Figure S2.
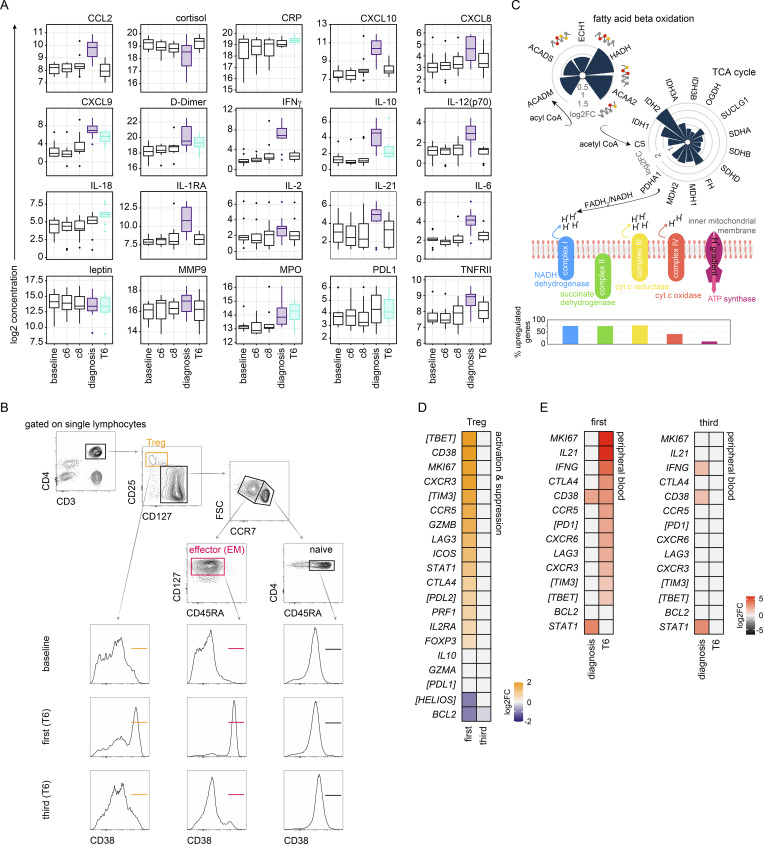
Activated T cells are released into circulation as inflammation resolves. (A) Mixed-effects models and linear regression were used to identify plasma analytes that vary significantly at diagnosis and/or T6 across the entire VAC063C dataset (all volunteers regardless of infection number). Kenward Roger approximation was used to calculate P values and multiple test correction was performed using the Benjamini–Hochberg method; significance (adj P < 0.05) is indicated by colored box plots (purple at diagnosis and turquoise at T6). Box (median and IQR) and whisker (1.5× upper or lower IQR) plots are shown with outliers as dots. (B) Gating strategy for sorting CD4+ T cell subsets during VAC063C. CD4+ T cells were sorted ex vivo (within 2 h of blood draw) into TRIzol for downstream RNAseq, and cells with a naive, effector (effector memory [EM]), or regulatory phenotype were sorted as shown at baseline and T6. Note that we did not use CD38 for sorting but subsequently used this marker to assess the level of activation within each subset at both time points. (C) RNAseq was used to analyze transcriptional regulation of fatty acid β-oxidation, the tricarboxylic acid (TCA) cycle, and oxidative phosphorylation (oxphos) in flow-sorted effector (effector memory) CD4+ T cells during first infection. The circular bar charts show the log2 fold-change (FC) of each major enzyme involved in fatty acid β-oxidation and the TCA cycle clockwise in reaction order 6 days after parasite clearance (T6 versus baseline). The vertical bar chart shows the proportion of oxphos enzymatic subunits that are transcriptionally upregulated at T6; all subunits required to form complex I to IV in the electron transport chain and ATP synthase are shown. The key molecules that connect these metabolic pathways are labeled. (D) RNAseq was used to identify differentially expressed genes in flow-sorted regulatory T cells (Treg) (CD4pos CD25hi CD127neg) in the first and third infection (T6 versus baseline) (adj P <0.05 and >1.5 fold-change). Heatmap shows the log2 fold-change of differentially expressed genes that control Treg activation and suppressor function. (E) RNAseq was used to identify differentially expressed genes in whole blood at diagnosis and T6 (relative to baseline) in the first and third infection (adj P < 0.05 and >1.5 fold-change). Signature genes associated with T cell activation, TH1 polarization, and cytokine production are shown. In A, n = 10 (3 first infection, 2 second infection, and 5 third infection). v1040 was excluded from plasma analysis because all samples failed QC. In C and D, n = 2 or 3 for the first infection (T6 and baseline, respectively) and n = 6 for the third infection (v313 was excluded at T6 because this sample failed QC). In E, n = 3 for the first infection and n = 6 for the third infection. In D and E, non-significant genes are displayed with a log2 fold-change of zero and square brackets indicate that common protein names have been used.