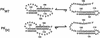Table 1.
Thermodynamic parameters obtained at 1 M KCl
| RNA | Observed
|
Predicted
|
|
|---|---|---|---|
| −ΔHtotal | −ΔGtotal | −ΔGtotal | |
| PKWT | 164.7 | 17.8 | 14.2 |
| PKDC | 124.0 | 11.2 | 8.2 |
| HPWT | 92.0 | 9.8 | 9.3*, 10.1† |
| HPDC | 99.0 | 10.5 | 9.3*, 10.1† |
| CR7WT | 88.9 | 9.7 | 11.2 |
| CR7DC | 87.0 | 5.3 | 7.1 |
ΔH is model-dependent van't Hoff enthalpy. ΔH and ΔG are reported in kcal mol−1. ΔG is calculated at 37°C and compared to nearest-neighbor predictions for stacking (35), pseudoknot loop penalties (36), tandem mismatches, and internal loop free energies (37).
Calculated for the expected hairpin with a six-Watson–Crick base-pair stem and a 17-nucleotide loop.
Calculated based on the observed base pairing, with the pyrimidine tract stability estimated from tandem mismatch data.
The pseudoknot-stem 1 hairpin equilibrium for WT and DC is illustrated below.