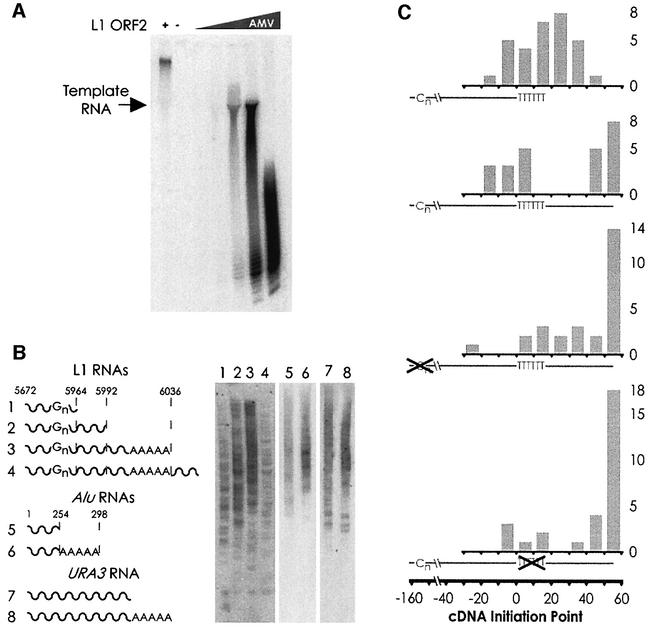
Fig. 4. (A) In a homopolymer RT assay (polyA RNA, oligo-dT primer), L1 ORF2p produces RT products far in excess of the molecular weight of the template, indicative of template switching activity; AMV RT does not. (B) TPRT of various RNA species. The end-point of the L1 and Alu RNAs are indicated with the dotted line (not to scale). RNA number 4 has 38 nt of vector RNA after the polyA tail. URA3, fragment of S.cerevisiae URA3 RNA with and without a polyA tail. (C) Distribution of cDNA initiation points. Shown below each histogram is a full-length cDNA (from 3′ to 5′), with the positions of the reverse-transcribed polyG and polyA tracts indicated by ‘Cn’ and ‘TTTTTT’, respectively. Position zero marks the end of L1 sequence and the beginning of the polyA tail. The actual positions of the cDNA initiation are plotted in the histogram grouped into 10 nt bins. For the first cDNA (generated from RNA number 1 in B), many cDNAs end beyond the designed position at nucleotide 14 due to the addition of extra A residues to the RNA by T7 polymerase (see Materials and methods). Transposition of RNA number 4 (the second cDNA from the top) yields two distinct populations of cDNA initiation points. Mutation of either the polyG or the polyA sequence (histograms three and four, respectively) removes the bias towards internal initiation of cDNA synthesis, although only the polyA mutation results in a statistically significant difference. Student’s two-tailed t-test comparison of wild type versus mutant, bins –20–30 with bins 30–60; polyG p-value = 0.17; polyA p-value = 0.05. For all three cDNAs from 3′ extended transcripts, all initiation points in the 50–60 bin occurred at nucleotide 53, the end of the transcript. The following number of highly truncated (endpoint <–30) cDNAs were excluded from the statistical analysis: wt 3′ extended RNA, 3; polyG mutant, 4; polyA mutant, 3.

An official website of the United States government
Here's how you know
Official websites use .gov
A
.gov website belongs to an official
government organization in the United States.
Secure .gov websites use HTTPS
A lock (
) or https:// means you've safely
connected to the .gov website. Share sensitive
information only on official, secure websites.