Figure 6.
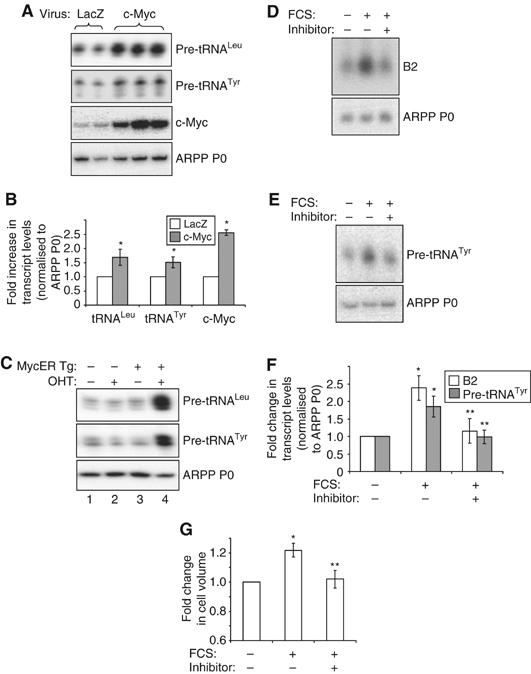
c-Myc regulates pol III transcription in cardiomyocytes. (A, B) Cultured cardiomyocytes were infected with adenovirus expressing Lac Z (control) or c-Myc, as indicated. Cells were grown in the absence of hypertrophic stimuli, then total RNA extracted and used to generate cDNAs by reverse transcription. cDNAs were amplified by PCR using primers specific for the indicated transcripts. (A) Representative PCRs are shown. (B) The average fold increases in transcript levels (normalised to ARPP P0), induced by c-Myc, are represented graphically (n=3; *significantly higher than LacZ-expressing control, P<0.05). (C) c-Myc induces pol III transcription in the heart. RNA was extracted from hearts derived from wild-type adult mice (lanes 1 and 2) or transgenic littermates expressing MycER specifically in cardiomyocytes (MycER Tg, lanes 3 and 4). Mice had either been exposed to 4-hydroxytamoxifen (OHT) (lanes 2 and 4) or vehicle (lanes 1 and 3) for 1 week before removal of hearts. RNA was analysed by RT–PCR using pre-tRNA- and ARPP P0-specific primers, as specified. (D–G) Cells were cultured in the absence of serum (SF) or stimulated with 10% FCS for 24 h. c-Myc inhibitor (‘+') or vehicle (‘−') were also included for 24 h where indicated. Whole-cell RNA was analysed by (D) Northern blotting or (E) RT–PCR. (F) tRNA and B2 levels were quantified by densitometry and normalised to ARPP P0, as represented graphically (n=3; *significantly higher than SF+vehicle, P<0.05; **significantly lower than FCS+vehicle, P<0.05). (G) Cell volume was assayed using a Z2 coulter counter and the average fold changes in cell size are shown. (n=3; *significantly higher than SF+vehicle, P<0.05; **significantly lower than FCS+vehicle, P<0.05).
