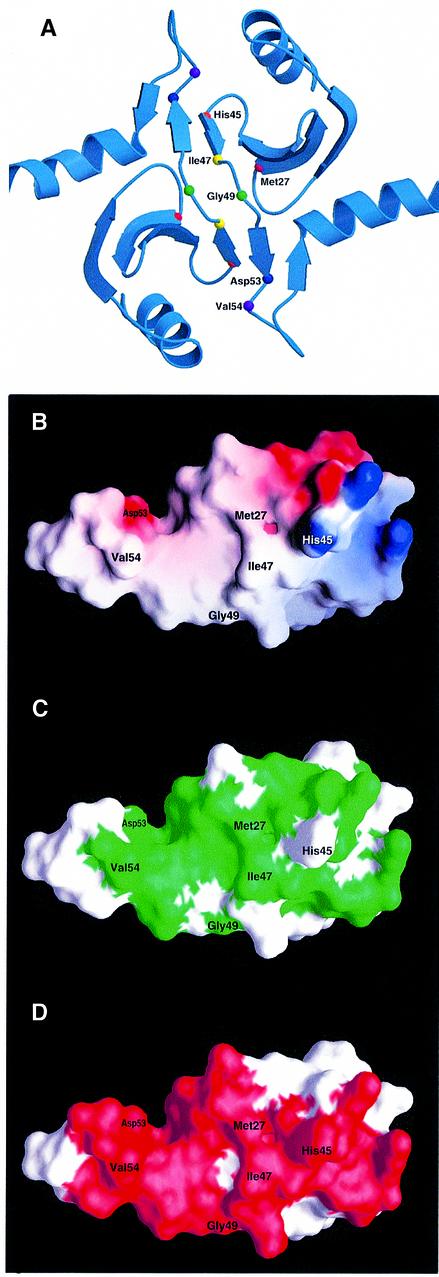
Fig. 3. Rational design of mutants in the dimerization domain of DsbC. (A) The crystal structure of DsbC is shown with the mutated residues represented as small spheres. The monomers interact via an N-terminal dimerization interface that consists of two very short consecutive β-strands from each monomer. Two extended β-sheets are formed, which consist of two strands from one monomer and four strands from the other. Two DsbC mutants that were found to complement a dsbA null mutant were mapped to G49 (drawn as a green sphere), which lies in between the interacting β-strands. These mutations replace G49 with either arginine or glutamate. An additional five residues, which were predicted to provide crucial interactions in the dimerization interface and were subject to mutagenesis, are also shown as small spheres. (B) Surface potential of DsbC’s dimerization interface. Only one monomer is shown. Values range from +7kT (blue) to 0 (white) to –7kT (red). M27, V54 and I47 were mutated to lysines. H45 forms a salt bridge with Asp53 on the other monomer (compare with Figure 3A), and was mutated to an aspartate. (C) Same as (B), except that the green color highlights conserved residues. Conserved residues were mapped according to the sequence alignment of 15 DsbC molecules. A BLAST search was performed with the entire DsbC protein sequence at the NCBI database of finished and unfinished bacterial genomes (http://www.ncbi.nlm.nih.gov/Microb_blast/unfinishedgenome.html). Fifteen microbial sequences were aligned with CLUSTAL W (Thompson et al., 1994). Identical and similar residues that are found in at least 60% of the aligned sequences are shaded in green. (D) Molecular surface of the dimerization domain showing residues involved in dimer formation. Note that most of the residues involved in dimerization are conserved.

An official website of the United States government
Here's how you know
Official websites use .gov
A
.gov website belongs to an official
government organization in the United States.
Secure .gov websites use HTTPS
A lock (
) or https:// means you've safely
connected to the .gov website. Share sensitive
information only on official, secure websites.