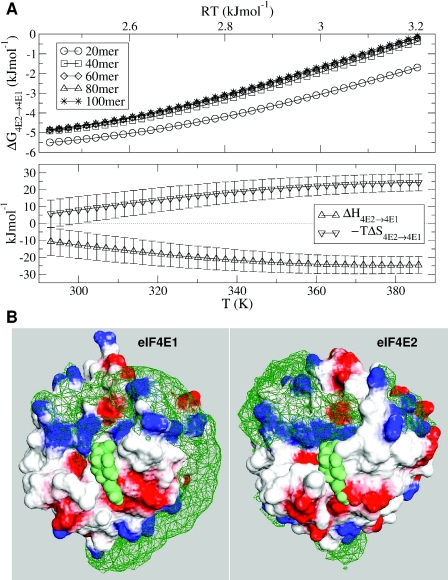
Figure 3.
Binding of mRNA to S.pombe eIF4E1/E2. (A) Temperature-dependence of binding differential, with ΔG4E2→4E1 plotted for variable polymer length. Simulations have been performed up to high (non-physiological) temperatures to facilitate the parallel tempering algorithm in allowing configurations to escape metastable ‘traps’. More favourable binding to S.pombe eIF4E1 is denoted by negative ΔG. (B) eIF4E surfaces with mRNA monomer density plots, as detailed in Figure 2, with spacefilled cap analogue (green).