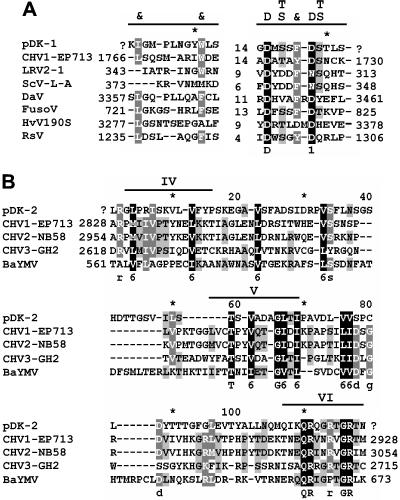
FIG. 3.
Conserved sequence motifs in the RNA-dependent RNA polymerases (RdRp) (A) and helicases (B) of positive-strand RNA viruses, related dsRNA viruses, and the 7.5-kb dsRNA of strain DK21. The tentative superpositions of the conserved motifs correspond to the motifs described by others (21, 30, 48). The number of amino acid residues between the motifs and the distances from the protein termini are shown (question marks show that numbers are unknown). In the consensus line above the sequences, “&” designates a bulky hydrophobic residue (either aliphatic or aromatic). GenBank accession numbers are as follows: CHV1-EP713 (Cryphonectria hypovirus 1), AAA67458; LRV2-1 (Leishmania RNA virus 1-1), AAB50024; ScV-L-A (Saccharomyces cerevisiae virus L-A), AAA50321; DaV (Diaporthe ambigua RNA virus 1), AAF22958; FusoV (Mycovirus FusoV from Fusarium solani), BAA09520; HvV190S (Helminthosporium victoriae virus 190S), AAB94791; RsV (Rhizoctonia solani virus), AF133290; CHV1-713 (Cryphonectria hypovirus EP713), AAA67458; CHV2-NB58 (Cryphenectria hypovirus 2-NB58), AAA20137; CHV3-GH2 (Cryphonectria hypovirus 3-GH2), AF13604; and BaYMV (Barley yellow mosaic virus), CAA49412.