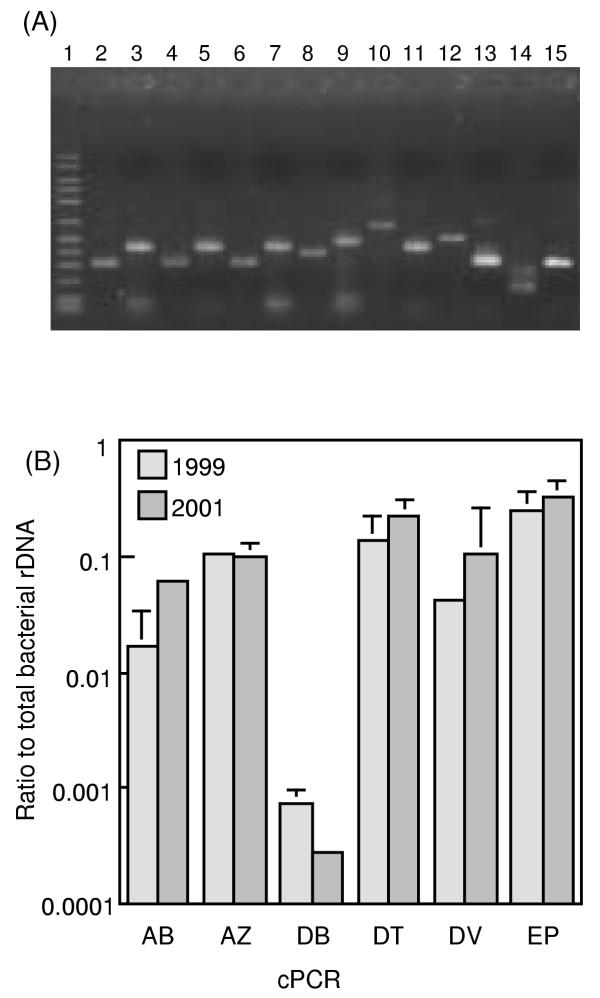
Figure 2.
cPCR assays for quantifying rDNA copies of major sequence types. The 6 cPCR systems used, namely AB, AZ, DB, DT, DV and EP, are explained in Table 3. (A) Target fragments amplified by PCR from the groundwater DNA (obtained in 1999) and competitor fragments. Lane 1, 50–2500 molecular size marker (FMC corp.); lanes 2, 4, 6, 8, 10, 12 and 14, target fragments; lanes 3, 5, 7, 9, 11, 13 and 15; competitor fragments. Lanes 2 and 3, DV; lanes 4 and 5, DB; lanes 6 and 7, AZ; lanes 8 and 9, EP; lanes 10 and 11, AB; lanes 12 and 13, DT; lanes 14 and 15, BC. (B) Rations of specific rDNA copies to the total bacterial rDNA copies (determined by the BC system in Table 3) in DNA extracted from the cavity groundwater. The total bacterial counts estimated by FISH were approximately 3 × 106 cells ml-1. The mean of three determinations is shown, and an error bar indicates a standard deviation.