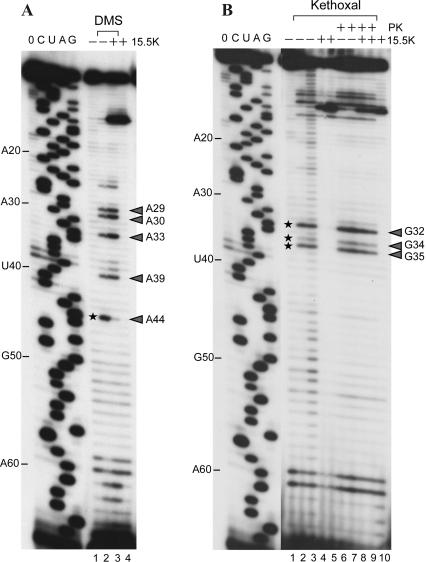
FIGURE 7.
Chemical RNA structure probing. (A) Primer extension analysis of U4 snRNA after DMS treatment of the RNA either in the absence or presence of recombinant 15.5K protein (lanes 2,3); (lanes 1,4) control lanes, no DMS modification. (B) Primer extension analysis of U4 snRNA after Kethoxal treatment either in the absence (lanes 2,3,6,7) or presence (lanes 4,5,8,9) of 15.5K protein; (lanes 6–9) RNA modification after Proteinase K digestion; (lanes 1,10) control lanes, no Kethoxal treatment; the modification of an RNA base results in a stop of the reverse transcriptase 1 nt before the site of attack (Ehresmann et al. 1987); modified nucleotides are indicated by an arrowhead; nucleotides that are clearly protected from chemical modification in the presence of bound 15.5K protein are marked by asterisks; the presence or absence of the 15.5K protein is indicated by “+” or “-,” respectively. C, U, A, and G refer to dideoxysequencing reactions and correspond to the sequence of human U4 snRNA; 0 indicates a control primer extension with unmodified U4 snRNA where no ddNTPs were added to the reaction; and the position of every tenth nucleotide of the U4 snRNA is indicated on the left in panels A and B, respectively.