TABLE 1.
Assessment of the importance-sampling methods
|
I1
|
I2
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| T | 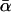 |
SD | SD↓ | SD↑ | ∉C.I. | 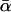 |
SD | SD↓ | SD↑ | ∉C.I. | ISI |
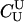 |
5.23e-08 | 0.89 | 0.92 | 0.95 | 9 | — | — | — | — | — | 2.4e+06 |
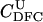 |
7.82e-07 | 1.27 | 1.09 | 1.35 | 13 | — | — | — | — | — | 7.9e+04 |
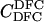 |
1.29e-05 | 1.20 | 1.06 | 1.24 | 11 | 7.76e-06 | 104.26 | 3.07 | 608.03 | 49 | 5.4e+03 |
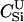 |
8.76e-06 | 2.20 | 1.80 | 4.47 | 6 | 4.39e-06 | 142.90 | 3.24 | 772.98 | 60 | 2.4e+03 |
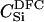 |
1.49e-04 | 1.57 | 1.34 | 2.47 | 9 | 1.47e-04 | 18.52 | 3.95 | 129.35 | 17 | 2.7e+02 |
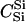 |
1.66e-03 | 1.46 | 1.29 | 2.08 | 6 | 1.67e-03 | 2.54 | 1.70 | 6.17 | 10 | 2.8e+01 |
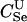 |
5.43e-04 | 14.72 | 4.34 | 67.70 | 24 | 5.53e-04 | 5.39 | 3.08 | 12.36 | 13 | 6.2e+00 |
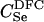 |
8.57e-03 | 6.02 | 2.76 | 30.61 | 12 | 8.59e-03 | 1.02 | 0.89 | 1.07 | 11 | 1.1e+01 |
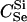 |
7.49e-02 | 3.25 | 1.89 | 9.92 | 9 | 7.48e-02 | 0.49 | 0.54 | 0.56 | 5 | 5.2e+00 |
| BU | 5.08e-04 | 24.40 | 4.82 | 137.21 | 27 | 5.58e-04 | 22.95 | 3.23 | 135.82 | 29 | — |
| BDFC | 2.76e-03 | 23.99 | 5.09 | 137.72 | 22 | 2.91e-03 | 2.10 | 1.52 | 4.83 | 10 | 7.8e+00 |
| BSi | 1.14e-02 | 54.21 | 4.28 | 508.81 | 33 | 1.21e-02 | 0.74 | 0.85 | 1.07 | 4 | 1.5e+01 |
| BSe | 1.04e-01 | 204.62 | 4.04 | 1859.42 | 58 | 1.18e-01 | 0.44 | 0.48 | 0.50 | 7 | 3.9e+00 |
| HU | 8.75e-06 | 2.03 | 1.72 | 3.53 | 8 | 6.64e-06 | 256.77 | 1.78 | 2439.39 | 61 | 2.8e+03 |
| HDFC | 1.48e-04 | 1.91 | 1.34 | 3.26 | 14 | 1.51e-04 | 50.34 | 4.45 | 472.74 | 38 | 1.9e+02 |
| HSi | 1.67e-03 | 1.54 | 1.29 | 3.19 | 8 | 1.66e-03 | 2.22 | 1.66 | 3.96 | 11 | 2.5e+01 |
| HSe | 7.45e-02 | 2.98 | 2.02 | 8.43 | 10 | 7.48e-02 | 0.54 | 0.54 | 0.57 | 8 | 4.2e+00 |
| D1 | 4.52e-02 | 79.70 | 3.89 | 468.64 | 41 | 7.27e-02 | 0.56 | 0.54 | 0.57 | 10 | 4.1e+00 |
| D2 | 1.42e-02 | 125.43 | 3.99 | 1052.07 | 59 | 1.17e-02 | 0.91 | 0.82 | 0.99 | 12 | 1.0e+01 |
| D3 | 1.93e-04 | 15.29 | 3.65 | 52.77 | 23 | 1.90e-04 | 20.17 | 4.13 | 161.01 | 23 | — |
| D4 | 1.67e-03 | 1.36 | 1.30 | 3.41 | 5 | 1.67e-03 | 2.96 | 1.68 | 15.78 | 8 | 3.2e+01 |
| D5 | 7.03e-05 | 162.84 | 4.72 | 1293.03 | 55 | 8.05e-05 | 12.33 | 3.94 | 58.29 | 19 | — |
| PU | 1.29e-09 | 1.33 | 1.26 | 1.31 | 9 | — | — | — | — | — | 4.4e+07 |
| PDFC | 8.79e-07 | 1.13 | 1.24 | 1.35 | 4 | 8.72e-07 | 2.46 | 2.15 | 4.84 | 9 | 8.9e+04 |
| PSi | 1.08e-04 | 1.42 | 1.24 | 1.44 | 14 | 1.08e-04 | 1.17 | 1.04 | 1.10 | 11 | 6.8e+02 |
| PF | 1.46e-04 | 285.69 | 3.04 | 2807.57 | 56 | 1.48e-04 | 1.21 | 1.06 | 1.11 | 14 | 4.6e+02 |
| PH | 6.88e-10 | 117.31 | 3.21 | 709.76 | 48 | 1.06e-09 | 30.62 | 4.68 | 166.84 | 27 | — |
For different relationships, given in the T column, 100 independent Monte Carlo estimates of α corresponding to β = 0.1 were made using both methods I1 and I2. Results for I1 appear on the left and those for I2 on the right. The description of the quantities given in the columns headed by 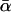 , SD, SD ↓, SD ↑, and ∉C.I. is given in the text. Values for the best-performing importance-sampling method are in italics. For four relationships—BU, D3, D5, and PH—neither I1 nor I2 provided an acceptable reduction in Monte Carlo error. The ISI column gives the factor by which the best importance-sampling algorithm speeds up the estimation of α relative to the naive Monte Carlo estimator of (6). All calculations were done assuming L = 60 loci with minor allele frequencies of q = 0.2.
, SD, SD ↓, SD ↑, and ∉C.I. is given in the text. Values for the best-performing importance-sampling method are in italics. For four relationships—BU, D3, D5, and PH—neither I1 nor I2 provided an acceptable reduction in Monte Carlo error. The ISI column gives the factor by which the best importance-sampling algorithm speeds up the estimation of α relative to the naive Monte Carlo estimator of (6). All calculations were done assuming L = 60 loci with minor allele frequencies of q = 0.2.
