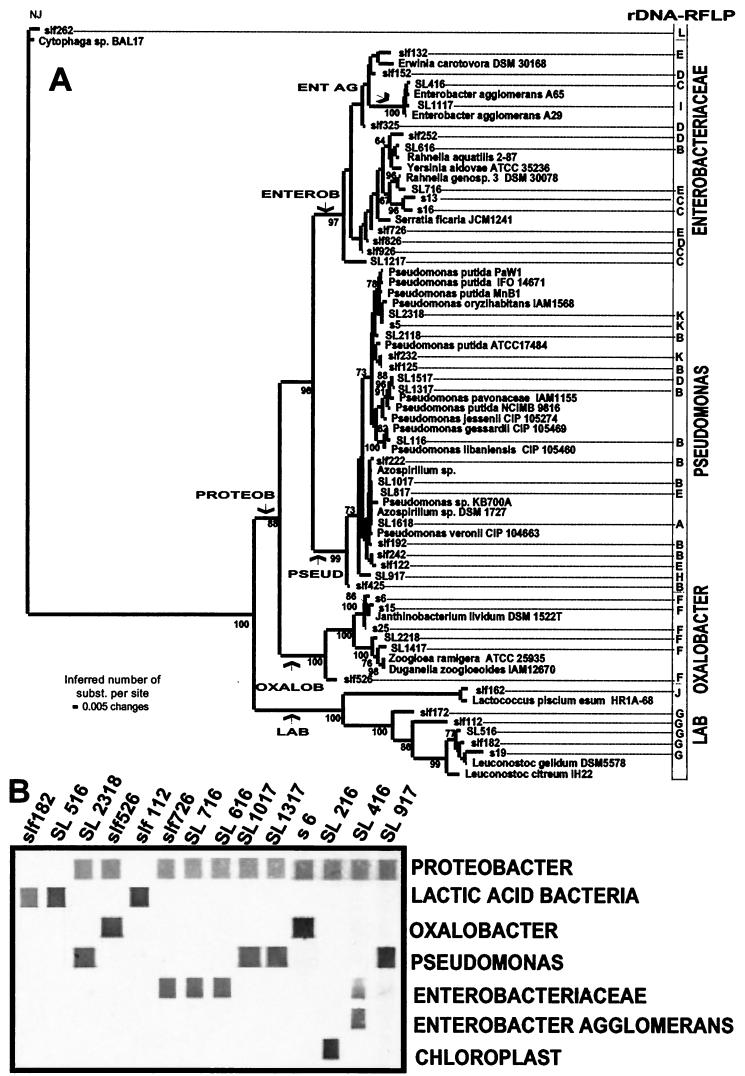
FIG.3.
Phylogenetic distance tree for 16S rDNA sequences (A) and specific detection with taxonomic probes (B). The distance tree (A) was built by using LogDet distances and neighbor-joining analysis for tree construction (31). The statistical support of the branches in the tree was tested by bootstrap analysis (11). The distance between two organisms, expressed in substitutions (subst.) per nucleotide, is obtained by adding the horizontal branches connecting them. The numbers at the nodes indicate the percentages of 500 bootstrap trees in which the cluster descending from the node was found. The arrows indicate the phylogenetic positions of the constructed probes. A selection of clones representing the major lineages in the phylogenetic tree shown in panel A was used for the probe labeling assay (B). RFLP, restriction fragment length polymorphism.