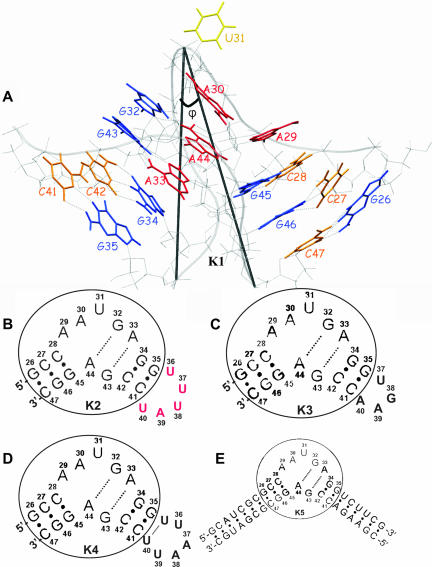
FIGURE 1.
Simulated RNA constructs. (A) The K-turn motif found in the crystal structure of the 5′ stem–loop of U4 snRNA bound to 15.5K protein (K1 RNA); the C-stem has three Watson-Crick base pairs (G26–C47, G46–C27, and G45–C28), the internal loop contains three unpaired nucleotides (A29, A30, U31) and two tandem-sheared G-A base pairs (G32–A44, G43–A33), and the NC-stem consists of two Watson-Crick base pairs (G34–C42 and G35–C41). Guanines are shown in blue, cytosines in orange, adenines in red, and the uracils in yellow. φ is the angle between the P atoms of C47, U31, and G35. (B) K2 RNA (naturally occurring): the pentaloop UUUAU (shown in red) is attached to the NC-stem of K1 RNA. (C) K3 RNA: the tetraloop UGAA was attached to the NC-stem of K1 RNA. (D) K4 RNA: the hexaloop UUAAUU was attached to the NC-stem of K1 RNA. (E) K5 RNA: the C-stem and the NC-stem of K1 RNA were extended with seven and six Watson-Crick base pairs. The core of the K-turn RNA observed in the crystal structure is encircled in B–E.