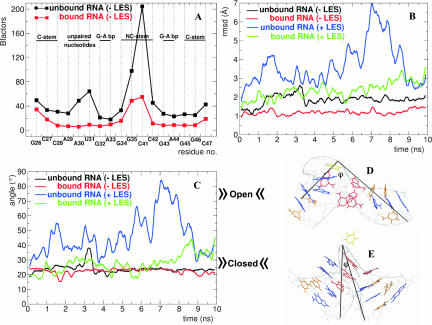
FIGURE 2.
General analysis of MD simulations (K2 RNA). (A) B-factors per residue for the RNA structure during the MD-LES trajectories of the bound (red curve) and unbound RNA (black curve). (B) Root mean square deviations (RMSD) of the RNA backbone from the initial structure for the bound and unbound RNA during the MD-LES (red and black curves) and MD+LES trajectories (green and blue curves). (C) φ angle of the bound and unbound RNA during the MD-LES (red and black curves) and MD+LES trajectories (green and blue curves). (D) Closed conformation of the K-turn (φ = 20°–40°). (E) Open conformation of the K-turn (φ = 50°–80°). In C and D the loop attached to the NC-stem is not shown and the φ angle is shown in black in C and D (see Fig. 1A).