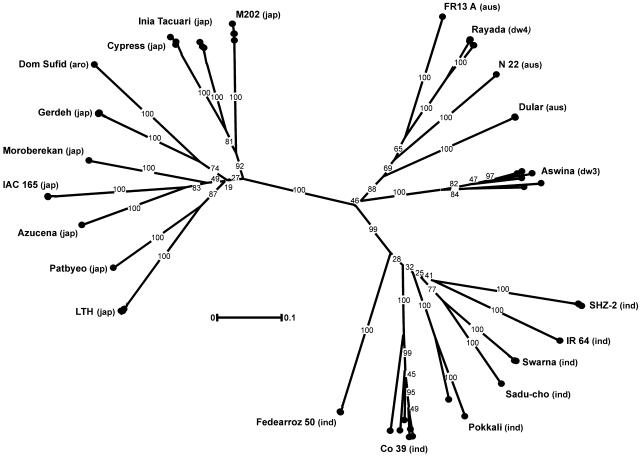
Figure 1.
Dendrogram of potential candidate varieties showing between- and within-variety diversity. For each variety, four to seven individual plants were genotyped using 49 simple sequence repeat markers distributed across the genome. Pairwise distances were calculated using the Dice similarity coefficient for 500 bootstrap replicates and subjected to clustering by unweighted pair group method with arithmetic means. Numbers on branches indicate the percentage of times that group occurred in the strict majority-rule consensus tree. Variety types are annotated as ind (indica), jap (japonica), aro (aromatic), aus (aus/boro), dp3 (deep-water III), and dp4 (deep-water IV).