pairs, where red and blue squares represent positive and negative correlations, respectively. The size of each square reflects the absolute value of the correlation coefficient. Lipid names in red or blue indicate positive or negative correlations, respectively, with lipid transport or oxidative stress indicators. The lines on the left represent the correlation between candidate gene expression and lipid content. Orange and blue lines indicate positive and negative correlations, respectively, with line thickness representing the strength of the correlation. Solid lines indicate significant correlations (P value < 0.05), while dashed lines indicate non-significant correlations (P value ≥ 0.05). (B-D) Selection sweep regions near the MGAT5 (B), GABRA4 (C), and LRRC4C (D) genes, and their expression levels in high and low RFI lines using RNA-seq and qRT-PCR. The values are the mean ± SEM, * represents P < 0.05, ** represents P < 0.01 and *** represents P < 0.001.