Abstract
Human retrovirus 5 (HRV-5) represented a fragment of a novel retrovirus sequence identified in human RNA and DNA preparations. In this study, the genome of HRV-5 was cloned and sequenced and integration sites were analyzed. Using PCR and Southern hybridization, we showed that HRV-5 is not integrated into human DNA. A survey of other species revealed that HRV-5 is present in the genomic DNA of the European rabbit (Oryctolagus cuniculus) and belongs to an endogenous retrovirus family found in rabbits. The presence of rabbit sequences flanking HRV-5 proviruses in human DNA extracts suggested that rabbit DNA was present in our human extracts, and this was confirmed by PCR analysis that revealed the presence of rabbit mitochondrial DNA sequences in four of five human DNA preparations tested. The origin of the rabbit DNA and HRV-5 in human DNA preparations remains unclear, but laboratory contamination cannot explain the preferential detection of HRV-5 in inflammatory diseases and lymphomas reported previously. This is the first description of a retrovirus genome in rabbits, and sequence analysis shows that it is related to but distinct from A-type retroelements of mice and other rodents. The species distribution of HRV-5 is restricted to rabbits; other species, including other members of the order Lagomorpha, do not contain this sequence. Analysis of HRV-5 expression by Northern hybridization and reverse transcriptase PCR indicates that the virus is transcribed at a low level in many rabbit tissues. In light of these findings we propose that the sequence previously designated HRV-5 should now be denoted RERV-H (for rabbit endogenous retrovirus H).
A 930-bp fragment of a new retrovirus genome was previously identified in human tissues and provisionally designated human retrovirus 5 (HRV-5) (15). HRV-5 has little sequence similarity with the known human retroviruses human T-cell lymphotropic virus types 1 and 2 (HTLV-1 and -2) (18, 29) and human immunodeficiency virus types 1 and 2 (HIV-1 and -2) (2, 9). Instead, HRV-5 is related to the simian D-type retroviruses (36), murine mammary tumor virus (26), and rodent intracisternal A-type particle (IAP) genes (20). Thus, based on sequence similarity, HRV-5 fits into the Betaretrovirus genus according to the present International Committee on Viral Taxonomy classification (16).
HRV-5 was first identified during an investigation into the potential role of retroviruses in Sjögren's syndrome (15). In that initial report, the cloning and sequencing of a short region of the pro and pol genes were described and this sequence was shown not to be endogenous in the human genome. This observation, together with other data such as a high degree of sequence variation between isolates from different individuals and conservation of open reading frames (ORFs) for gag, pro, and pol, led to the conclusion that HRV-5 was infectiously acquired and actively replicating (15). Since then, the preferential detection of HRV-5 in inflammatory diseases, particularly rheumatoid arthritis (RA) and systemic lupus erythematosus (SLE), has been reported (14). However, the virus has also been detected in other inflammatory conditions (6, 14, 31), in lymphoma (19, 27, 30), and occasionally in normal tissue and blood (6, 14). HRV-5 was initially identified in British patients, but viral sequences have since been independently detected in individuals from several European countries and the United States (19, 27; D. J. Griffiths and P. J. W. Venables, Author's Reply, Arthritis Rheum. 42:2493-2494, 1999), suggesting that this virus had a wide geographical prevalence in humans. Taken together, these data indicated that HRV-5 was a common infection presenting at a greater load in certain disease states.
Further evaluation of the role of HRV-5 in RA, SLE, and other inflammatory diseases required the cloning and sequencing of the viral genome in order to establish robust serological assays and other reagents for studying HRV-5 infection. As described here, we cloned the HRV-5 genome from clinical samples using PCR-based cloning techniques for DNA walking. Once the genome had been sequenced, we continued to use these methods to clone virus integration sites. However, analysis of sequences flanking HRV-5 proviruses revealed that this virus is not of human origin after all but is instead an endogenous retrovirus (ERV) of rabbits. ERVs are germ line elements with sequence similarity to known infectious retroviruses and are present in all vertebrate species that have been studied (4), although little has been published previously on retroviruses in rabbits. Therefore, this is the first report to describe a retroviral element at the molecular level in this species.
MATERIALS AND METHODS
Human DNA samples.
DNA was extracted from patient samples using standard proteinase K digestion and phenol extraction methods (32). Three DNA samples were used as the source of the HRV-5 integration site sequences. Sample A was extracted from whole blood of an apparently normal individual. Sample B was from resected colon tissue of a patient with ulcerative colitis. Sample C was extracted from peripheral blood of a patient with SLE.
Cell lines and rabbit tissue samples.
Cells were grown at 37°C in a humidified atmosphere containing 5% CO2 and were passaged two or three times weekly. IOWA (pig, Sus scrofa), CHO (Chinese hamster, Cricetulus griseus), BHK (Syrian hamster, Mesocricetus auratus), BALB3T12 (mouse, Mus musculus), Rat-2 (rat, Rattus norvegicus), PG-4 (cat, Felis catus), COS-SP6 (African green monkey, Cercopithecus aethiops), HeLa (human, Homo sapiens), QT 35 (Japanese quail, Coturnix coturnix japonica), and SIRC and EREp (both European rabbit [Oryctolagus cuniculus]) cell lines were grown in Dulbecco's modified Eagle medium (Gibco) containing 10% fetal calf serum (FCS; Biological Industries), 1% nonessential amino acids (NEA; Gibco), and antibiotics (100 U of penicillin and 100 μg of streptomycin per ml; Gibco). MLA 144 (gibbon ape, Hylobates lar), ChM114 (chimpanzee, Pan troglodytes), L-363 (human, H. sapiens), and SC300 (fat-tailed dunnart, Sminthopsis crassicaudata) cell lines were grown in RPMI 1640 (Gibco) containing 10% FCS, NEA, and antibiotics. DK (dog, Canis familiaris), Mv1-lu (mink, Mustela vison), NBL-6 (horse, Equus caballus), Tb1.lu (free-tailed bat, Tadarida brasiliensis), FRhK (rhesus macaque, Macaca mulatta), and MDBK (cow, Bos taurus) cell lines were grown in Eagle's minimum essential medium (Gibco) containing 10% FCS, NEA, and antibiotics. FCS was heat inactivated by incubation at 56°C for 1 h prior to use.
Samples of rabbit tissues (O. cuniculus, New Zealand White strain) were obtained from two rabbits and were supplied by the Institute of Cancer Research (Sutton, United Kingdom). Spleen tissue from a third rabbit was provided by J.-Y. Madec (CNEVA, Lyon, France). Spleen samples from European brown hare (Lepus europaeus) and Afghan pika (Ochotona rufescens) were obtained from the Zoological Society of London, (London, United Kingdom). Tissues from black-tailed jackrabbit (Lepus californicus) and pygmy marmoset (Callithirix pygmaea) were a kind gift from J. Martin (Imperial College, London, United Kingdom).
PCR-directed cloning of HRV-5 and integration sites.
Several PCR-based methods for DNA walking were used to clone the HRV-5 genome and flanking sequences from human DNA. Full details of the genome cloning procedure are presented in supplementary material available from http://www.ucl.ac.uk/windeyer-institute/Research/Griffiths.htm or directly from the authors on request. Three integration site fragments were cloned, each by a different PCR method. Clone 3A was obtained using a three-stage, nested Vectorette PCR (1) with ClaI-digested DNA from sample A using a kit from Genosys as recommended. First-stage primers were CACGTCACTGTAGATACATATTCAG and the first-round Vectorette primer that was supplied (Genosys) (40 cycles of 94°C, 1 min 10 s; 60°C, 1 min 10 s; and 72°C, 3 min with an initial denaturation at 94°C for 4 min). Second-stage primers were GGTGTAGTTATGGCCACAGCCATG and the second-stage Vectorette primer that was supplied (Genosys) (conditions same as for first stage). Third-stage primers were AACACTGCTTGCAGGCTTTTGCAG and the Vectorette sequencing primer that was supplied (35 cycles of 94°C, 1 min 10 s; 52°C, 1 min 10 s; and 72°C, 2 min with an initial denaturation at 94°C for 4 min). One microliter was transferred between the stages of the nested PCR. Following gel electrophoresis, an 1,100-bp band was purified, blunt end cloned into EcoRV-digested pBluescript II KS(−), and sequenced.
Clone 5A was obtained from sample B using an arbitrarily primed PCR technique (C. Cousens, University of Edinburgh, Edinburgh, United Kingdom, personal communication). Prior to PCR, 1 μg of DNA was added to a single primer extension reaction containing primer GGACCGGACGCTTGTTGTTCCCT with a 500 μM concentration of each deoxynucleoside triphosphate and the supplied buffer 3 and Pwo DNA polymerase from the Expand Long Template PCR system (Roche Molecular) and was incubated at 94°C, 1 min; 55°C, 1 min; and 68°C, 3 min for 60 cycles. One-tenth of the products of the extension reaction was added to a first-stage PCR containing HRV-5 primer LTR-R1 (TCTCTCCTCCTATATAGTCCTCTCC) and the arbitrary primer SV6 (GGAAAGTCCTTGGGGTCTTCTACC) and was cycled at low stringency (30 cycles of 94°C, 30 s; 30°C, 30 s; and 72°C, 3 min). One microliter of the products was reamplified in a second-stage reaction with primers LTR-R2 (CTGGAGCGGCTCTTGCAGCTTATC) and SV6 and was cycled at higher stringency (30 cycles of 94°C, 30 s; 60°C, 30 s; and 72°C, 2 min). One microliter of these products was then further amplified by adding to a third-stage reaction containing primers LTR-R3 (TCCTCTCCCAGCGCCACATTGTGG) and SV6 (25 cycles, as for second stage). The third-stage products were transferred to nylon membrane by Southern blotting, probed with a digoxigenin-labeled oligonucleotide (U3-R2, GTATAGTCCGAGTTGCTCCCCACAG) which was 3′ end labeled, and detected using digoxigenin system reagents (Roche Molecular) as recommended. A product of around 400 bp was detected, gel purified from the remaining third-stage products, and cloned into EcoRV-digested pBluescript II KS(−) (Stratagene) before sequencing.
Clone 5B was amplified from sample C using a capture PCR procedure (21). DNA was digested to completion with SspI and ligated to a double-stranded DNA linker (as described in reference 21). The ligated products were added to a single primer extension reaction containing the 5′-biotinylated primer GAG6R1 (GTAGCCAAAGAACTCCATTGTCTG) and reagents from the Expand Long Template PCR System (Roche Molecular Systems) and were cycled at 92°C, 1 min; 55°C, 1 min; and 68°C, 5 min for 40 cycles. The biotinylated extension products were purified on streptavidin-coated magnetic beads (KilobaseBINDER; Dynal, Oslo, Norway, used as recommended), and the clone 5B fragment was amplified by heminested PCR. The first-stage primers were LTR-R1 and Linker-1 (GCGGTGACCCGGGAGATCTGAATTC), and the second-stage primers were LTR-R3 and Linker-1. Conditions for both stages were as for the primer extension reaction. The products were blotted to nylon and were probed with digoxigenin-labeled LTR-R4 (GATGCATAGAATAAGTCTTAATTCCAG) to reveal a band of around 680 bp, which was then cloned and sequenced.
Southern analysis.
High-molecular-weight DNA was extracted from cultured cells or tissue samples using the Genomic DNA Extraction kit Genomic Tip (Qiagen) or standard phenol-choroform extractions (32). DNA was digested to completion with HindIII (Promega), and digested DNA (9 μg) was electrophoresed on a 0.8% agarose gel and transferred to a nylon membrane (Genescreen Plus; New England Nuclear) by the alkaline transfer procedure as recommended by the manufacturer. [32P]dCTP-radiolabeled probes were prepared with a kit (Ready-to-Go DNA labeling beads; Amersham Pharmacia) as recommended by the manufacturer. For genomic Southern blots, the probes used were as follows: a 357-bp fragment of HRV-5 gag (matrix [MA] domain, nucleotides 797 to 1148 of the reference clone, GenBank entry AF480924), a 268-bp fragment of the clone 3A flanking sequence (nucleotides 1174 to 1441 of GenBank entry AF480927), a 277-bp fragment of the clone 5A flanking sequence (nucleotides 111 to 387 of GenBank entry AF480928), and a 257-bp fragment of the clone 5B flanking sequence (nucleotides 329 to 585 of GenBank entry AF480929). Hybridizations were performed at 65°C for 16 h in 1 M sodium chloride, 10% dextran sulfate, 100 μg of salmon sperm DNA/ml, and 0.1% sodium dodecyl sulfate (SDS). Wash conditions for these probes were two 15-min washes at 65°C in 0.1% SDS-0.2× SSC (1× SSC is 0.15 M NaCl plus 0.015 M sodium citrate) and three 30-min washes in 0.1% SDS-0.2× SSC at room temperature. A 300-bp rabbit glyceraldehyde-3-phosphate dehydrogenase (GAPDH) probe was PCR amplified from rabbit liver RNA with primers GAPDH-F (GATGGTGAAGGTCGGAGTGAACG) and GAPDH-R (AAGACGCCAGTGGATTCCACCAC) (30 cycles of 94°C, 45 s; 55°C, 45 s; and 72°C, 45 s) and was hybridized at lower stringency (two 15-min washes in 0.1% SDS-2× SSC at 50°C and three 30-min washes in 0.1% SDS-2× SSC at room temperature) in order to detect homologs in all species.
PCR analysis on genomic DNA.
DNA for PCR was extracted from cell lines by lysis of 5 × 105 to 1 × 106 cells (trypsinized where appropriate and prewashed once in phosphate-buffered saline) and suspended in 250 μl of rapid lysis buffer (10 mM Tris-Cl, pH 8.0; 50 mM potassium chloride; 0.05% NP-40; and 0.05% Tween 20). Proteinase K (25 μg; Qiagen) was added, and the samples were incubated at 56°C for 24 h. Samples were then heated to 80°C for 20 min to inactivate the proteinase and were centrifuged for 15 min at 13,000 × g to pellet cellular debris. Ten-microliter aliquots of the supernatants were used in each PCR. DNA from tissues was extracted using the Qiagen DNA mini kit as recommended. One hundred nanograms of tissue DNA was added to each PCR.
PCRs were performed in a 50-μl reaction with Taq polymerase (Promega) in the supplied buffer with 2 mM MgCl2, a 200 μM concentration of each deoxynucleoside triphosphate, and 10 pmol of each primer. PCR primers and conditions were as follows: protease/reverse transcriptase (PR/RT), pol-F (TCAGGTGCTTCATTGGCAGGATCA) and pol-R (TAAAATTTGTACTTTTGGGCACTGCTG) (40 cycles of 94°C, 45 s; 63°C, 45 s; and 72°C, 45 s with an initial denaturation for 4 min at 94°C); long terminal repeat (LTR), U3-F (CTGTGGGGAGCAACTCGGACTATAC) and U5-R (CTTGCTGCTCCTCCGCACGCGG) (same conditions as for PR/RT but annealed at 55°C); and integrase (IN), IN-F (GCCATCCATTTGTTAACTTCTGCAC) and IN-R (GGTAAATTAGAATGAGCTCTTATGTG) (same conditions as for PR/RT but annealed at 59°C). To confirm their identity as HRV-5, the PCR products obtained were analyzed by Southern hybridization with [32P]dCTP-labeled internal probes that were, for pol, nucleotides 3620 to 3746; for LTR, nucleotides 6215 to 6407; and, for PR/RT, nucleotides 4995 to 4756 of the reference clone (GenBank accession no. AF480924). The suitability of the DNA for PCR was confirmed by PCR using a degenerate primer set for gammaretroviral pol sequences (present in most mammalian genomes) (ACCAAIGAYTAYMGRCCWGTICARGA and TTGAAMCCYTGKGGIARYCKIGTCCA, where I is inosine, Y is T/C, M is A/C, R is A/G, and K is G/T, same conditions as for the LTR primers but with 30 cycles). For amplification of rabbit mitochondrial DNA, the first-stage primers were CTAGTACACTTACCCAACATAGAG and ATGGGGATAGGAAGTCACATTGAT (cycles were same as for IN primers) and second-stage primers were TAGCCATAGCATTCCTCACCTTAG and GCGATAATAAAGAGTAGCGGAGAG (same as for IN but with 30 cycles).
Cloning of HRV-5 proviral clones from rabbit DNA.
HRV-5 proviral clones were amplified from 200 ng of SIRC DNA using the Expand Long Template PCR kit (Roche). The 3A locus was amplified with primers G7F2 (TGGTGCCGTGACTCGGATAGGA) and L1R1 (TTAAGGACAGAGATCCCACATAAG), the 5A locus using primers K12F1 (CAGCCCTGGCCGGAACTGTTGCAG-3) and U3R3 (TTGATACGCGCCTGTGTAAAGTTC), and the 5B locus using primers FLANK-R1 (GCTGGAGTCACATATATAAGTGACC) and U3R3. Cycling conditions for each PCR were 30 cycles of 94°C, 1 min; 55°C, 1 min; and 68°C, 13 min with an initial denaturation at 94°C for 3 min. PCR products were gel purified from agarose using the QIAquick gel extraction kit (Qiagen). Clone 3A PCR products were cloned into EcoRV-digested pBluescript II KS(−) (Stratagene) using standard methods (32). Clone 5B PCR products were digested by HindIII and EcoRV, and the three restriction fragments obtained were blunt ended and cloned into EcoRV digested pBluescript II KS(−).
Copy number estimation.
A serial dilution of SIRC genomic DNA was transferred onto a nylon membrane (Genescreen Plus; New England Nuclear) using a slot blot system (Bio-Rad). As a standard, we used the reference HRV-5 plasmid pHRV5.6 linearized by digestion with NotI at 25 ng, 5 ng, 1 ng, 200 pg, 40 pg, 8 pg, and 1.6 pg (1 ng is 108 copies). The membranes were hybridized with [32P]dCTP-labeled probes derived from HRV-5 gag, pro, pol, and LTR and washed under conditions of high stringency as described for the Southern blots. Following autoradiography, the films were photographed and analyzed with a digital imaging system (ds ID version 3; Kodak Digital Sciences). The copy number of HRV-5 in the rabbit genome was estimated with the assumption that the size of the rabbit genome was 3 × 109 bp.
RNA extractions.
RNA extractions were performed by the acidified guanidinium isothiocyanate procedure (8) with the RNA Bee reagent (Biotecx Laboratories Inc.). RNA was extracted from cells in culture by using RNA Bee as recommended by the manufacturer. RNA from tissue was also prepared as recommended except that tissue fragments were homogenized on ice in RNA Bee (1 ml per 50 mg of tissue) with an Ultra-Turrax T25 tissue grinder (IKA Labortechnik) at maximum speed prior to the addition of chloroform. RNA pellets were resuspended in nuclease-free water and were stored at −80°C. Prior to RT-PCR, RNA was treated with RQ1 RNase-free DNase (10 U per μg of RNA; Promega) as recommended.
Northern hybridization.
Twelve-microgram aliquots of total RNA were electrophoresed on a 1.2% agarose gel containing 2% formaldehyde and were blotted onto a nylon membrane (Genescreen Plus; New England Nuclear) as recommended by the manufacturer. The blots were hybridized with the 357-bp HRV-5 MA probe, and the 300-bp rabbit GAPDH probe was used for the Southern blots under stringent conditions (final wash, 65°C, 0.1% SDS-2× SSC).
RT-PCR.
RT-PCR was performed essentially as described previously (15). One microgram of RQ1-treated RNA was reverse transcribed using a mixture of reverse primers (GAPDH-R, IN-R, and nucleocapsid [NC]-R1 [CTGAAATTGTTTCYGCCCTCACCT]), and one-fifth of the products was amplified by PCR for IN, GAPDH, and NC. For IN and GAPDH PCR primers and conditions used were the same as described above, except that IN PCR was performed for 45 cycles. NC PCR used primers NC-F3 (GCAGGGGCATCTAATGAGGGAATG) and NC-R1 (35 cycles of 94°C, 45 s; 60°C, 45 s; and 72°C, 45 s). Control reactions were performed in which RT was omitted in order to check for DNA contamination of the RNA preparations. The products were analyzed by electrophoresis through a 3% ethidium-stained agarose gel prior to cloning and sequencing.
DNA sequencing.
Fluorescent DNA sequencing was performed on a Beckman-Coulter CEQ2000 automated DNA sequencer. Computer-aided analysis of nucleotide sequences was performed using the Sequencher program (Gene Code). Multiple sequence alignments were prepared using CLUSTAL W (37).
Nucleotide sequence accession number.
The following sequences have been deposited in the GenBank database: composite complete HRV-5 sequence obtained from human samples (plasmid pHRV5.6) (AF480924), integration site 5A (AF480928), integration site 5B (AF480929), integration site 3A (AF480927), HRV-5/rabbit ERV H (RERV-H) clone A from rabbits (AF480525), and HRV-5/RERV-H clone B from rabbits (AF480926).
RESULTS
Cloning the HRV-5 genome and viral integration sites.
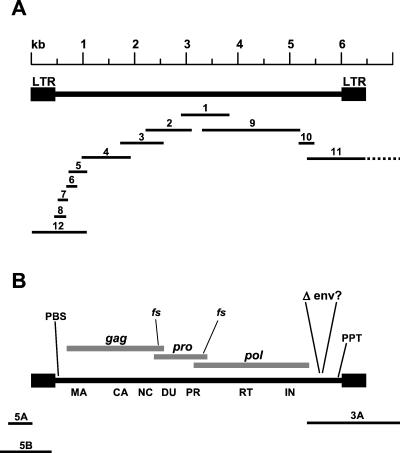
In a previous report (15), the identification of a 930-bp fragment of the HRV-5 pro/pol sequence was described. Although HRV-5 has since been detected in DNA extracted from many patients by independent groups (6, 14, 19, 27), the virus load in these samples was always too low to permit cloning the viral genome by library hybridization using the pro/pol fragment as a probe. Therefore, additional regions of the viral genome were cloned as a series of overlapping fragments using PCR-directed DNA walking (Fig. 1A). Complete details of the methods used to clone the HRV-5 genome are presented in supplementary material (available from http://www.ucl.ac.uk/windeyer-institute/Research/Griffiths.htm or on request from the authors). The sequence obtained had ORFs for gag, pro, and pol and two LTRs, but no env gene was detected in clones obtained independently from four individuals. Within each gene there was sequence variation between independent PCR products as previously observed (15). A composite viral genome was assembled from these PCR fragments (plasmid pHRV5.6, GenBank accession no. AF480924 [Fig. 1B]).
FIG. 1.
Cloning the HRV-5 genome and proviral flanking sequences. (A) HRV-5 was cloned as a series of overlapping PCR fragments (see supplementary material), which are shown in relation to the complete cloned genome. Information for fragment 1 was published previously (15). Fragments 2, 9, and 10 were cloned using degenerate primers for conserved regions of NC and IN. Fragments 3, 4, 8, and 11 were cloned using the Vectorette procedure (1). Fragments 5, 6, and 7 were cloned using a 5′ rapid amplification of cDNA ends modified for amplifying from genomic DNA. Fragment 12 was cloned using specific primers. Each new fragment was confirmed as contiguous with the previously identified sequence by PCR across the junction using specific primers. The dotted line represents DNA flanking the provirus that was cloned as part of fragment 11 (clone 3A). (B) Diagram of HRV-5 genome assembled from the various PCR fragments and cloned as plasmid pHRV5.6 (GenBank accession no. AF480924). The genome has ORFs for gag, pro, and pol and two LTRs but lacks env. The locations of putative protein-coding domains are marked, including MA, CA, NC, deoxyuridine triphosphatase (DU), PR, RT/RNase H (RT), and IN. Regulatory motifs including a polypurine tract (PPT), a primer binding site (PBS; specific for tRNA-His), and heptanucleotide frameshifting sequences (fs) between gag/pro and pro/pol are present. The locations of three PCR products containing integration sites are indicated (5A, 5B, and 3A).
Once the LTRs of HRV-5 had been identified, we cloned regions flanking the proviruses to examine their integration sites. Three flanking sequences were obtained, each from a different patient sample, again using PCR techniques for DNA walking (Fig. 1B). The first, a 3′ flanking sequence, was cloned from sample A by Vectorette PCR (1) using primers anchored in IN (denoted clone 3A). The second (clone 5A), a 5′ flanking sequence, was cloned from sample B by an arbitrarily primed PCR method (C. Cousens, University of Edinburgh, personal communication) with primers located in the 5′ LTR. The third (clone 5B) was cloned from sample C by capture PCR (21) also anchored in the viral 5′ LTR.
Of the three integration sites obtained, none was present in the human genome sequence databases (22, 38), although clones 3A and 5B had limited similarity with repetitive elements. These were a rabbit LINE element (GenBank accession no. X15966.1, 85% similarity with clone 3A over 387 bp) and a human MER2 repeat (in a human BAC clone, GenBank accession no. AC019206, 100% similarity with clone 5B over 38 bp), respectively. Clone 5A had no significant match in GenBank.
We then designed specific PCR assays for these integration site sequences and tested a range of human DNA samples. We reasoned that, if these sequences are from human sources, these assays should detect them in human DNA from any source, despite their absence from sequence databases. All samples tested were negative, except for those from which each integration site had originally been cloned (data not shown). This result indicated that sequences flanking HRV-5 proviruses in our samples were not of human origin and raised doubts about the human origin of HRV-5. To investigate this further, we expanded an earlier survey (13) of the presence of HRV-5 in other species.
Sequences indistinguishable from those of HRV-5 are present in rabbit genomic DNA.
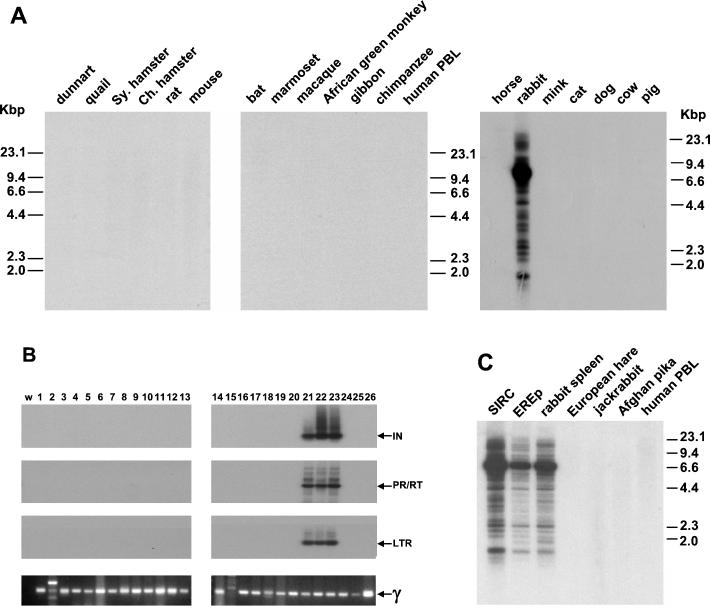
DNA was extracted from cell lines or tissues representing over 20 mammalian and avian species and was analyzed by Southern hybridization with a probe derived from the 5′ region of gag (Fig. 2A). As a loading control, the blots were stripped and hybridized with a rabbit GAPDH probe, which is sufficiently conserved to hybridize to all the species present under low-stringency wash conditions (not shown). We also tested the same samples by PCR using several sets of HRV-5-specific primers (Fig. 2B). To check for DNA quality, we used a degenerate primer set for gammaretroviral pol sequences, which are present in most mammalian genomes. With both methods we found that, of the species studied, only the European rabbit (O. cuniculus) contained HRV-5 sequences (Fig. 2A and B). It is notable that HRV-5 sequences are not present in other members of the order Lagomorpha tested, namely, the Afghan pika (O. rufescens), the European brown hare (L. europaeus), and the black-tailed jackrabbit (L. californicus) (Fig. 2B and C).
FIG. 2.
HRV-5 is an ERV of rabbits. Genomic DNA from several mammalian and avian species was tested for the presence of HRV-5 sequences. (A) Southern blots probed with the MA domain of HRV-5 gag. Kbp, kilobase pairs. (B) PCR products obtained from the same species with primers derived from HRV-5 PR/RT, IN, and LTR and with degenerate primers for a gammaretrovirus pol sequence (γ). Following PCR, the HRV-5 products were transferred to a nylon membrane and were probed with an internal fragment specific for each product. Lanes: w, water; 1, fat-tailed dunnart; 2, quail; 3, Syrian hamster; 4, Chinese hamster; 5, rat; 6, mouse; 7, bat; 8, marmoset; 9, macaque; 10 African green monkey; 11, gibbon; 12, chimpanzee; 13, human (donor blood); 14, human (HeLa); 15, horse; 16, mink; 17, cat; 18, dog; 19, cow; 20, pig; 21, rabbit (SIRC); 22, rabbit (EREp), 23, rabbit spleen; 24, European hare; 25, black-tailed jackrabbit; and 26, Afghan pika. HRV-5 sequences were detected in rabbit DNA from three sources. (C) Southern blot of several species of Lagomorpha probed with the MA domain of HRV-5 gag.
To estimate the copy number of HRV-5 in the rabbit genome, we performed hybridization experiments with several HRV-5 probes on titrated rabbit DNA and compared this with a titration of the pHRV5.6 plasmid as a standard. The results indicate that HRV-5 has increased to around 5,000 copies per haploid genome since it entered rabbits (data not shown). In addition, Southern blot results from three different sources of rabbit DNA showed slightly different banding patterns (Fig. 2C), indicating that some polymorphism exists between different rabbits.
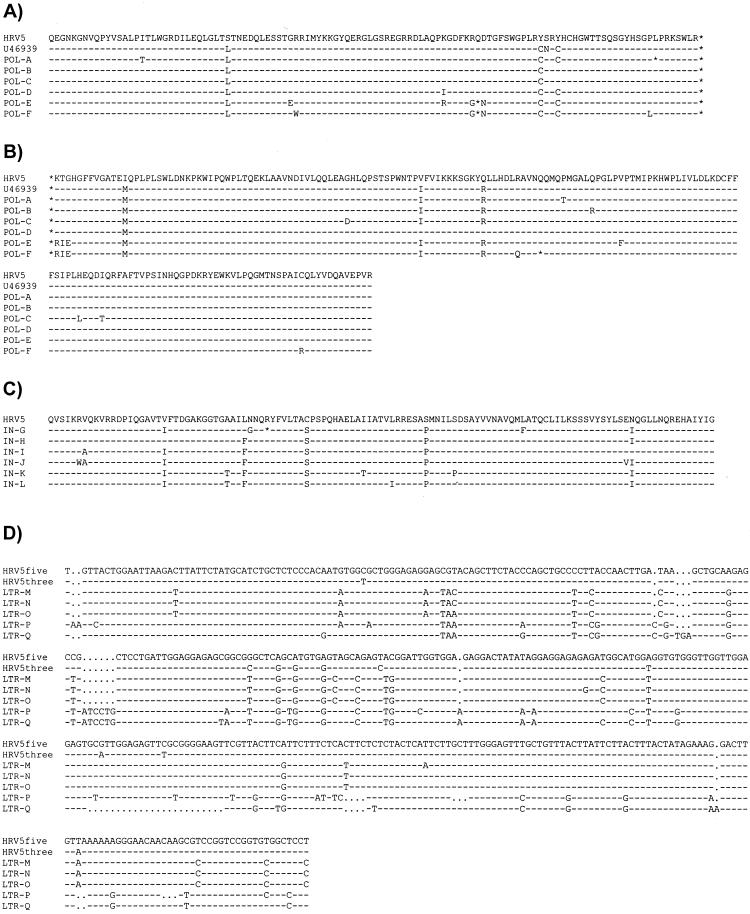
The PCR products from HRV-5 PR/RT, IN, and LTR that were amplified from rabbit DNA were cloned into a plasmid vector, and several clones of each product were sequenced. For PR/RT and IN, the clones obtained were almost identical (>97% nucleotide similarity) to the sequences previously cloned from human DNA (Fig. 3), although the ORFs were not maintained in every clone. For the LTR, up to 15% divergence was observed in the clones sequenced. These data confirmed that the HRV-5 sequences identified in rabbits by Southern hybridization and PCR were indistinguishable from HRV-5 sequences found in humans.
FIG. 3.
HRV-5 sequences amplified from rabbit DNA. Sequences amplified from rabbit DNA by PCR (Fig. 2B) are shown aligned with the reference clone (HRV-5). (A) Deduced amino acid sequence of the pro ORF amplified by PR/RT primers (clones A to F). U46939, previously published clone (15). (B) Deduced amino acid sequence of the pol ORF amplified by PR/RT primers. (C) Deduced amino acid sequence of the region of pol amplified by IN primers (clones G to L). (D) Nucleotide sequence amplified by LTR primers (clones M to Q). Both the 5′ (HRV5five) and 3′ (HRV5three) LTRs of the reference clone are shown. The sequences of several representative plasmid clones are presented. A dash indicates identity with the reference clone (GenBank accession no. AF480924); a period indicates the presence of a gap in the alignment. Differences are marked and stop codons are indicated by an asterisk. Primer sequences are omitted.
Rabbit DNA is present in human DNA samples.
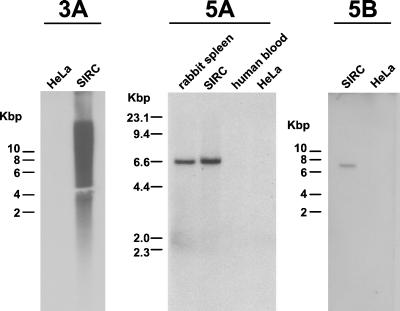
The identification of HRV-5 sequences in rabbit DNA led us to further evaluate the viral integration site sequences obtained from human samples. Using PCR (data not shown) and Southern hybridization, we found that all three flanking sequences were present in rabbit but absent from human DNA (Fig. 4). Thus, this experiment confirmed that the sequences flanking HRV-5, which were originally cloned from human DNA preparations, were actually rabbit genomic sequences. Since there is no known mechanism for a retrovirus to carry both its 5′ and 3′ flanking regions with it during infection, this indicated that our human DNA samples contained rabbit DNA. To test this, we developed a PCR assay specific for rabbit mitochondrial sequences and tested a selection of our DNA preparations. Four of the five samples tested were positive with this assay, confirming that a trace amount of rabbit DNA (and therefore HRV-5 itself) was present in at least some human DNA preparations as contaminating sequences.
FIG. 4.
Sequences flanking HRV-5 proviruses are present in rabbit genomic DNA. Southern blots of rabbit and human DNA were hybridized with probes derived from the three HRV-5 integration site sequences cloned from patient DNA. Each of the integration site clones (3A, 5A, and 5B) hybridizes only with rabbit DNA (from spleen or SIRC cells) and not with human DNA (from blood or HeLa cells), confirming that they are derived from rabbits only. Clones 5A and 5B appear to be single-copy elements, while clone 3A is a rabbit LINE element present at a high copy number.
HRV-5 RNA is expressed in rabbit tissues.
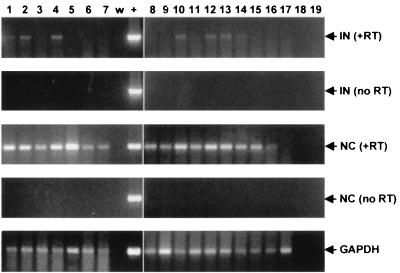
In order to characterize HRV-5 further, we analyzed the transcription of this ERV in a variety of rabbit tissues by RT-PCR with HRV-5-specific primers (Fig. 5). Of the 15 tissues and one cell line studied, all had detectable HRV-5 gag RNA and some also had detectable pol RNA. The PCR signal for pol (obtained with primers for IN) was weaker than for gag (NC), even when 45 cycles of amplification were used. This could be due to the expression of a defective transcript with a deletion in pol but is likelier due to a difference in the efficiency of the two primer sets. All samples were negative by Northern analysis when hybridized with a probe derived from the MA domain of Gag (data not shown), indicating that the expression level was low in these tissues.
FIG. 5.
HRV-5 RNA expression in rabbit tissues. The expression of HRV-5 RNA was analyzed in several rabbit tissues by RT-PCR with primer sets from gag (NC) and pol (IN). Lanes: 1, spleen; 2, kidney; 3, liver; 4, ovary; 5, stomach; 6, colon; 7, pancreas; 8, heart; 9, skeletal muscle; 10, lung; 11 skin; 12, brain; 13, placenta; 14, mammary gland; 15, lymph node; 16, SIRC (rabbit corneal epithelial cell line), 17, human L363 cell RNA; w, PCR water control; +, positive control (rabbit DNA); 18, water control from the DNase treatment step; and 19, water control from the primer-annealing step. Experiments were performed with or without RT in the cDNA synthesis reaction to control for contamination with cellular DNA. PCR primers for rabbit GAPDH were used as controls for RNA quality.
Cloning of HRV-5 proviruses from rabbits.
To obtain additional sequence information on the HRV-5 proviruses in rabbits, we used PCR to amplify the proviruses flanking the three known integration site sequences. PCR from the proviruses adjacent to flanking sequences 3A and 5B produced full-length (but lacking env) HRV-5 proviruses. PCR with the 5A sequence did not yield a band representing an HRV-5 provirus. The 3A PCR fragment was cloned into a plasmid vector, and a single clone was sequenced (denoted clone A). Attempts to clone the 5B PCR fragment in a plasmid vector were unsuccessful, and so this product was digested with EcoRV and HindIII and cloned as three separate fragments. A single plasmid of each of these three fragments was sequenced, and the assembled sequences were denoted clone B.
Sequence analysis of HRV-5 clones A and B amplified from rabbits showed that they are both defective proviruses. Clone A has complete ORFs for pro and pol but has one frameshift mutation (a 37-bp deletion) in the capsid (CA) domain of gag and one in-frame nonsense mutation in NC. Clone B has a defective gag due to two insertions (4 and 11 bp) close to the N terminus, an intact pro and a defective pol due to two in-frame nonsense mutations, and two frameshift mutations (one single-nucleotide insertion and one single-nucleotide deletion). Clones A and B have 97.7 and 96.3% sequence identity, respectively, with the reference HRV-5 clone amplified from human DNA samples and 96.5% identity with each other.
DISCUSSION
HRV-5 is a RERV.
To date, four distinct infectious retroviruses of humans have been discovered, HTLV-1 (29) and -2 (18) and HIV-1 (2) and -2 (9). In addition, simian retroviruses, such as Mason-Pfizer monkey virus (5) and simian foamy viruses (24), have been reported in humans, although these infections appear to be rare zoonoses that are not passed from human to human. There have also been numerous other claims for the existence of infectious retroviruses of humans, most of which were later shown to be mistaken (39), while others have yet to be confirmed (10, 17, 28).
Despite the controversial history of human retrovirology, the discovery of HRV-5 sequences in patients with inflammatory disease (14, 15) appeared to represent a genuine human retrovirus, and this hypothesis was supported by several lines of evidence. Firstly, the original pro and pol sequence fragments of HRV-5 were identified by RT-PCR from RNA isolated from salivary gland homogenates fractionated on a sucrose density gradient (15). The fractions containing HRV-5 RNA had a density of 1.16 g/ml, which is the typical density of enveloped retrovirus virions. Secondly, genome fragments amplified from human samples had complete ORFs for gag, pro, and pol and showed up to 10% sequence divergence between amplicons (14, 15, 19, 27, 30, 31). Such variation with maintenance of ORFs would be expected of an actively replicating retrovirus. Thirdly, HRV-5 sequences have been identified in numerous (almost 100) individuals using nested PCR, although it is clear that HRV-5 is not an ERV in the human genome (see Fig. 2A and reference 15). Indeed, in almost all the patients where it has been detected, HRV-5 is present at an extremely low proviral load (14, 15). Finally, other groups have independently detected HRV-5 sequences in their own clinical samples, although to date only two laboratories (working together) have published their data. Blomberg, Murovska, and their coworkers described an association between HRV-5 and non-Hodgkin's lymphomas of both T- and B-cell origin (19, 27). In addition, we have anecdotal evidence of HRV-5 detection in human DNA samples from five other laboratories in the United States and Europe (Griffiths et al., reply), although at least one group has been unable to detect HRV-5 in RA (11).
Taken together, the previous data on HRV-5 supported the notion that this is an infectious retrovirus of humans; our objective at the outset of the present study was to confirm this hypothesis by cloning the viral genome and integration sites (Fig. 1). Our results led us to the unexpected finding that HRV-5 is in fact an ERV family from rabbits (Fig. 2 and 3). At first glance, this finding might suggest that the occasional detection of HRV-5 sequences in humans is a result of a zoonotic infection from rabbits. However, the identification of rabbit DNA sequences unrelated to HRV-5 in human DNA preparations (Fig. 4 and mitochondrial PCR data) indicates that this may not be the case and that contamination or some other mechanism is responsible for the presence of HRV-5 sequences in human DNA samples. Although we have not proved that zoonotic transfer of HRV-5 to humans cannot occur, attempts to identify human integration sites in our samples using Alu-PCR (25) have so far been unsuccessful (data not shown).
How did rabbit DNA come to be present in human DNA preparations? The simplest explanation is that contamination occurred in the laboratory. Certainly there is a potential for this, since rabbit-derived reagents such as antisera and microsomal membrane preparations are widely used in many laboratories and are likely to contain traces of rabbit DNA. In addition, rabbits are housed in many research institutions for production of antibodies and other studies, so this also presents a source of rabbit DNA that could contaminate human samples. However, laboratory contamination by DNA cannot explain either the initial detection of HRV-5 as an RNA element (15) or its more frequent detection in samples from specific inflammatory diseases and lymphomas (14, 19, 27).
During the course of our studies on HRV-5, we have acquired tissue and DNA samples from a variety of collaborating centers and from our own patients, with DNA extracted in house or by the collaborating laboratory. Thus, it could be argued that there is a chance that samples from RA and SLE were contaminated while sample sets from other diseases or controls remained clean, perhaps because samples from RA and SLE are handled more frequently than control samples in the collaborating laboratories. However, although there may have been greater opportunity for contamination of some sample sets than for others, in most cases test samples were extracted at the same time as controls in the same laboratory using the same reagents and yet still a disease bias was observed. Therefore, while it appears to be the most obvious explanation, laboratory contamination during DNA extraction is in fact an unlikely source of HRV-5 sequences in human samples.
If rabbit DNA was not introduced into our human samples during or after extraction, we must assume that these sequences were present in the tissues prior to their purification from the clinical material. Could normal contact between humans and rabbits, e.g., as pets or food, lead to sufficient transfer of rabbit material to explain our data? This is difficult to estimate, but we can speculate that rabbit meat eaten as food could be a potential source of rabbit DNA in some of our human samples. It has been shown previously that M13 bacteriophage DNA fed to mice can later be detected in the peripheral blood and organs (33). Furthermore, in the same study M13 sequences in spleen were found to be integrated into mouse chromosomal DNA. In humans, it is well established that gut-derived bacterial antigens and DNA can be detected in the peripheral blood and may accumulate in joints (12, 34, 35). Clearance of this material from joints may be less efficient in patients with inflammatory disease. Despite these precedents, we consider that this hypothesis is rather tenuous, although a search for other food-derived DNA in human samples may be worthwhile. Unfortunately, at this time we can offer no other plausible explanation for the selective detection of HRV-5 in samples from RA and SLE.
RERV.
Very little has previously been reported on retroviruses in rabbits. Particles with manganese-dependent RT activity and antigenic similarity with mammalian C-type retroviruses have been described in rabbit lymphosarcoma cell lines and placenta (3). These particles have not been characterized further but are most likely expressed by a RERV related to the gammaretroviruses. HRV-5 is therefore the first retrovirus of rabbits to be described at the molecular level, and we suggest that HRV-5 now be denoted RERV-H, indicating the tRNA-His specificity of the primer binding site.
Phylogenetic analysis based on the sequence of the RT protein has shown that HRV-5/RERV-H clusters in the betaretrovirus genus (14) and is most closely related to (though clearly distinct from) IAPs of mice and other rodents. While this may suggest a common origin with IAPs, the absence of RERV-H from other lagomorphs (Fig. 2C) indicates that rabbits acquired this virus after the divergence of rabbits and hares some 7 million years ago (7) and thus long after the separation of Lagomorpha and Rodentia. This time scale for the entry of RERV-H into the rabbit genome would be relatively recent for an ERV and is consistent with the retention of long ORFs for the viral coding sequences. Future work on RERV-H will aim to determine the age of these elements more accurately, for instance by analysis of LTR sequence divergence for individual proviruses. In addition, the distribution of RERV-H will be investigated in greater detail. The family Leporidae contains 58 different species (23), some of which are more closely related to O. cuniculus than are the hares tested in this study; thus, it remains possible that some other species also possess RERV-H elements.
Our failure to identify env sequences for HRV-5/RERV-H in human samples can perhaps be explained in light of the discovery that this is a RERV, since ERVs commonly lose env at many, if not all, proviral loci (4). We have attempted to clone RERV-H env from rabbit genomic DNA and cDNA using primers based in pol and the 3′ LTR but without success thus far (data not shown). Nevertheless, we would predict that some RERV-H loci do contain env but that these are not detectable by our PCR systems due to mutations or polymorphisms.
Transcription of HRV-5/RERV-H RNA was detected in many rabbit tissues and a corneal cell line by RT-PCR but not by Northern hybridization (Fig. 5), indicating that, although most tissues express RERV-H RNA, expression is not highly activated, even in placenta, which is a common site of ERV expression in other species. It should, however, be noted that almost all these tissues came from a single rabbit (placenta was from a second rabbit) and that RERV-H expression may be increased in different rabbit strains or under different pathological or physiological conditions. Since at least some RERV-H loci have retained long ORFs, it is possible that some of these elements are capable of retrotransposition, and this is worthy of further investigation. Such activity may perhaps explain the polymorphism observed on Southern blots of DNA from different rabbits (Fig. 2C), although this could also simply be due to postintegration mutations creating restriction fragment length polymorphisms.
The etiology of autoimmune and inflammatory diseases such as SLE and RA is unknown. Retroviruses have been cited as potential environmental triggers or modulatory factors in these diseases, due to their ability to establish chronic persistent infections, their ability to cause immune dysfunction, and the occurrence of inflammatory disease in HIV and HTLV infections (10). However, strong experimental evidence linking retroviruses with these diseases is still lacking. The identification of HRV-5/RERV-H in samples from patients with Sjögren’s syndrome, SLE, and RA initially appeared to provide a credible candidate for an etiological viral agent in these diseases. By focusing on the characterization of the virus genome and integration sites, we have determined the provenance of HRV-5 as nonhuman, casting doubt on its role as an etiological agent in human disease. Our studies on HRV-5 illustrate the potential of using PCR technology for discovering and characterizing novel viruses but demonstrate the importance of validating the role of any candidate pathogen by additional means.
Acknowledgments
We thank Christina Cousens for stimulating discussions and for providing unpublished protocols for DNA walking. We are grateful to Shona Douglas, Jean-Yves Madec, Jo Martin, and The Zoological Society of London for providing animal tissues for this study and to Alex Brand, Kevin Davies, Ruth Jarrett, Robin Patel, and Andrew Wakefield for providing clinical tissues and DNA. We thank Richard Jenner for providing L-636 cell RNA and Stuart Neil for useful comments and discussion on the manuscript.
This work was supported by the Arthritis Research Campaign and Abbott Laboratories.
REFERENCES
- 1.Allen, M. J., A. Collick, and A. J. Jeffreys. 1994. Use of vectorette and subvectorette PCR to isolate transgene flanking DNA. PCR Methods Appl. 4:71-75. [DOI] [PubMed] [Google Scholar]
- 2.Barre-Sinoussi, F., J. C. Chermann, F. Rey, M. T. Nugeyre, S. Chamaret, J. Gruest, C. Dauguet, C. Axler-Blin, F. Vezinet-Brun, C. Rouzioux, W. Rozenbaum, and L. Montagnier. 1983. Isolation of a T-lymphotropic retrovirus from a patient at risk for acquired immune deficiency syndrome (AIDS). Science 220:868-871. [DOI] [PubMed] [Google Scholar]
- 3.Bedigian, H. G., R. R. Fox, and H. Meier. 1978. Induction of type C RNA virus from cultured rabbit lymphosarcoma cells. J. Virol. 27:313-319. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Boeke, J. D., and J. P. Stoye. 1997. Retrotransposons, endogenous retroviruses, and the evolution of retroelements. In J. M. Coffin, S. H. Hughes, and H. E. Varmus (ed.), Retroviruses. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y. [PubMed]
- 5.Bohannon, R. C., L. A. Donehower, and R. J. Ford. 1991. Isolation of a type D retrovirus from B-cell lymphomas of a patient with AIDS. J. Virol. 65:5663-5672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Brand, A., D. J. Griffiths, C. Herve, E. Mallon, and P. J. Venables. 1999. Human retrovirus-5 in rheumatic disease. J. Autoimmun. 13:149-154. [DOI] [PubMed] [Google Scholar]
- 7.Casane, D., N. Dennebouy, H. de Rochambeau, J. C. Mounolou, and M. Monnerot. 1997. Nonneutral evolution of tandem repeats in the mitochondrial DNA control region of lagomorphs. Mol. Biol. Evol. 14:779-789. [DOI] [PubMed] [Google Scholar]
- 8.Chomczynski, P., and N. Sacchi. 1987. Single-step method of RNA isolation by acid guanidinium thiocyanate-phenol-chloroform extraction. Anal. Biochem. 162:156-159. [DOI] [PubMed] [Google Scholar]
- 9.Clavel, F., D. Guetard, F. Brun-Vezinet, S. Chamaret, M. A. Rey, M. O. Santos-Ferreira, A. G. Laurent, C. Dauguet, C. Katlama, C. Rouzioux, D. Klatzmann, J. L. Champalimaud, and L. Montagnier. 1986. Isolation of a new human retrovirus from West African patients with AIDS. Science 233:343-346. [DOI] [PubMed] [Google Scholar]
- 10.Garry, R. F., A. M. Krieg, W. P. Cheevers, R. C. Montelaro, H. Golding, C. D. Fermin, and W. R. Gallaher. 1995. Retroviruses and their roles in chronic inflammatory diseases and autoimmunity, p. 491-603. In J. A. Levy (ed.), The Retroviridae, vol. 4. Plenum Press, New York, N.Y. [Google Scholar]
- 11.Gaudin, P., F. Moutet, P. W. Tuke, and J. A. Garson. 1999. Absence of human retrovirus 5 in French patients with rheumatoid arthritis. Arthritis Rheum. 42:2492-2494. [DOI] [PubMed] [Google Scholar]
- 12.Gerard, H. C., Z. Wang, G. F. Wang, H. El-Gabalawy, R. Goldbach-Mansky, Y. Li, W. Majeed, H. Zhang, N. Ngai, A. P. Hudson, and H. R. Schumacher. 2001. Chromosomal DNA from a variety of bacterial species is present in synovial tissue from patients with various forms of arthritis. Arthritis Rheum. 44:1689-1697. [DOI] [PubMed] [Google Scholar]
- 13.Griffiths, D. J. 1996. Investigation of a novel retroviral element isolated from human salivary gland. Ph.D. thesis. University of London, London, United Kingdom.
- 14.Griffiths, D. J., S. P. Cooke, C. Herve, S. P. Rigby, E. Mallon, A. Hajeer, M. Lock, V. Emery, P. Taylor, P. Pantelidis, C. B. Bunker, R. du Bois, R. A. Weiss, and P. J. Venables. 1999. Detection of human retrovirus 5 in patients with arthritis and systemic lupus erythematosus. Arthritis Rheum. 42:448-454. [DOI] [PubMed] [Google Scholar]
- 15.Griffiths, D. J., P. J. Venables, R. A. Weiss, and M. T. Boyd. 1997. A novel exogenous retrovirus sequence identified in humans. J. Virol. 71:2866-2872. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Hunter, E., J. Casey, B. Hahn, M. Hayami, B. Korber, R. Kurth, J. Neil, A. Rethwilm, P. Sonigo, and J. Stoye. 2000. Retroviridae, p. 369-387. In M. H. van Regenmortel et al. (ed.), Virus taxonomy: the classification and nomenclature of viruses. Seventh report of the International Committee on Taxonomy of Viruses. Academic Press, San Diego, Calif.
- 17.Iversen, O. J. 1990. The expression of retrovirus-like particles in psoriasis. J. Investig. Dermatol. 95:41S-43S. [DOI] [PubMed] [Google Scholar]
- 18.Kalyanaraman, V. S., M. G. Sarngadharan, M. Robert-Guroff, I. Miyoshi, D. Golde, and R. C. Gallo. 1982. A new subtype of human T-cell leukemia virus (HTLV-II) associated with a T-cell variant of hairy cell leukemia. Science 218:571-573. [DOI] [PubMed] [Google Scholar]
- 19.Kozireva, S., S. Lejniece, J. Blomberg, and M. Murovska. 2001. Human retrovirus type 5 sequences in non-Hodgkin's lymphoma of T cell origin. AIDS Res. Hum. Retrovir. 17:953-956. [DOI] [PubMed] [Google Scholar]
- 20.Kuff, E. L., and K. K. Lueders. 1988. The intracisternal A-particle gene family: structure and functional aspects. Adv. Cancer Res. 51:183-276. [DOI] [PubMed] [Google Scholar]
- 21.Lagerstrom-Fermer, M., U. Pettersson, and U. Landegren. 1993. Molecular basis and consequences of a deletion in the amelogenin gene, analyzed by capture PCR. Genomics 17:89-92. [DOI] [PubMed] [Google Scholar]
- 22.Lander, E. S., et al. 2001. Initial sequencing and analysis of the human genome. Nature 409:860-921. [DOI] [PubMed] [Google Scholar]
- 23.Macdonald, D. 2001. The new encyclopaedia of mammals. Oxford University Press, Oxford, United Kingdom.
- 24.Meiering, C. D., and M. L. Linial. 2001. Historical perspective of foamy virus epidemiology and infection. Clin. Microbiol. Rev. 14:165-176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Minami, M., K. Poussin, C. Brechot, and P. Paterlini. 1995. A novel PCR technique using Alu-specific primers to identify unknown flanking sequences from the human genome. Genomics 29:403-408. [DOI] [PubMed] [Google Scholar]
- 26.Moore, R., M. Dixon, R. Smith, G. Peters, and C. Dickson. 1987. Complete nucleotide sequence of a milk-transmitted mouse mammary tumor virus: two frameshift suppression events are required for translation of gag and pol. J. Virol. 61:480-490. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Murovska, M., S. Lejniece, S. Kozireva, M. Koulikovska, H. Yin, and J. Blomberg. 2000. Human retrovirus 5 sequences in peripheral blood cells of patients with B-cell non-Hodgkin's lymphoma. Int. J. Cancer 85:762-770. [DOI] [PubMed] [Google Scholar]
- 28.Perron, H., J. A. Garson, F. Bedin, F. Beseme, G. Paranhos-Baccala, F. Komurian-Pradel, F. Mallet, P. W. Tuke, C. Voisset, J. L. Blond, B. Lalande, J. M. Seigneurin, and B. Mandrand. 1997. Molecular identification of a novel retrovirus repeatedly isolated from patients with multiple sclerosis. Proc. Natl. Acad. Sci. USA 94:7583-7588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Poiesz, B. J., F. W. Ruscetti, A. F. Gazdar, P. A. Bunn, J. D. Minna, and R. C. Gallo. 1980. Detection and isolation of type C retrovirus particles from fresh and cultured lymphocytes of a patient with cutaneous T-cell lymphoma. Proc. Natl. Acad. Sci. USA 77:7415-7419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Rigby, S. P., D. J. Griffiths, R. F. Jarrett, R. A. Weiss, and P. J. Venables. 1998. A new human retrovirus: a role in lymphoma? Am. J. Med. 104:99-100. [DOI] [PubMed] [Google Scholar]
- 31.Rigby, S. P., D. J. Griffiths, R. A. Weiss, and P. J. Venables. 1997. Human retrovirus-5 proviral DNA is rarely detected in salivary gland biopsy tissues from patients with Sjogren's syndrome. Arthritis Rheum. 40:2016-2021. [DOI] [PubMed] [Google Scholar]
- 32.Sambrook, J., E. F. Fritsch, and T. Maniatis. 1989. Molecular cloning: a laboratory manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
- 33.Schubbert, R., D. Renz, B. Schmitz, and W. Doerfler. 1997. Foreign (M13) DNA ingested by mice reaches peripheral leukocytes, spleen, and liver via the intestinal wall mucosa and can be covalently linked to mouse DNA. Proc. Natl. Acad. Sci. USA 94:961-966. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Schumacher, H. R., Jr. 1998. Reactive arthritis. Rheum. Dis. Clin. N. Am. 24:261-273. [DOI] [PubMed] [Google Scholar]
- 35.Schumacher, H. R., Jr., T. Arayssi, M. Crane, J. Lee, H. Gerard, A. P. Hudson, and J. Klippel. 1999. Chlamydia trachomatis nucleic acids can be found in the synovium of some asymptomatic subjects. Arthritis Rheum. 42:1281-1284. [DOI] [PubMed] [Google Scholar]
- 36.Thayer, R. M., M. D. Power, M. L. Bryant, M. B. Gardner, P. J. Barr, and P. A. Luciw. 1987. Sequence relationships of type D retroviruses which cause simian acquired immunodeficiency syndrome. Virology 157:317-329. [DOI] [PubMed] [Google Scholar]
- 37.Thompson, J. D., D. G. Higgins, and T. J. Gibson. 1994. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22:4673-4680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Venter, J. C., et al. 2001. The sequence of the human genome. Science 291:1304-1351. [DOI] [PubMed] [Google Scholar]
- 39.Weiss, R. A. 1984. The search for human RNA tumor viruses, p. 1205-1281. In R. Weiss, N. Teich, H. Varmus, and J. Coffin (ed.), RNA tumor viruses, 2nd ed., vol. 1. Cold Spring Harbor Laboratory, Cold Spring Harbor, N.Y. [Google Scholar]