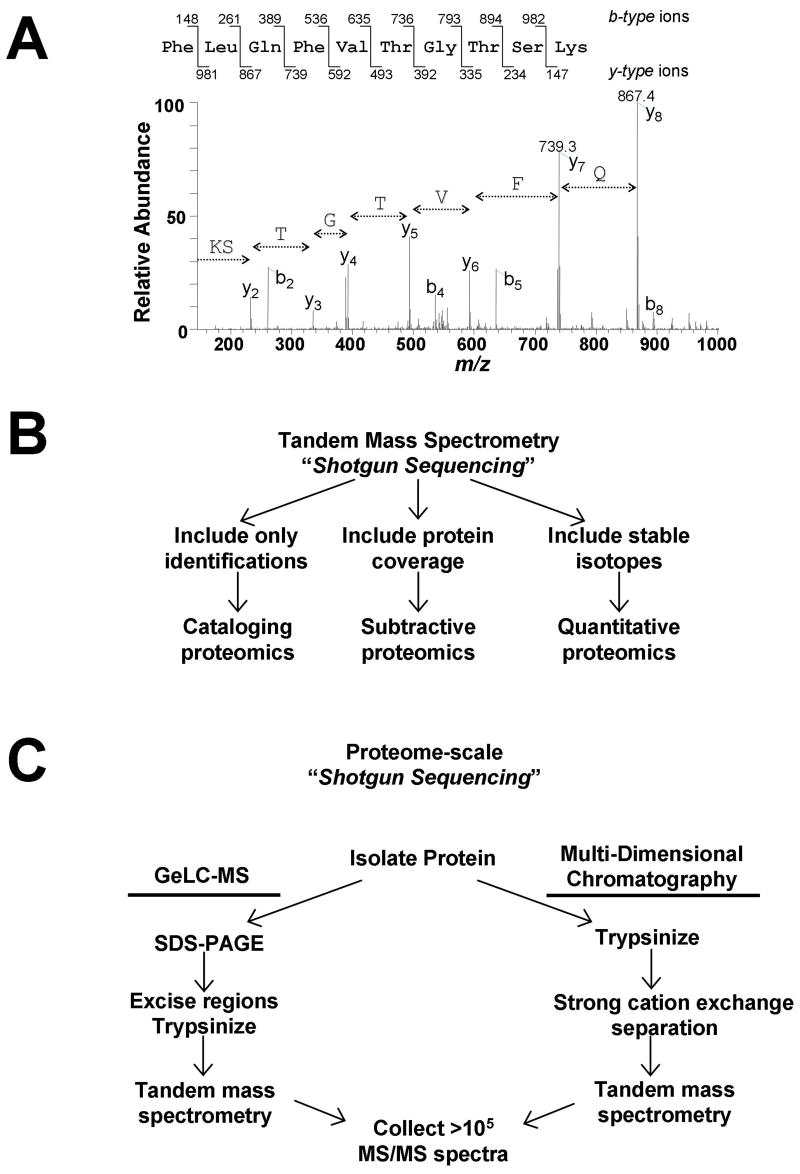
Figure 2.
Overview of shotgun sequencing from complex mixtures by mass spectrometry. a) Representative tandem mass (MS/MS) spectrum of a peptide from the protein Urebp1. Amino acid sequence for a single peptide can be deduced from the series of fragment ions present in the spectrum. b) Large-scale peptide detection via shotgun sequencing can be interpreted in three different ways to provide either lists (cataloging proteomics), differential identifications (subtractive proteomics), or abundance comparisons (quantitative proteomics). c) For proteome-scale analyses, huge numbers of MS/MS spectra are collected. Only two routes have proven successful for identifying thousands of proteins from a single sample. Both strategies utilize multiple[dk33] steps to fractionate the original sample prior to MS analysis. In one strategy, SDS-PAGE separation followed by tandem MS analysis of many gel regions (sometimes called GeLC-MS) is used. When using multiple-dimensional chromatography, protein mixtures are directly proteolyzed, and the peptide mixture is separated first by strong cation exchange [dk34]chromatography. In both cases, the final step involves reversed-phase separation of peptides from multiple samples, followed by tandem MS analysis of multiple samples. Both techniques provide the opportunity to collect hundreds of thousands of MS/MS spectra from a single sample in less than 24 hrs.