TABLE 2.
Results from a reciprocal analysis
| Analytical model | α | β | r | α̂ | β̂ | r̂ |
|
%1 | %2 | %3 | |
|---|---|---|---|---|---|---|---|---|---|---|---|
| Data simulated by the Luo et al. model | |||||||||||
| Wu-Ma | 0.10 | 0.14 | 0.10 | 0.101 (0.028) | 0.139 (0.027) | 0.105 (0.028) | −687 | 100 | 0 | 0 | |
| Luo et al. | 0.10 | 0.14 | 0.10 | 0.100 (0.026) | 0.139 (0.020) | 0.103 (0.029) | −688 | 100 | 0 | 0 | |
| Data simulated by the Wu-Ma model | |||||||||||
| Wu-Ma | 0.10 | 0.14 | 0.10 | 0.097 (0.031) | 0.141 (0.028) | 0.100 (0.025) | −742 | 100 | 0 | 0 | |
| Luo et al. | 0.10 | 0.14 | 0.10 | 0.089 (0.026) | 0.134 (0.021) | 0.113 (0.034) | −765 | 100 | 0 | 0 | |
| Wu-Ma | 0.10 | 0.30 | 0.25 | 0.096 (0.023) | 0.295 (0.034) | 0.256 (0.036) | −683 | 100 | 0 | 0 | |
| Luo et al. | 0.10 | 0.30 | 0.25 | 0.109 (0.035) | 0.209 (0.092) | 0.347 (0.112) | −691 | 97 | 0 | 3 | |
The data simulated by our (Wu-Ma) and Luo et al.'s models are analyzed by the two models, respectively. The marker cross type for a full-sib family of size n = 200 is the one used by Luo et al., expressed as 1220/1222 × 3455/1130. 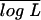 is the mean of the log-likelihood values obtained from 200 simulations. See Table 1 for explanations of the other parameters.
is the mean of the log-likelihood values obtained from 200 simulations. See Table 1 for explanations of the other parameters.