Figure 5.
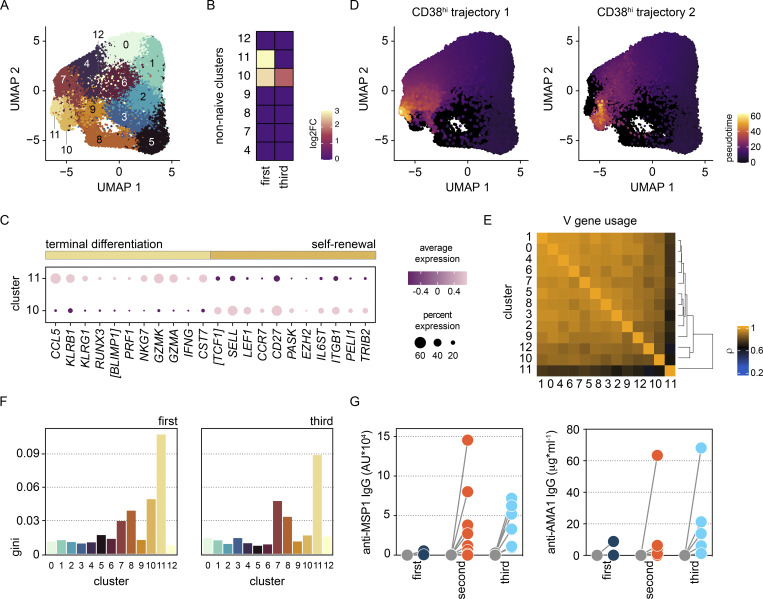
Stem-like memory CD4 + T cells respond to rechallenge. Droplet-based single-cell RNAseq was carried out during VAC063C on flow-sorted CD4+ T cells obtained at baseline and T6 from volunteers undergoing their first or third infection of life. (A) Data from all volunteers and time points was concatenated for UMAP analysis and FlowSOM identified 13 discrete clusters of CD4+ T cells across the dataset (each given a unique color). (B) Heatmap showing the differential abundance of non-naive CD4+ T cell clusters at T6 (versus baseline) in first and third infection (FDR < 0.05). Note that non-significant clusters are shown with a log2 fold-change (FC) of zero and the identification of non-naive clusters is shown in Fig. S3 C. (C) Dot plot showing differentially expressed signature genes in clusters 10 and 11 (adj P < 0.05). Square brackets indicate that common protein names have been used. (D) Trajectory inference with Slingshot revealed clusters 10 and 11 as discrete non-overlapping endpoints of CD4+ T cell activation and differentiation (analysis was performed on concatenated data and CD38hi cells were set as the endpoint; Slingshot identified two possible non-overlapping routes). (E) Spearman correlation matrix showing shared V gene usage (TRAV and TRBV) across all CD4+ T cell clusters (the order of features was determined by unsupervised hierarchical clustering). (F) Gini plot showing the equality of V gene usage in each CD4+ T cell cluster in the first and third infection; zero denotes perfect equality, which indicates a diverse TCR repertoire. (G) Class-switched antibodies (IgG) recognizing the malaria antigens MSP1 and AMA1 were quantified in serum by ELISA at baseline (grey dots) and 1 mo after challenge (colored dots). Samples were obtained from volunteers undergoing their first, second, or third infection during VAC063A, VAC063B, and VAC063C, respectively, and these data are presented in Salkeld et al. (2022). In A–F, n = 3 for the first and third infections, whereas in G, n = 10 for first and second infections and n = 6 in third infection.