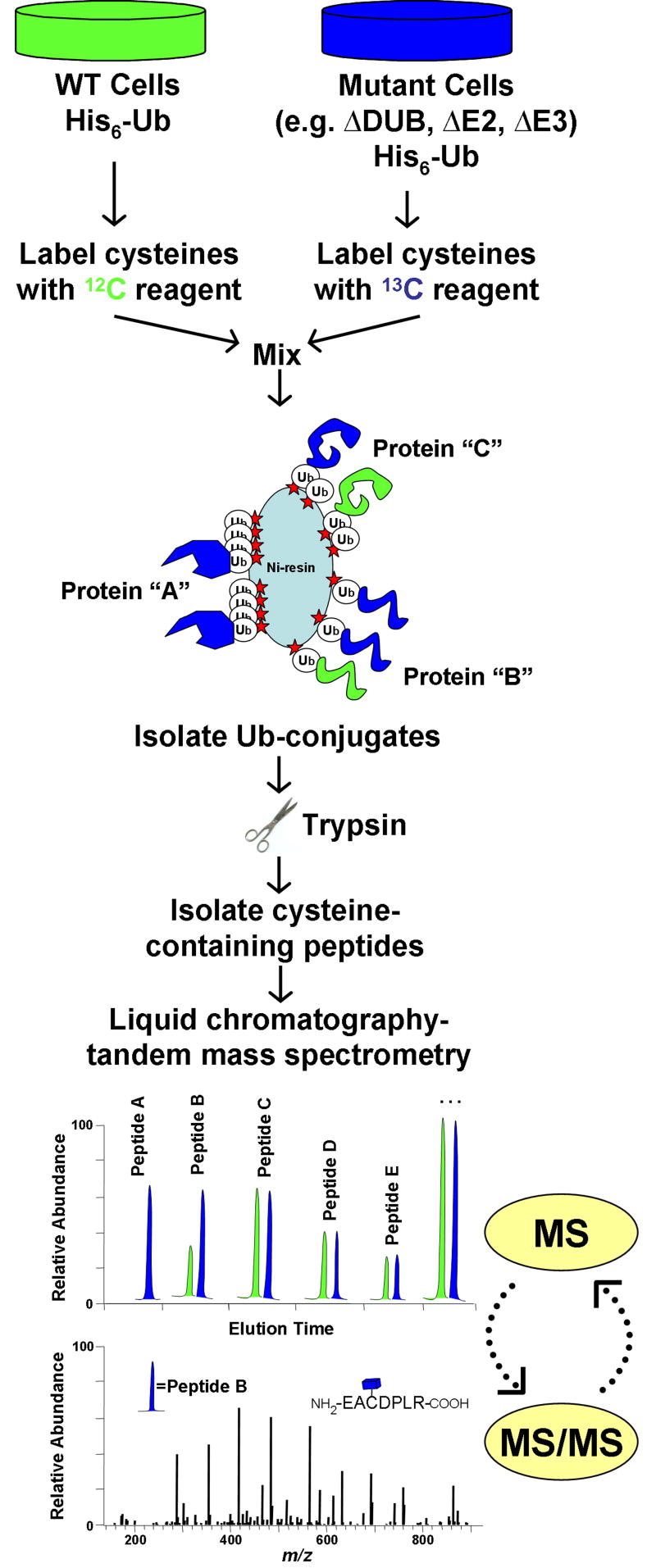
Figure 3.
Quantitative profiling of ubiquitin-conjugates using stable isotopes. The isotope coded affinity tag (ICAT) strategy is shown for comparing ubiquitinated proteins between wild type cells and mutant cells lacking a ubiquitin pathway enzyme (e.g. DUB, E2, E3). In an ICAT experiment, protein is harvested from two samples and differentially labeled at cysteinyl residues with either a 12C- or 13C-containing reagent. After labeled proteins are mixed, ubiquitinated-proteins are affinity-purified and digested into peptides. The ICAT label allows for further enrichment of cysteine-containing peptides, thus eliminating all peptides derived from ubiquitin. Since 12C- and 13C-containing peptides co-elute by reverse-phase chromatography, they can be simultaneously quantified during MS analysis. Rapid cycling between MS and MS/MS modes allows for the acquisition of both sequence and abundance information for isotopic peptide pairs. In this example, protein “A” is exclusively ubiquitinated in mutant cells, while protein “B” is ubiquitination is increased in mutant cells. Protein “C” is ubiquitinated equally in both samples. Other potential strategies would utilize similar work-flows with minor modifications. For example, metabolic labeling (e.g. SILAC) involves incorporation of isotopes into living cells prior to harvest, and would not utilize the cysteine-enrichment step. A benefit of metabolic labeling would be quantification of non-cysteine containing peptides, which would include most –GG signature peptides.