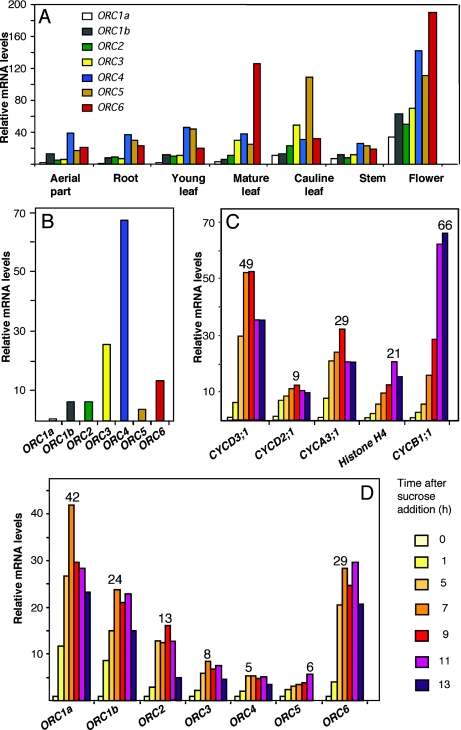
Figure 2.
Organ- and cell cycle-dependent expression of AtORC genes. (A) Expression pattern of AtORC genes in different organs. Measurements were normalized to the amount of UBQ10 or ACT2 and, then all the AtORC values made relative to the amount of AtORC1a present in the sample of aerial part of these seedlings (the lowest of all). Samples were prepared from aerial parts and root system of 12 day-old seedlings, young and mature rosette leaves, cauline leaves, stems, flowers at different stages or growth. (B) A.thaliana MM2d suspension cultured cells were sucrose-starved for 24 h and the amount of different AtORC mRNAs was determined by real-time RT–PCR, using the normalization procedure described for panel A. (C–D) A.thaliana MM2d suspension cultured cells, sucrose-starved for 24 h, were stimulated to re-enter the cell cycle, as described (19). The amount of mRNA of several cell cycle marker genes (31) was determined at the indicated times after sucrose addition by real-time RT–PCR, as described in panel A. CYCD3;1 and CYCD2;1 were used as G1 markers, CYCA3;1 and histone H4, as S-phase markers and CYCB1;1, as a G2/M marker (panel C). The mRNA levels of each AtORC gene (panel D) were determined at the indicated times after sucrose addition by real-time RT–PCR, as described in panel A. Numbers on top of the bars in panels C and D indicate the fold increase at the maximum level of expression relative to the value, in each case, obtained at time zero (arrested cells). In all cases, the RT–PCR measurements were repeated, at least, 2–3 times but error bars have been omitted for simplicity.