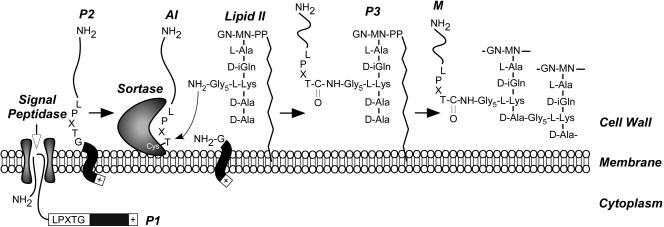
FIG. 5.
Cell wall sorting pathway of surface proteins in gram-positive bacteria. Surface proteins are first synthesized in the bacterial cytoplasm as full-length precursors (P1) containing an N-terminal signal sequence and a C-terminal sorting signal. The signal sequence directs the cellular export of the polypeptide through the Sec system and, upon translocation, is cleaved by signal peptidase. The product of this reaction, the P2 precursor harboring only the C-terminal sorting signal, is retained within the secretory pathway via its C-terminal hydrophobic domain (black box) and positively charged tail (+). Sortase, a membrane-anchored transpeptidase with active-site cysteine, cleaves the peptide bond between the threonine (T) and the glycine (G) of the LPXTG motif, generating an acyl intermediate (AI). Lipid II, the peptidoglycan biosynthesis precursor, and its pentaglycine cross bridge (Gly5) amino group attack the acyl intermediate, linking the C-terminal threonine of the surface protein to lipid II (P3 precursor) and regenerating the active site of sortase. The P3 precursor functions as a substrate for penicillin binding proteins and is incorporated into the cell wall envelope to generate mature anchored surface protein (M), which is also displayed on the bacterial surface. This pathway is universal in many gram-positive bacteria, and the functional elements of cell wall cross bridges, LPXTG motif, sortase, and penicillin binding proteins are conserved.