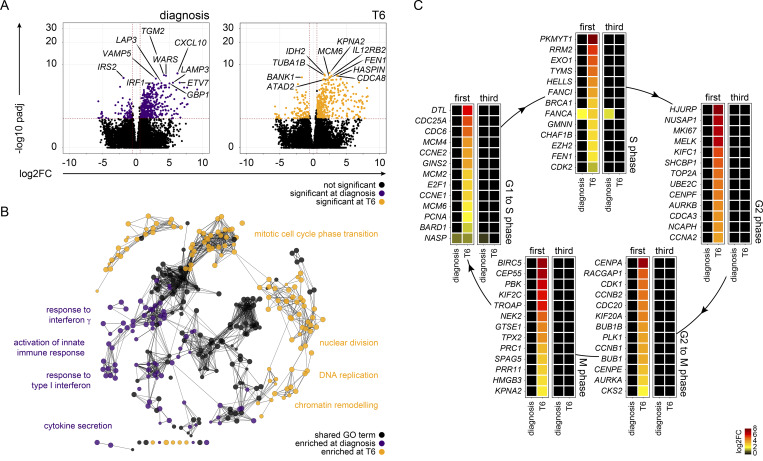
Figure 3.
Inflammation is followed by proliferation in the circulation. (A) RNAseq was used to identify differentially expressed genes in whole blood at diagnosis and T6 (versus baseline) during first infection in the VAC063C study (adj P < 0.05 and >1.5 fold-change [FC]). Volcano plots show all differentially expressed genes (colored dots), and the dashed lines represent the significance/fold-change cutoffs. The top 10 differentially expressed genes (lowest adj P) at each time point are labeled. (B) Differentially expressed genes at T6 and diagnosis were combined for GO analysis of first infection and ClueGO was used to construct a merged functional gene ontology network. Each node represents a GO term and nodes are colored according to whether their associated genes were majoritively (>60%) derived from the diagnosis or T6 time point. GO terms that were shared between time points are shown in black. Four leading GO terms (each from a unique functional group) are labeled for each time point. (C) Heatmaps show differentially expressed genes at diagnosis and T6 (versus baseline) during first and third infection (adj P < 0.05 and >1.5 fold-change). The log2 fold-change of key genes associated with each phase of the cell cycle is shown. Non-significant genes are displayed with a log2 fold-change of zero. In A and B, n = 3 (first infection only), and in C, n = 3 (first infection) and n = 6 (third infection).