Abstract
Proteins of the GFP (green fluorescent protein) family demonstrate a great spectral and phylogenetic diversity. However, there is still an intense demand for red-shifted GFP-like proteins in both basic and applied science. To obtain GFP-like chromoproteins with red-shifted absorption, we performed a broad search in blue-coloured Anthozoa species. We revealed specimens of Actinia equina (beadlet anemone) exhibiting a bright blue circle band at the edge of the basal disc. A novel blue chromoprotein, aeCP597, with an absorption maximum at 597 nm determining the coloration of the anemone basal disk was cloned. AeCP597 carries a chromophore chemically identical with that of the well-studied DsRed (red fluorescent protein from Discosoma sp.). Thus a strong 42-nm bathochromic shift of aeCP597 absorption compared with DsRed is determined by peculiarities of chromophore environment. Site-directed and random mutagenesis of aeCP597 resulted in far-red fluorescent mutants with emission maxima at up to 663 nm. The most bright and stable mutant AQ143 possessed excitation and emission maxima at 595 and 655 nm respectively. Thus aeCP597 and its fluorescent mutants set a new record of red-shifted absorption and emission maxima among GFP-like proteins.
Keywords: beadlet anemone (Actinia equina), blue chromoprotein, coloration, fluorescent labelling, green fluorescent protein (GFP), red fluorescent protein from Discosoma sp. (DsRed)
Abbreviations: aeCP597, blue chromoprotein from Actinia equina; asFP595, purple chromoprotein from Anemonia sulcata; DsRed, red fluorescent protein from Discosoma sp.; GFP, green fluorescent protein
INTRODUCTION
GFP (green fluorescent protein) from the hydroid jellyfish Aequorea victoria, its mutants and homologues are used intensely in biotechnology. They represent the only available fluorescent tag fully encoded in a single gene. GFP-like proteins are utilized for in vivo labelling of organisms, cells, organelles and proteins, as well as in molecular sensors that probe different aspects of intracellular environment [1]. Recent studies demonstrated broad spectral and phylogenic diversity of GFP-like proteins [2,3]. In addition to the well-known green fluorescent proteins, several novel spectral variants such as naturally yellow and red fluorescent proteins and non-fluorescent chromoproteins were characterized [4,5]. Mutagenesis efforts resulted in far-red fluorescent proteins emitting at close to 650 nm [6,7].
Structural studies revealed unexpected chromophore diversity in red-shifted GFP-like proteins. Already, four chemically distinct types of red-shifted chromophores are known: DsRed (red fluorescent protein from Discosoma sp.)-like [8], Kaede (green fluorescent protein from Trachyphyllia geoffroyi)-like [9], zFP538 (yellow fluorescent protein from Zoanthus sp.)-like [10] and asFP595 (purple chromoprotein from Anemonia sulcata)-like [11]. The DsRed-like chemical structure appeared to be the most common chromophore in red-shifted GFP-like proteins. Crystallographic studies revealed three possible conformations (planar cis [12,13], planar trans [14] and non-planar trans [15]) of DsRed-like chromophore within proteins with distinct spectral properties. In particular, the non-planar trans conformation is characteristic of non-fluorescent chromoprotein Rtms5 (from Montipora efflorescens) [15] and probably of many other chromoproteins [3,16].
Red-shifted fluorescent proteins attract particular attention from both scientific and practical points of view. In practice, red and far-red fluorescent proteins are important for multi-coloured labelling and applications based on FRET (fluorescence resonance energy transfer). Far-red fluorescence is in especially high demand for whole-mount labelling since long-wave light penetrates the tissues more easily.
The present paper describes a novel blue GFP-like chromo-protein and its mutagenesis resulting in the most red-shifted fluorescent protein to date.
EXPERIMENTAL
Gene cloning and mutagenesis
Total RNA was isolated from a small blue-coloured piece of a single specimen of Actinia equina (beadlet anemone) with NucleoSpin RNA II kit (Clontech). Synthesis and amplification of cDNA was performed by SMART PCR cDNA synthesis kit (Clontech). A fragment of cDNA encoding a novel GFP homologue was obtained by PCR with degenerate primers using a strategy described previously [4]. A step-out RACE (rapid amplification of cDNA ends) PCR method [17] was used to clone the 5′-end region of the target cDNA. The cDNA nucleotide sequence encoding the novel GFP-like protein aeCP597 from A. equina has been deposited in GenBank® database under the accession number DQ159069. For bacterial expression of aeCP597, the full-length coding region was amplified using specific primers and was cloned into pQE30 vector (Qiagen).
Site-directed mutagenesis was performed by PCR using the overlap extension method, with primers containing appropriate target substitutions [18]. For random mutagenesis, we used Diversity PCR random mutagenesis kit (Clontech) under conditions optimal for five or six mutations per 1000 bp. Escherichia coli colonies expressing mutant proteins were visually screened in a fluorescent stereomicroscope SZX-12 (Olympus) equipped with spectrophotometer SMS 2 VIS (Pannhoff Optische Messtechnik) that was used to record emission spectra for individual colonies. The brightest and the most red-shifted variants were selected and subjected to the next rounds of random mutagenesis.
Protein expression and spectral measurements
N-terminal His6-tagged proteins were expressed in E. coli XL1 Blue strain (Invitrogen) and purified using TALON metal affinity resin (Clontech). Absorption and excitation–emission spectra were recorded with Beckman DU520 UV/VIS spectrophotometer and Varian Cary Eclipse fluorescence spectrophotometer respectively. A fluorescent stereomicroscope SZX-12 (Olympus) equipped with spectrophotometer SMS 2 VIS was used to measure emission spectra for individual E. coli colonies as well as absorption spectra for coloured body parts of A. equina specimens.
Heated and unheated protein samples in standard reducing sample buffer were subjected to common SDS/PAGE (15% gel) at 15 mA/gel.
Expression in HeLa cell line
AQ143 mutant was cloned into pEGFP-C1 vector (Clontech) between AgeI and BglII restriction sites in lieu of the EGFP (enhanced GFP)-coding region. HeLa cells were transfected using Lipofectamine™ reagent (Invitrogen) and were probed 20 h post-transfection. Leica confocal inverted microscope TCS SP2 was used for cell imaging. The red fluorescent signal was acquired using a 543 nm excitation laser line and detection at 570–700 nm. Fluorescence spectrum for individual cells was measured in x-y-λ mode by collecting 50 image stacks in 10-nm increments covering the range 560–732 nm.
RESULTS AND DISCUSSION
Cloning cDNA encoding aeCP597
All wild-type GFP-like chromoproteins known to date possess absorption peaks at less than 580 nm, which corresponds to a crimson–purple–lilac appearance to the human eye. To clone a red-shifted chromoprotein, we decided to find specimens with truly blue coloured parts that should correspond to absorption maxima at 600 nm or more. Many Anthozoa specimens were visually inspected in Moscow sea aquaria as well as in natural habitats on Osprey Reef in the Coral Sea, off Australia. Unfortunately, the overwhelming majority of specimens with non-fluorescent coloration were not of the colour desired, although they often looked dark blue in natural habitats owing to ineffective red radiation penetration in sea water. Finally, we spotted a species possessing dark blue coloration, in Moscow Zoo aquarium. It appeared to be A. equina originally collected in the Black Sea.
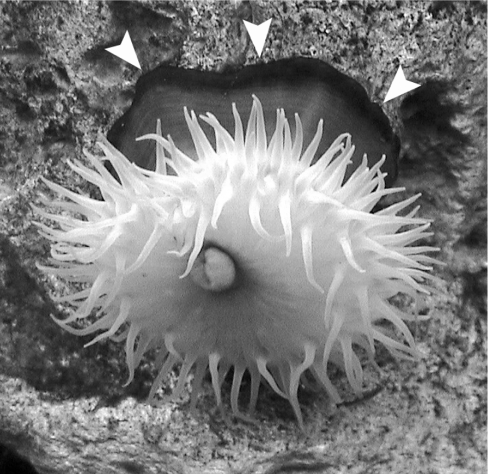
Generally, beadlet anemones demonstrate a broad variety of body coloration [19]. Our specimen of A. equina possessed a blue circular stripe at the edge of the basal disc (Figure 1). Such a pattern is rather unusual for GFP-like protein localization in sea anemones; expression of fluorescent and chromoproteins at the tentacles and the oral disc is much more common. We dissected a small piece of coloured tissue and used it for RNA isolation. Cloning of target cDNA was performed by PCR with degenerated primers corresponding to conservative amino acid regions as described in [4]. As a result, the cDNA sequence encoding the new GFP homologue was obtained. We named the new chromo-protein aeCP597 in accordance with nomenclature suggested by Wiedenmann et al. [20] for GFP-like chromoproteins (Actinia equina chromoprotein with absorption maximum at 597 nm; see below). AeCP597 is highly homologous with a purple chromo-protein asFP595 [5] from sea anemone Anemonia sulcata (68% amino acid identity). As in asFP595, the chromophore-forming triad in aeCP597 is Met63-Tyr64-Gly65 (Figure 2). In the asCP597 sequence, one can find all the typical features of sea anemone chromoproteins: a 3-amino-acid shorter N-terminal part, cysteine residues just in front of chromophore-forming amino acids, and, most importantly, Cys143 and Ser158 (positions 148 and 165 in accordance with GFP numbering), which are rather common in chromoproteins, but are never present in fluorescent proteins [5,14,15].
Figure 1. A. equina specimen.
The arrowheads point to the blue stripe on the basal disc that appears black in this black-and-white image.
Figure 2. Amino acid sequence alignment of the new chromoprotein aeCP597 and a number of GFP homologues.
Sequence numbering corresponds to aeCP597 above the alignments and GFP beneath the alignments. Introduced gaps are represented by dashes. The residues with side chains forming the interior of the β-can are shaded in grey. Chromophore-forming residues are indicated by the dashed box. Mutated positions in aeCP597 sequence are boxed, and corresponding substitutions are shown above the boxes. In asFP595 and hcCP (Heteractis crispa non-fluorescent chromoprotein) sequences, amino acids that were mutated to produce red fluorescent mutants AsRed2 and HcRed1 (Clontech) are underlined. eqFP611, far-red fluorescent protein from the sea anemone Entacmaea quadricolor.
Spectral properties of aeCP597
For heterologous expression of aeCP597, the full-length coding region was cloned into pQE30 vector. E. coli colonies expressing aeCP597 possessed bright blue coloration, but produced no detectable fluorescence. The absorbance spectrum for purified aeCP597 peaked at 597 nm (Figure 3A). Thus aeCP597 is the GFP-like protein with the farthest red-shifted absorption described to date. The absorbance spectrum measured directly from the blue stripe on A. equina's body overlaid convincingly with the aeCP597 spectrum (Figure 3A). We concluded that the blue coloration of the A. equina basal disc edge is determined by aeCP597.
Figure 3. Spectral properties of aeCP597 and its fluorescent mutants.
(A) Absorbance spectra for wild-type aeCP597 (solid line) and for the blue-coloured basal disc of the A. equina specimen (dotted line). (B) Excitation (dotted line) and emission (solid line) spectra for the AQ14 mutant. (C) Time-course of AQ14 absorbance spectrum change. Curve 1, freshly purified protein sample; curves 2–4, the same protein sample after 21, 62 and 183 h at 30 °C. (D) Excitation (dotted line) and emission (solid line) spectra for the AQ143 mutant.
Generally, spectral properties of GFP-like proteins are determined by both chemical chromophore structure and the protein shell of the chromophore. To investigate the structural basis for red-shifted absorption in aeCP597, we performed several simple tests. First, to check for possible protein fragmentation, purified chromoprotein was subjected to denaturing gel electrophoresis. The unheated protein sample produced a high-molecular-mass (>100 kDa) coloured band, suggesting a tetrameric structure of the native aeCP597. In the heated sample, electrophoresis yielded a major band at 27 kDa, corresponding to full-length aeCP597 with a His6-tag, and two minor bands at 8 and 19 kDa, corresponding to the breaking polypetide (Figure 4). The pattern resembled that described for DsRed. Appearance of the lower bands is explained by partial hydrolysis of the acylimine bond in DsRed chromophore during protein heating before electrophoresis [8]. In contrast, all other known chemically distinct red-shifted chromophores in GFP-like proteins imply a protein backbone break in the native state [9–11], so that the corresponding 8 and 19 kDa bands dominate in gel electrophoresis. Secondly, to investigate the spectral features of the naked chromophore, absorption spectra for acid- and alkali-denatured aeCP597 were obtained. In alkali, the absorption maximum was at 450 nm, while, in mild acid, a fast transition from the 450-nm to the 380-nm form was detected (results not shown). Again, this conversion is very similar to that of proteins with DsRed-like chromophores [3] and different from other red-shifted proteins. These data suggest that aeCP597 carries a DsRed-like chromophore. Thus the strong 42-nm red-shift compared with DsRed absorption maximum is determined by aeCP597 protein shell features.
Figure 4. Protein gel electrophoresis analysis of aeCP597 and its mutant AQ14 (Coomassie Blue staining).
Lanes 1 and 2, heated and unheated samples of recombinant purified aeCP597. Lines 3 and 4, heated and unheated samples of AQ14 after the ‘red-into-green’ spontaneous conversion. Positions of molecular-mass markers are shown on the left (sizes in kDa). The asterisk marks bands that are coloured/fluorescent before staining and correspond to native tetrameric protein in the unheated samples. The black triangle shows denatured full-length protein bands. The white triangles indicate positions of minor bands corresponding to 19 and 8 kDa fragments of broken polypeptide chain.
Generally, GFP-like chromoproteins possess more red-shifted absorption than red fluorescent proteins. Recent crystallographic studies revealed a trans non-planar conformation of chromophore to be characteristic for the chromoproteins [11,15]. One can propose that such a chromophore conformation forced by the protein environment is at least partially responsible for red-shifted spectra. Other possible causes for the aeCP597 strong bathochromic shift might be Met63 {corresponding Met66 is important for the red-shift in DsRed mutants [7,21]; also, the most red-shifted wild-type fluorescent protein eqFP611 (far-red fluorescent protein from the sea anemone Entacmaea quadricolor) [22] carries the corresponding Met63} and His197 (aromatic residues at corresponding position 203 are key substitution sites in yellow mutants of GFP [23]).
Direct evolution of aeCP597 into far-red fluorescent state
We demonstrated previously that GFP-like chromoproteins can be transformed into red and far-red fluorescent state by means of mutagenesis [5,6,24]. The most important positions for such conversion were found to be 148 and 165 in accordance to GFP numbering. To make aeCP597 fluorescent, we randomized residues at the corresponding positions 143 and 158. Approx. 5000 clones expressing mutant proteins (theoretical diversity was 400 variants) were visually screened using a fluorescent stereomicroscope. Several of the brightest red fluorescent clones were selected. As a result, two amino acid combinations were found to ensure a fluorescent state with an emission maximum at approx. 645 nm: Ser143/Ala158 and Ser143/Cys158. In addition, aggregation-reducing substitutions K6T and K7E [25] were introduced. Resulting mutants were subjected to random mutagenesis at the full coding region. Selection was directed at both enhanced brightness and red-shifted fluorescence. In order to monitor absorption spectra, we used a fluorescent stereomicroscope equipped with spectrofluorimeter allowing us to measure emission spectrum from individual colonies. As a result of two rounds of random mutagenesis, we selected a protein named AQ14 (mutations K6T, K7E, Y101S, C143S, M146I, S158A compared with the wild-type protein) demonstrating an excitation maximum at 595 nm and emission maximum at 663 nm (Figure 3B). This is the farthest red-shifted fluorescence described to date for GFP-like proteins.
Unfortunately, AQ14 fluorescence was highly sensitive to light irradiation. Moreover, even in the dark at room and higher temperatures, AQ14 red fluorescence decreased dramatically over several days of storage. We investigated this process in detail and discovered a new type of ‘red-into-green’ spectral transition never described for red fluorescent proteins. Monitoring of the absorbance spectrum in purified mature AQ14 showed that the absorption peak at 588 nm gradually transformed into a peak at 510 nm (Figure 3C). In addition, subtraction of the spectra clearly demonstrated a decrease of the 450 nm peak and an increase of the peak at 400 nm. It is well known that protonation of phenolic oxygen of the chromophore-forming tyrosine residue greatly affects the absorption peak position in GFP-like proteins. In GFP, for instance, anionic (deprotonated) and neutral (protonated) chromophores absorb at 475 and 395 nm respectively [26–28]. Analogously, AQ14 absorption peaks at 588 and 450 nm can probably be attributed to anionic and neutral states of ‘red’ chromophore that transform into anionic and neutral ‘green’ chromophore states absorbing at 510 and 400 nm respectively. Interestingly, very little green fluorescence was observed for AQ14 samples after red-into-green conversion. Thus the GFP-like chromophore within this protein exists in a ‘chromo’ state ensuring effective light absorption, but a low fluorescence quantum yield.
The following possible chemical mechanism of AQ14 red-intogreen conversion can be suggested. Compared with GFP-like chromophore, DsRed-like chromophore carries an additional acylimine moiety [8]. Thus red-into-green transition should result in the disruption of acylimine structure. For example, the acylimine C=N bond can undergo hydrolysis that results in protein backbone break [8]. However, denaturing gel electrophoresis of AQ14 demonstrated that the protein chain remained unbroken after the conversion (Figure 4). Taking into account this observation, we propose that the blue shift in AQ14 results from the hydration of the C=N bond of the chromophore acylimine moiety by an inner water molecule without subsequent protein backbone break.
To overcome AQ14 instability, we applied additional rounds of random mutagenesis. To select light-insensitive fluorescent proteins, screening of the E. coli libraries was performed after 2-day plate storage under direct sunlight. As a result, a mutant AQ143 (mutations S3P, K6T, K7E, V19I, Y101S, C143S, M146I, S158A and N168D compared with the wild-type protein) was selected. AQ143 demonstrated excitation and emission peaks at 595 and 655 nm respectively (Figure 3D), and no longer underwent the spontaneous degradation into the non-fluorescent form. A rational explanation of mutagenesis-introduced mutations (i.e. S3P, V19I, Y101S, M146I and N168D) on protein brightness and thermo- and photo-stability is still lacking. Amino acids at positions 101 and 146 probably participate in hydrophobic and hydrophilic tetramer interfaces respectively [2,12,13]. So, AQ14-characteristic mutations at these positions may adjust the tetramer structure after site-directed mutagenesis of aeCP597 inner positions 143 and 158. Among the random mutations, only one, V19I, represents an inner residue (Figure 2). Ile19 may ensure a more stable hydrophobic core and thus enhanced stability of AQ143. Finally, mutations S3P and N168D cluster together with corresponding mutations introduced randomly near the N-terminal end and in the loop between the eighth and ninth β-strands in asFP595 and hcCP (Heteractis crispa non-fluorescent chromoprotein) chromo-proteins in the course of their transformation to red fluorescent proteins AsRed2 (a variant of Anemonia sulcata red fluorescent protein) and HcRed1 (a red fluorescent mutant of Heteractis crispa non-fluorescent chromoprotein) (Clontech) respectively (Figure 2).
The brightness of AQ143 is not high (quantum yield 0.04 and molar absorption coefficient 90000 M−1·cm−1) similar to all other far-red fluorescent mutants [6,7]. However, AQ143 far-red fluorescence can be detected easily in cells (Figure 5). Thus it provides additional colour for multicoloured labelling of cells and organelles.
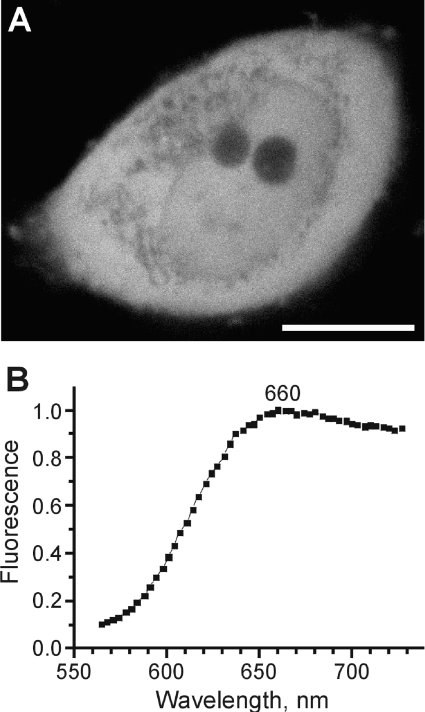
Figure 5. Confocal image of a HeLa cell expressing AQ143 (A) and emission spectrum of this cell measured using a 543-nm excitation laser line (B).
Scale bar in (A), 10 μm.
Acknowledgments
We thank Dr Anya Satin for organizing the expedition to Osprey Reef, Tatyana Chepurnykh for cell culture work and Maria Bulina for help in manuscript preparation. This work was supported by European Commission FP-6 Integrated Project LSHG-CT-2003-503259, Russian Academy of Sciences for the program ‘Molecular and Cell Biology’, Russian Foundation for Basic Research (grant 05-04-49316 to Y.A.L.), and the grant of the President of the Russian Federation MK-3835.2004.4 (to N.G.G.).
References
- 1.Lippincott-Schwartz J., Patterson G. H. Development and use of fluorescent protein markers in living cells. Science. 2003;300:87–91. doi: 10.1126/science.1082520. [DOI] [PubMed] [Google Scholar]
- 2.Verkhusha V. V., Lukyanov K. A. The molecular properties and applications of Anthozoa fluorescent proteins and chromoproteins. Nat. Biotechnol. 2004;22:289–296. doi: 10.1038/nbt943. [DOI] [PubMed] [Google Scholar]
- 3.Shagin D. A., Barsova E. V., Yanushevich Y. G., Fradkov A. F., Lukyanov K. A., Labas Y. A., Semenova T. N., Ugalde J. A., Meyers A., Nunez J. M., et al. GFP-like proteins as ubiquitous metazoan superfamily: evolution of functional features and structural complexity. Mol. Biol. Evol. 2004;21:841–850. doi: 10.1093/molbev/msh079. [DOI] [PubMed] [Google Scholar]
- 4.Matz M. V., Fradkov A. F., Labas Y. A., Savitsky A. P., Zaraisky A. G., Markelov M. L., Lukyanov S. A. Fluorescent proteins from nonbioluminescent Anthozoa species. Nat. Biotechnol. 1999;17:969–973. doi: 10.1038/13657. [DOI] [PubMed] [Google Scholar]
- 5.Lukyanov K. A., Fradkov A. F., Gurskaya N. G., Matz M. V., Labas Y. A., Savitsky A. P., Markelov M. L., Zaraisky A. G., Zhao X., Fang Y., et al. Natural animal coloration can be determined by a non-fluorescent green fluorescent protein homolog. J. Biol. Chem. 2000;275:25879–25882. doi: 10.1074/jbc.C000338200. [DOI] [PubMed] [Google Scholar]
- 6.Gurskaya N. G., Fradkov A. F., Terskikh A., Matz M. V., Labas Y. A., Martynov V. I., Yanushevich Y. G., Lukyanov K. A., Lukyanov S. A. GFP-like chromoproteins as a source of far-red fluorescent proteins. FEBS Lett. 2001;507:16–20. doi: 10.1016/s0014-5793(01)02930-1. [DOI] [PubMed] [Google Scholar]
- 7.Wang L., Jackson W. C., Steinbach P. A., Tsien R. Y. Evolution of new nonantibody proteins via iterative somatic hypermutation. Proc. Natl. Acad. Sci. U.S.A. 2004;101:16745–16749. doi: 10.1073/pnas.0407752101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Gross L. A., Baird G. S., Hoffman R. C., Baldridge K. K., Tsien R. Y. The structure of the chromophore within DsRed, a red fluorescent protein from coral. Proc. Natl. Acad. Sci. U.S.A. 2000;97:11990–11995. doi: 10.1073/pnas.97.22.11990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Mizuno H., Mal T. K., Tong K. I., Ando R., Furuta T., Ikura M., Miyawaki A. Photo-induced peptide cleavage in the green-to-red conversion of a fluorescent protein. Mol. Cell. 2003;12:1051–1058. doi: 10.1016/s1097-2765(03)00393-9. [DOI] [PubMed] [Google Scholar]
- 10.Remington S. J., Wachter R. M., Yarbrough D. K., Branchaud B., Anderson D. C., Kallio K., Lukyanov K. A. zFP538, a yellow-fluorescent protein from Zoanthus, contains a novel three-ring chromophore. Biochemistry. 2005;44:202–212. doi: 10.1021/bi048383r. [DOI] [PubMed] [Google Scholar]
- 11.Quillin M. L., Anstrom D. M., Shu X., O'Leary S., Kallio K., Chudakov D. M., Remington S. J. Kindling fluorescent protein from Anemonia sulcata: dark-state structure at 1.38 Å resolution. Biochemistry. 2005;44:5774–5787. doi: 10.1021/bi047644u. [DOI] [PubMed] [Google Scholar]
- 12.Wall M. A., Socolich M., Ranganathan R. The structural basis for red fluorescence in the tetrameric GFP homolog DsRed. Nat. Struct. Biol. 2000;7:1133–1138. doi: 10.1038/81992. [DOI] [PubMed] [Google Scholar]
- 13.Yarbrough D., Wachter R. M., Kallio K., Matz M. V., Remington S. J. Refined crystal structure of DsRed, a red fluorescent protein from coral, at 2.0-Å resolution. Proc. Natl. Acad. Sci. U.S.A. 2001;98:462–467. doi: 10.1073/pnas.98.2.462. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Petersen J., Wilmann P. G., Beddoe T., Oakley A. J., Devenish R. J., Prescott M., Rossjohn J. The 2.0-Å crystal structure of eqFP611, a far red fluorescent protein from the sea anemone Entacmaea quadricolor. J. Biol. Chem. 2003;278:44626–44631. doi: 10.1074/jbc.M307896200. [DOI] [PubMed] [Google Scholar]
- 15.Prescott M., Ling M., Beddoe T., Oakley A. J., Dove S., Hoegh-Guldberg O., Devenish R. J., Rossjohn J. The 2.2 Å crystal structure of a pocilloporin pigment reveals a nonplanar chromophore conformation. Structure. 2003;11:275–284. doi: 10.1016/s0969-2126(03)00028-5. [DOI] [PubMed] [Google Scholar]
- 16.Martynov V. I., Maksimov B. I., Martynova N. Y., Pakhomov A. A., Gurskaya N. G., Lukyanov S. A. A purple-blue chromoprotein from Goniopora tenuidens belongs to the DsRed subfamily of GFP-like proteins. J. Biol. Chem. 2003;278:46288–46292. doi: 10.1074/jbc.M306810200. [DOI] [PubMed] [Google Scholar]
- 17.Matz M., Shagin D., Bogdanova E., Britanova O., Lukyanov S., Diatchenko L., Chenchik A. Amplification of cDNA ends based on template-switching effect and step-out PCR. Nucleic Acids Res. 1999;27:1558–1560. doi: 10.1093/nar/27.6.1558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Ho S. N., Hunt H. D., Horton R. M., Pullen J. K., Pease L. R. Site-directed mutagenesis by overlap extension using the polymerase chain reaction. Gene. 1989;77:51–59. doi: 10.1016/0378-1119(89)90358-2. [DOI] [PubMed] [Google Scholar]
- 19.Haylor G. S., Thorpe J. P., Carter M. A. Genetic and ecological differentiation between sympatric colour morphs of the common intertidal sea anemone Actinia equina. Mar. Ecol. Prog. Ser. 1984;16:281–289. [Google Scholar]
- 20.Wiedenmann J., Elke C., Spindler K. D., Funke W. Cracks in the β-can: fluorescent proteins from Anemonia sulcata (Anthozoa, Actinaria) Proc. Natl. Acad. Sci. U.S.A. 2000;97:14091–14096. doi: 10.1073/pnas.97.26.14091. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Shaner N. C., Campbell R. E., Steinbach P. A., Giepmans B. N., Palmer A. E., Tsien R. Y. Improved monomeric red, orange and yellow fluorescent proteins derived from Discosoma sp. red fluorescent protein. Nat. Biotechnol. 2004;22:1567–1572. doi: 10.1038/nbt1037. [DOI] [PubMed] [Google Scholar]
- 22.Wiedenmann J., Schenk A., Rocker C., Girod A., Spindler K. D., Nienhaus G. U. A far-red fluorescent protein with fast maturation and reduced oligomerization tendency from Entacmaea quadricolor (Anthozoa, Actinaria) Proc. Natl. Acad. Sci. U.S.A. 2002;99:11646–11651. doi: 10.1073/pnas.182157199. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Ormo M., Cubitt A. B., Kallio K., Gross L. A., Tsien R. Y., Remington S. J. Crystal structure of the Aequorea victoria green fluorescent protein. Science. 1996;273:1392–1395. doi: 10.1126/science.273.5280.1392. [DOI] [PubMed] [Google Scholar]
- 24.Bulina M. E., Chudakov D. M., Mudrik N. N., Lukyanov K. A. Interconversion of Anthozoa GFP-like fluorescent and non-fluorescent proteins by mutagenesis. BMC Biochem. 2002;3:7. doi: 10.1186/1471-2091-3-7. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Yanushevich Y. G., Staroverov D. B., Savitsky A. P., Fradkov A. F., Gurskaya N. G., Bulina M. E., Lukyanov K. A., Lukyanov S. A. A strategy for the generation of non-aggregating mutants of Anthozoa fluorescent proteins. FEBS Lett. 2002;511:11–14. doi: 10.1016/s0014-5793(01)03263-x. [DOI] [PubMed] [Google Scholar]
- 26.Heim R., Prasher D. C., Tsien R. Y. Wavelength mutations and posttranslational autoxidation of green fluorescent protein. Proc. Natl. Acad. Sci. U.S.A. 1994;91:12501–12504. doi: 10.1073/pnas.91.26.12501. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Niwa H., Inouye S., Hirano T., Matsuno T., Kojima S., Kubota M., Ohashi M., Tsuji F. I. Chemical nature of the light emitter of the Aequorea green fluorescent protein. Proc. Natl. Acad. Sci. U.S.A. 1996;93:13617–13622. doi: 10.1073/pnas.93.24.13617. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Brejc K., Sixma T. K., Kitts P. A., Kain S. R., Tsien R. Y., Ormo M., Remington S. J. Structural basis for dual excitation and photoisomerization of the Aequorea victoria green fluorescent protein. Proc. Natl. Acad. Sci. U.S.A. 1997;94:2306–2311. doi: 10.1073/pnas.94.6.2306. [DOI] [PMC free article] [PubMed] [Google Scholar]