Abstract
Using a combination of single molecule and bulk solution measurements, we have examined the DNA translocation activity of a helicase, the Type I restriction modification enzyme EcoR124I. We find that EcoR124I can translocate past covalent interstrand crosslinks, inconsistent with an obligatory unwinding mechanism. Instead, translocation of the intact dsDNA occurs principally via contacts to the sugar-phosphate backbone and bases of the 3′–5′ strand; contacts to the 5′–3′ strand are not essential for motion but do play a key role in stabilising the motor on the DNA. A model for dsDNA translocation is presented that could be applicable to a wide range of other enzyme complexes that are also labelled as helicases but which do not have actual unwinding activity.
Keywords: DNA translocation, helicase, motor protein, restriction enzyme, single molecule
Introduction
The ATP-dependent translocation of motor proteins along polynucleotides is a fundamental feature of DNA metabolism. One important class of such motors are the helicases (Lohman and Bjornson, 1996). As defined on the basis of characteristic protein motifs, helicases encompass a large enzyme group with diverse genetic roles. For a great many of these proteins, the catalysis of nucleic acid strand separation is key to their translocation mechanism (Soultanas and Wigley, 2001); they are ‘helicases' as originally defined (Kuhn et al, 1979). The strand separation activity of these classical helicases is often measured using synthetic polynucleotide substrates. However, there are also many enzymes classified as helicases which fail to unwind such substrates. Occasionally this may be because a vital nucleotide or protein component has not been included in the assay. But more often, the lack of unwinding activity reflects the fact that ssDNA or ssRNA production is not required for the cellular role of the enzyme. A more all-encompassing definition of a helicase is that the protein motifs actually form an energy-coupling module based around RecA-folds that converts the binding and hydrolysis of NTPs into mechanical events on DNA or RNA (Singleton and Wigley, 2002).
While the mechanisms of DNA unwinding by helicases have been studied extensively, much less is known about how mechanochemical coupling occurs in other processes catalysed by helicases; for example, the remodelling of polynucleotide structures (e.g., RecG; Singleton et al, 2001) and nucleoprotein complexes (e.g., RSC; Lia et al, 2006). Is nucleic acid strand separation still required for these processes? What mechanistic properties, if any, are shared with the orthodox unwindases? Here, we have addressed these questions using EcoR124I, an Escherichia coli Type I restriction-modification (RM) enzyme that can be defined on the basis of primary amino-acid sequence as a helicase (McClelland and Szczelkun, 2004) and which has DNA translocation activity (Firman and Szczelkun, 2000; Seidel et al, 2004).
EcoR124I is a multifunctional, hetero-oligomeric enzyme complex comprising two main components: a core methyltransferase (MTase), which undertakes sequence-specific DNA recognition and modification; and two HsdR subunits, which are loaded by the MTase onto the adjacent nonspecific DNA and which carry out ATP hydrolysis, DNA translocation and endonuclease activities (Szczelkun et al, 1996; Firman and Szczelkun, 2000; Seidel et al, 2004, 2005; McClelland et al, 2005). ATP hydrolysis is coupled to DNA translocation by the superfamily 2 (SF2) helicase motifs in HsdR (McClelland and Szczelkun, 2004). Each HsdR motor translocates independently on the DNA away from the core MTase (Szczelkun et al, 1996; Firman and Szczelkun, 2000; Seidel et al, 2004, 2005; McClelland et al, 2005). As the HsdRs remain bound to the MTase, which in turn remains bound to the EcoR124I binding site, two DNA loops are extruded (van Noort et al, 2004). The MTase–DNA complex is much longer lived than the MTase–HsdR complex so that termination of a translocation event results in HsdR dissociation from both the DNA and the MTase (Seidel et al, 2005). Another motor event is then initiated by the MTase loading a new HsdR molecule. On viral DNA, motor events will fire repeatedly until cleavage is triggered by the collision of two converging HsdRs. Cleavage of the host genome is prevented by attenuation of productive motor events (Makovets et al, 2004; Seidel et al, 2005).
No evidence of DNA unwinding by Type I enzymes has been reported using classical oligoduplex helicase assays. However, it is not clear whether the substrates utilised were appropriate to the analysis of a heteroligomeric system such as EcoR124I. One significant problem is that motor activity must be measured in the presence of the MTase loading complex, HsdR has no motor activity in isolation (Firman and Szczelkun, 2000). Since HsdR loading requires the MTase to be bound at a specific dsDNA binding site, it is difficult to imagine how strand separation could be observed using a simple dsDNA substrate. Short oligoduplex substrates, as used in most helicase assays, may also fail because their limited size does not support initiation of productive motor events. Our alternative approach has been to investigate the motor kinetics of EcoR124I on long (>1 kbp) modified DNA using a combination of single molecule and bulk solution techniques (Figure 1). Our results are consistent with a mechanism in which EcoR124I translocates without unwinding along the 3′–5′ strand of intact duplex DNA. Contacts to the 5′–3′ strand play a supplementary ‘clamp' role, stabilising the translocating complex. Since Type I enzymes have provided a model system for other helicases, such as the chromatin remodelling factors (Lia et al, 2006), we believe that the dsDNA translocation scheme we present may be generally applicable to other helicase-based motors which do not demonstrate unwinding activity.
Figure 1.

Approaches for examining the translocation of EcoR124I on modified DNA. (A) Magnetic tweezers assay. A DNA molecule with a single EcoR124I binding site and containing suitable modification(s) is tethered between a magnetic bead and a glass slide. Magnets are used to elongate the DNA by the application of different forces. EcoR124I translocation causes loop extrusion that, in turn, shortens the apparent end-to-end length of the DNA, which can be measured using video microscopy. Due to the close location of the EcoR124I site to the magnetic bead (275 bp for ICLs, 90 bp for gaps) mainly unidirectional DNA translocation towards the modification(s) is probed. Examples are shown of: ICLs (red circle) and accompanying monoadducts (red lollipops); and, a 4 nucleotide gap in the 5′–3′ strand. (B) Triplex displacement assay. A labelled triplex is introduced downstream of an EcoR124I binding site, with modifications to the DNA made in the intervening region. Translocation and collision with triplex leads to displacement and an increase in fluorescence, which can be measured using a stopped-flow fluorimeter. Inhibition of translocation can be inferred from changes to the triplex displacement profiles. Examples are shown of: a 4 nucleotide gap in the 3′–5′ strand; and two abasic groups in 3′–5′ strand introduced by annealing a modified oligonucleotide (red strand) into a gap.
Results
Is DNA unwinding a prerequisite of DNA translocation by Type I RM enzymes?
DNA unwinding by an orthodox helicase is facilitated by ssDNA translocation (Lohman and Bjornson, 1996; Soultanas and Wigley, 2001); intact dsDNA cannot be accommodated by the motor. Accordingly, if interstrand crosslinks (ICLs) are introduced into duplex DNA, an obligate unwindase cannot progress (e.g., DNA unwinding by RecBCD is completely blocked on crosslinked substrates; Figure 2A; Karu and Linn, 1972). But what is the situation for a dsDNA translocase? For one such enzyme, E. coli RuvB, bypass of an ICL has been observed, suggesting that ssDNA does not play an extensive role in translocation (George et al, 2000). It has also been reported that Type I RM enzymes can still cleave crosslinked DNA (Karu and Linn, 1972; Endlich and Linn, 1985), although experimental data were never presented. It was therefore concluded that ICLs have no effect on translocation. However, it has been shown subsequently that Type I enzymes cleave DNA when translocation is impeded (Janscak et al, 1999; also, see below). Therefore, ICLs may actually induce endonuclease activity through inhibition of translocation. To resolve these issues, we analysed EcoR124I on DNA in which ICLs were randomly introduced by treatment with 4′-hydroxymethyl-4,5′,8′-trimethylpsoralen (HTMP) and UV light (Karu and Linn, 1972; Hearst, 1981), using conditions which resulted in either a low or high density of corresponding HTMP monoadducts (see Materials and methods).
Figure 2.
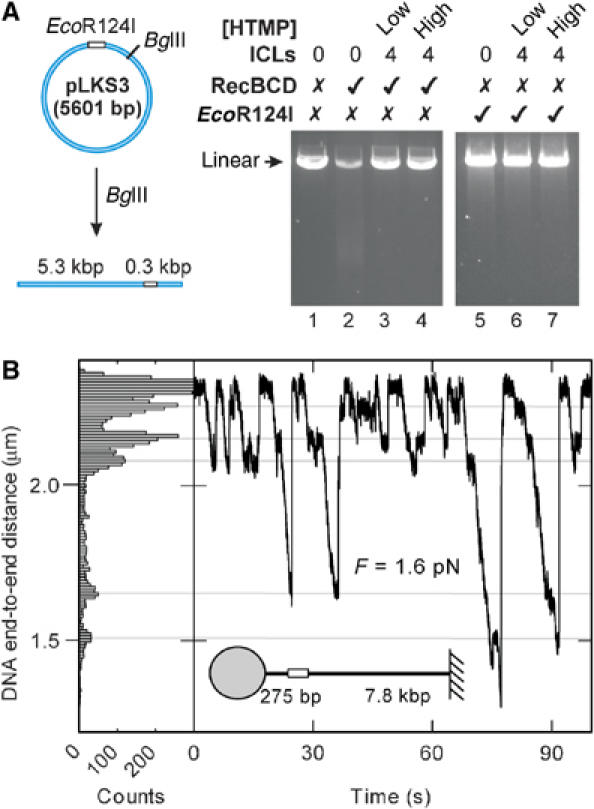
The effect of DNA ICLs on EcoR124I and RecBCD activity. (A) pLKS3 was linearised with BglII to produce a linear DNA substrate. ∼4 ICLs per DNA were introduced at two HTMP concentrations: either <0.033 HTMP/dT (low) or >1 HTMP/dT (high). RecBCD conditions were chosen so that DNA unwinding would occur without DNA cleavage. EcoR124I reactions conditions were chosen that would allow cleavage of circular DNA. DNA substrates and products were separated by gel electrophoresis. (B) Time trace (right panel) recorded in the magnetic tweezers at 20 nM MTase and 160 nM HsdR using a DNA substrate as shown with ICLs introduced at ∼1 ICL/kbp using a low concentration of HTMP (<0.033 HTMP/dT). Multiple motor events are observed over a 90 s window, showing multiple stall events occurring at the same sites on the DNA. (In comparison, on unmodified DNA there is only one stall event corresponding to collision with the bead.) In the left panel is the occupancy of different DNA lengths (all detected lengths of a time trace binned at 10 nm intervals). Peaks in the counts correspond to the full length DNA at ∼2.3 μm (i.e., when the enzyme is not translocating) and correlated stalls at ICLs/adducts (grey lines) and the bead during translocation. If stalling were independent of sequence then the former peaks would not be observed.
We first investigated if ICLs could induce cleavage by analysing linear DNA carrying a single EcoR124I site (Figure 2A). Such substrates are normally not cleaved under the reaction conditions used here (Szczelkun et al, 1996). At an average crosslinking density of ∼1 ICL per 1.4 kbp, no cleavage was observed following extensive incubation with EcoR124I (Figure 2A, lanes 5–7). Therefore, neither ICLs nor the accompanying HTMP monoadducts activate EcoR124I endonuclease activity. We next tested circular substrates with a single EcoR124I site that, in contrast to linear DNA, can be cut under the conditions used here (Szczelkun et al, 1996). As has been suggested previously (Karu and Linn, 1972; Endlich and Linn, 1985), ICLs had no effect on the rate or amplitude of DNA cleavage, although elevated monoadduct levels did block EcoR124I binding (Supplementary Figure 1). We also examined the distribution of cleavage sites on circular DNA as a function of increased crosslinking; no significant differences were observed with between 0 and 8 ICLs per DNA (Supplementary Figure 1). Therefore, during cleavage of crosslinked circular DNA, the same collision/cleavage events must be occurring irrespective of multiple ICLs/adducts.
To observe directly the effects of ICLs on translocation, we measured motor activity of single EcoR124I molecules using magnetic tweezers (Figure 1A; Seidel et al, 2004, 2005). This setup allows us to monitor the end-to-end distance of a single DNA molecule in real time, with EcoR124I translocation seen as characteristic saw tooth-shaped DNA shortening events (Seidel et al, 2004). We introduced ∼6 ICLs into an 8 kbp linear DNA substrate (Figure 2B) under conditions that produced a low monoadduct density (see Supplementary Methods). While typical profiles on non-crosslinked DNA exhibit translocation events with stalling only observed at the DNA bead (Seidel et al, 2004, 2005), profiles on crosslinked DNA exhibited multiple short temporary stalls within a single translocation event (Figure 2B). There was a strong correlation in stall location between successive events, suggesting that they are dependent upon substrate- and site-specific modifications. In many instances, translocation continued after stalling, showing that the modifications could be bypassed. Given our crosslinking conditions, both randomly distributed ICLs and/or monoadducts could be responsible for these events. Nonetheless, for the longest motor runs observed (representing translocation of ≫1 kbp), at least one ICL will have been overcome on average. We also examined the ATPase activity of EcoR124I on modified and unmodified linear DNA; no differences were observed (data not shown).
There is no indication from the single molecule profiles that bypass of DNA damage occurs by DNA release followed by long-range re-association. The same conclusion can be made from the mapping studies (Supplementary Figure 1), where significant changes in motor protein distribution would have been revealed as changes in cleavage site distribution. We confirmed that long-range ‘hopping' by EcoR124I is highly unlikely by testing translocation on a DNA catenane (Supplementary Figure 2). Although these data do not completely exclude short-range hopping, this mechanism of bypass seems unlikely for a Type I enzyme as release of the topologically strained expanding loop(s) would most likely result in a rapid diffusion of the HsdR and DNA away from one another, making rebinding unlikely. This is particularly true in the tweezers where separation of the HsdR and loop would be strongly supported by the applied force (1.6 pN in Figure 2B), making any hopping involving a transient disengagement practically impossible. An alternative mechanism of bypass would be for the motor to ‘step over' the adduct (see Discussion for detailed definitions of protein step sizes). Given that HTMP crosslinking of thymine residues across a dTA dinucleotide produces a bulky major groove adduct, a protein would most likely need a step size of 3 bp or more to reliably overcome the damage. Since we present evidence in the discussion that EcoR124I steps along DNA in increments of 2 bp or less, we also disfavour this mechanism of bypass.
Is there a strand polarity of DNA translocation by Type I RM enzymes?
From the data above, strand separation during translocation is highly unlikely and EcoR124I can be classified as a true dsDNA translocase. But how do the HsdRs contact DNA? For the orthodox helicases, the binding and hydrolysis of NTPs has been suggested to drive the opening and closing of a cleft between RecA-like motor domains, which in turn drives motion of ssDNA in one of two polarities—either 5′–3′ or 3′–5′—across the domains (the ‘inchworm mechanism'; Soultanas and Wigley, 2001). Thus, helicases can be defined as having a distinct strand polarity (Lohman and Bjornson, 1996). In a similar manner, a dsDNA motor could contact and directionally convey just one strand, the other strand being passively transported. Alternatively, a dsDNA motor could make contacts with both strands.
To investigate these alternatives, we introduced ssDNA gaps downstream of an EcoR124I site and measured the effects on DNA translocation using complementary single molecule and bulk solution approaches (Figure 1). For both sets of DNA substrates, the gaps were introduced by modifications of a published method (see Wang and Hays, 2001; Materials and methods; Supplementary Figure 3). One consequence of this method is that the relative locations of the gaps differ from substrate-to-substrate (Supplementary Tables 1–3). However, we show later by the analysis of polyethylene glycol (PEG) spacers that the relative location of an adduct does not affect the outcome (Figure 6A).
Figure 6.

Stopped flow measurements of DNA translocation over DNA adducts. Triplex displacement profiles were recorded using DNA substrates containing the modifications indicated (see Supplementary Table 3). Experimental conditions were as given in Figure 5. (A) ΔTlag values measured on DNA containing two C18 PEG spacers (replacing 6 nt of DNA) in one or other strand. The relative location of the spacers was varied from 601–611 bp downstream of the EcoR124I binding site. The dashed lines represent 〈ΔTlag〉 for each strand. (B) ΔTlag values measured on DNA containing the modifications indicated. All error bars in the figure represent the standard error of the mean calculated from ⩾2 independent experiments.
Translocation on gapped DNA measured using magnetic tweezers
EcoR124I motor events on DNA containing a 20 nt gap in one or other strand are shown in Figure 3A. A clear difference between the substrates emerged. (1) On the 3′–5′ modified DNA, only short-range motor events were observed (upper panel). The maximum distance translocated was 1210±20 bp, corresponding to the 1180 bp distance between the EcoR124I site and the gap, with the majority of events terminating almost exactly at this point (Supplementary Figure 4). In comparison, on unmodified DNA, a significant number of translocation events are observed beyond this distance (Supplementary Figure 4). The simplest explanation is that EcoR124I can translocate as far as the 3′–5′ gap but then cannot continue. (2) On the 5′–3′ modified DNA, a significant proportion of events also terminated upon reaching the gap (lower panel). However, in 22±6% of events bypass was observed, with termination occurring downstream of the gap (Supplementary Figure 4). This suggests that EcoR124I can only bypass a 20 nt gap when it is on the 5′–3′ strand. We note that DNA cleavage at the gaps was not observed for either substrate.
Figure 3.

Single molecule measurements of DNA translocation on gapped DNA. (A) Two time traces recorded in the magnetic tweezers at 20 nM MTase and 160 nM HsdR using the DNA substrate as shown with a 20 nt gap in one or other strand. EcoR124I translocation towards the bead is lost in the thermal noise so the only events observed directly are those towards the gap. The grey lines represent the approximate locations of the gaps. (B) Classification of different motor events as observed at the gap. Broken line represents location of the gap. (C) Statistics of different motors events on each strand as a function of gap length (see Supplementary Methods for details of statistical analysis). Bypass and stall+bypass events were scored together.
Four different events could be classified as occurring at a gap (Figure 3B): bypass without visible stalling (bypass); stalling followed by bypass (stall+bypass); stalling followed by dissociation (stalling); and dissociation without visible stalling (dissociation). The statistics of these events were calculated for gaps in each strand, and for variations in gap size (Figure 3C). Bypass and stall+bypass events were scored together. Bypass events were always more efficient when the gap was in the 5′–3′ strand, even when the modification was only a break in the phosphodiester backbone (top panel). This suggests that the principal motor contacts are made to the 3′–5′ strand. This conclusion is reinforced by the stalling probability, which was much greater with gaps on the 3′–5′ strand (middle panel)—a break in the translocated strand would necessarily stall a motor, whereas a break in the non-translocated strand could be overcome by translocation across the intact strand. In contrast, dissociation events were more likely when the gap was in the 5′–3′ strand (lower panel). This suggests that contacts to this strand are more important in stabilising a translocating complex. Given that even a dephosphorylated nick can have a significant effect on the translocation kinetics, it also seems likely that EcoR124I moves in small steps (see below and Discussion).
An alternative view of the data is that translocation along the 5′–3′ strand would also lead to an increase in dissociation at 5′–3′ gaps as the motor would dissociate from the end of the DNA. However, in most cases helicase motors tend to be stabilised at DNA ends (e.g., Dillingham et al, 2000), as there are still sufficient contacts between the DNA and the rear of the motor. Data on modified DNA presented below (Figure 6) supports the notion that the 3′–5′ strand plays a more important role than the 5′–3′ strand. Therefore, the overall conclusion we draw from our data is that EcoR124I translocates dsDNA principally by contacts to the 3′–5′ strand, with additional stability coming from contacts to the 5′–3′ strand. We have also obtained further support for 3′–5′ translocation by measuring the directionality of EcoR124I motion at DNA branch points (Louise K Stanley and Mark D Szczelkun, unpublished observations).
Is applied force critical to the probability of gap bypass?
A prerequisite of measuring translocation by a Type I enzyme in the tweezers assay is that the DNA substrate must be extended by the application of force parallel to the chain (1.6 pN in Figure 3A; Seidel et al, 2004). This is critical to our interpretation of the above data as applied force could effect the motor reaction itself; for EcoR124I, initiation rates and processivity are force dependent (Seidel et al, 2004, 2005). Additionally, applied force could alter the conformational dynamics of the DNA (Blumberg et al, 2005; Sankararaman and Marko, 2005; Yan et al, 2005), which might influence the efficiency of gap bridging.
To investigate if applied tension affected gap bypass, we re-examined EcoR124I translocation on the 20 nt 3′–5′ gap substrate at a reduced force (F=0.4 pN, Figure 4A). In contrast to the complete inhibition at 1.6 pN (Figure 3A), gap bypass was now observed in ∼12% of events (Figure 4B). Therefore, DNA tension seems to inhibit gap bypass. How these observations tally with possible translocation mechanisms is discussed below.
Figure 4.

Force dependence of gap bypass on the 3′–5′ strand. (A) Time trace recorded in the magnetic tweezers at 0.4 pN using the DNA substrate as shown in Figure 3A with a 20 nt gap in the 3′–5′ strand. Enzyme concentrations were 20 nM MTase and 80 nM HsdR. EcoR124I translocation towards the bead is lost in the thermal noise so the only events observed directly are those towards the gap. The grey line represents the estimated location of the gap (1190±40 bp, Supplementary Figure 4). Note that compared to Figure 3A, at the lower force there is an increase in noise and a decrease in apparent distances due to the reduced DNA stretching. (Inset) Detail from the full time trace (dotted box) showing stall events at the gap region and one bypass event. (B) Statistics of 3′–5′gap bypass over 20 nt at different forces. The 1.6 pN data is taken from Figure 3C. Calculation of statistics was carried out as in Figure 3. At least some of the reduction in Pstall at 0.4 pN comes from the increased probability of bypass. There is also an increase in Pdissociation, which might reflect complexes that terminate during bypass of 20 nt ssDNA loops.
Translocation on gapped DNA measured using triplex displacement
To provide supporting data for the interpretations of the single molecule experiments, we also measured translocation on gapped DNA using triplex displacement (Figure 1B; Firman and Szczelkun, 2000; McClelland et al, 2005; Seidel et al, 2005). We generated 5.6 kbp linear DNA substrates containing an EcoR124I site, a gap site 0.6 kbp downstream, followed by a triplex binding site a further 0.6 kbp downstream (Figure 5A; Supplementary Figure 3). Displacement of a fluorescent triplex by a translocating enzyme can be measured and is preceded by a lag phase, the size of which reflects initiation and translocation events (McClelland et al, 2005).
Figure 5.

Stopped flow measurements of DNA translocation on gapped DNA. (A) Triplex displacement profiles from the stopped flow experiments. The final solution contains 1 nM DNA (0.5 nM triplex), 30 nM MTase, 300 nM HsdR, and 25 μM ATP. (B) Changes in the triplex displacement profiles with different gap sizes, with ΔTlag=Tlagmodified−Tlagunmodified, where 〈Tlagunmodified〉=16.28±0.14 s (the average of Tlag from intact and nicked DNA, which were identical within error, data not shown). A gap size of zero represents a dephosphorylated nick. Due to the method used to make the gaps (Materials and methods, Supplementary Figure 3 and Table 2), all spacings shown are dephosphorylated except for the 30 nt spacing. To within experimental error, phosphorylated gaps gave identical results (data not shown). (C) Comparison of DNA translocation and gap cleavage using a substrate with a 150 nt 3′–5′ gap. A triplex displacement profile from a hand-mixed radioactive triplex assay (5 nM DNA (2.5 nM triplex), 30 nM MTase and 300 nM HsdR) is compared to the appearance of product DNA cleaved at a gap from an agarose gel assay using 3H-labelled DNA (5 nM DNA, 30 MTase and 90 nM HsdR). Similar traces were obtained for the 5′–3′ gap and the 300 nt spacings (data not shown). (D) ΔTlag values calculated for long gaps (all phosphorylated). All error bars in the figure represent the standard error of the mean calculated from ⩾2 independent experiments.
Complete triplex displacement was observed on all substrates, irrespective of gap size or strand polarity (e.g., 30 and 5 nt gaps in Figure 5A). We did not observe irreversible inhibition at longer 3′–5′ spacings as in Figure 3A. Since Pbypass is critically sensitive to DNA tension (Figure 4), the higher efficiency in the triplex assay most likely reflects the conformation of the DNA under the zero force regime in bulk solution (also, see Discussion). Nonetheless, we observed in nearly every case that the modifications altered the lag phase preceding triplex displacement. Unlike our most recent study (Seidel et al, 2005), we did not use full kinetic models to extract translocation rate constants as simulations showed that, due to model complexity, indistinguishable profiles were returned using completely different schemes. Instead, we estimated the lag time (Tlag) for each substrate using an established exponential fit method (Firman and Szczelkun, 2000; McClelland et al, 2005) and plotted the difference in Tlag to the unmodified substrate as ΔTlag=Tlagmodified−Tlagunmodified (Figure 5B).
While very little difference was observed between intact and nicked DNA (ΔTlag≈0), all other modifications, including a single missing phosphate, showed positive ΔTlag values (Figure 5B). This reflects slower translocation events at the gap. Note, however, that since dissociation does not alter Tlagmodified (McClelland et al, 2005), changes in Pdissociation will not alter ΔTlag. If stalling is more frequent at 3′–5′ gaps, as suggested by the tweezers measurements (Figure 3C), then ΔTlag3′–5′ should increase relative to ΔTlag5′–3′. Indeed, at short spacings (<10 nt), ΔTlag3′–5′ was four- to five-fold higher than ΔTlag5′–3′, consistent with the single-molecule experiments. However, as the spacings were increased (⩾10 nt), ΔTlag3′–5′ decreased, until at the 30 nt there was little difference between the strands. This trend appears to be the opposite of that expected given the data in Figure 3. While ΔTlag3′–5′ varied with gap size, ΔTlag5′–3′ was largely invariant (〈ΔTlag5′–3′〉=∼0.5 s). Note that although the relative positions of the gaps are not identical (Supplementary Table 2), there is no evidence for position-dependent effects with EcoR124I (Figure 6A). We could also show that the kinetics were unaffected by 5′ phosphorylation of the gap (data not shown). A small amount of gap cleavage was observed (<10% after 1 h incubation), independent of strand polarity or gap length (data not shown).
Bypass of long gaps in preference to short gaps seemed surprising, so we lengthened the gaps by exonuclease III treatment (Materials and methods). This produces gaps that are distributive in size and represent a range rather than an absolute length. Complete triplex displacement was still observed with ssDNA gaps in either strand of ∼150 and ∼300 nt, although the kinetics were considerably slower (Figure 5C and D, other data not shown). There was a small, strand-dependent difference in ΔTlag at 150 nt, but not at 300 nt. Significant cleavage at the gaps was also observed, but on timescales significantly slower than translocation (Figure 5C); in other words, bypass of a large gap could occur multiple times before it were cleaved. We suggest that EcoR124I remains at the gap for long periods of time during which slow cleavage could occur (Janscak et al, 1999). This might represent long-lived stall states or slow ssDNA translocation.
Overall, the triplex data confirm increased stalling (or hindered translocation) at the 3′–5′ gap relative to the 5′–3′ gap and support the proposition above that the principal EcoR124I motor contacts are made with the 3′–5′ strand (although both strands are clearly important). Also, in line with the results above, a dephosphorylated nick produced significant increases in ΔTlag, indicative of a small step size. However, it seems that EcoR124I can, under conditions of zero force, translocate over considerable gaps in either strand.
What nucleotide contacts are made during DNA translocation by Type I RM enzymes?
For the classical monomeric helicases, there are two possible modes of binding ssDNA: for SF1 enzymes the principal contacts come from DNA bases, which flip into binding pockets where hydrophobic stacking interactions are made with aromatic amino acids (the ‘Mexican Wave' mechanism; Soultanas and Wigley, 2001), although electrostatic interactions with the phosphodiester backbone are not completely superfluous (Eoff et al, 2005); in contrast, for SF2 enzymes the principal contacts appear to be with the backbone (Kim et al, 1998; Kawaoka et al, 2004). It has been suggested that backbone tracking has allowed adaptation to translocation on either ss- or dsDNA (Kawaoka et al, 2004). A recently determined structure of the catalytic core of a Rad54-homologue bound to DNA has revealed an even more complex picture for a dsDNA translocase (Dürr et al, 2005), with evidence for base and backbone contacts in both strands.
Here, we used triplex displacement to measure the effect of backbone and base adducts on EcoR124I translocation. This was achieved by rescuing our gapped substrates with synthetic oligonucleotides containing different adducts (see Materials and methods, Supplementary Table 3). We first replaced a 6 nt stretch with an equivalent length of PEG spacer (thus removing completely the nucleic acid contacts), with the position of the spacer relative to the EcoR124I site varied within an 11 bp window (Figure 6A). As observed with gapped DNA, modifications in either polarity were overcome, with those on the 3′–5′ strand slowing translocation to a greater degree than those on the 5′–3′ strand (∣ΔTlag3′–5′∣〉∣ΔTlag5′–3′∣). Within experimental error, the relative location of the spacer did not significantly alter the strand-dependent differences in ΔTlag; bypass of 3′–5′ adducts never became more efficient than 5′–3′ adducts. At first, it may seem contradictory that a motor can bypass a region of the translocated strand devoid of any nucleotide contacts. However, we believe that because of the conformational flexibility of the PEG spacer, bypass of this adduct occurs in a similar manner as for gapped DNA (see Discussion).
We next investigated stretches of stable abasic DNA (1 or 5 nt of tetrahydrofuran), reversed backbone polarity (5 nt of 3′–5′ linkage within the 5′–3′ chain, or vice versa) or peptide nucleic acid (5 nt of PNA). For each adduct, the B-form DNA conformation is also likely to be altered (Tackett et al, 2001; Eoff et al, 2005). Nonetheless, none of the substrates irreversibly blocked EcoR124I translocation (data not shown). In every case, ΔTlag3′–5′ was greater than ΔTlag5′–3′, supporting strongly a defined translocation strand polarity (Figure 6B). The ΔTlag3′–5′ values obtained with these substrates are somewhat higher than those obtained on the DNA with short gaps, although not as high as those obtained on long gaps (Figure 5). The range of ΔTlag values obtained most likely reflects underlying differences in the mechanisms of bypass. In contrast to some other helicase motors (e.g., see Kawaoka et al, 2004), we did not observe a striking difference in kinetics between sugar-phosphate backbone and base analogues. Where ΔTlag3′–5′ did increase noticeably with the PNA modification (Figure 6B), ΔTlag5′–3′ also increased proportionally, suggesting that this adduct has a greater influence on overall DNA structure (Tackett et al, 2001). It therefore appears that, as suggested for Rad54 (Dürr et al, 2005), EcoR124I must utilise contacts to the backbone and bases of both strands during translocation.
Discussion
When is a helicase not a helicase?
We have investigated the motor activity of an SF2 helicase, the Type I RM enzyme EcoR124I, using both single molecule and bulk solution measurements. Overall we find excellent consistency between our approaches. Where we have observed incongruity, this can be explained in terms of quantifiable technical differences (e.g., the application of force in the tweezers setup). In fact, the discrepancies have highlighted important mechanistic features that would otherwise have been overlooked, as discussed below. Key features of the EcoR124I translocation mechanism can be highlighted and incorporated into a view of dsDNA motor activity (Figure 7A):
Figure 7.

Model for strand-specific dsDNA translocation by a Type I RM motor. (A) Schematic of the motor contacts and DNA transitions. As a function of ATP binding and hydrolysis, downstream dsDNA is pumped via the 3′–5′ strand into the expanding loop. Secondary clamp contacts to the 5′–3′ strand from the HsdR subunit and/or other parts of the enzyme complex stabilise the dsDNA translocation and maintain processivity. DNA strand separation is not necessary during steady-state translocation but may play a role in initiation. The DNA is twisted as it passes the motor, 360° for every ∼11 bp translocated (Seidel et al, 2004). Accordingly, DNA in the expanding loop is undertwisted and becomes negatively supercoiled. (B) Schematic mechanisms for gap bypass by ssDNA looping (upper panel) or ssDNA translocation (lower panel). See main text for more details.
Translocation of intact dsDNA leads to changes in DNA twist. The HsdR motor domain interacts tightly with dsDNA and, in an ATP-dependent reaction, pumps DNA downstream of the complex into an expanding DNA loop (Figure 7A). Although there is evidence that non-duplex DNA is formed transiently in the initial expanding loop to compensate for induced strain (van Noort et al, 2004), ICLs do not inhibit translocation during the subsequent loop expansion (Figure 2; Supplementary Figure 1). Therefore, despite the presence of characteristic helicase motifs in HsdR, EcoR124I is very unlikely to be an obligate unwindase.
Another important feature of the model in Figure 7A is that the motor follows the helical twist of the DNA (Seidel et al, 2004). As the HsdR unit is fixed to the MTase, this alters the topology of the up- and downstream DNA (Figure 7A; Halford et al, 2004). Every motor that moves on DNA will do so in incremental steps along the polynucleotide; this is defined as the motor step size (d bp). If the stepping motion also results in DNA twisting, then we can also define a twisting step size (Halford et al, 2004); this is the number of base pairs tracked before the DNA is twisted by 360° (StepTw bp). For EcoR124I, StepTw has been measured as ∼11 bp (Seidel et al, 2004). For a protein to rotate the DNA 360° in 11 bp, it must track the DNA helix, which in turn can only occur if the motor has a relatively small translocation step size (d≪11 bp). We suggest here for EcoR124I that d is only 1–2 bp. This is consistent with our observations that loss of even a single phosphate or base can have an appreciable effect on translocation (Figures 3C and 5B). Moreover, our kinetics were insensitive to incremental changes in the relative position of a PEG spacer (Figure 6A); a larger step size might have produced a more conspicuous periodic relationship (Bianco and Kowalczykowski, 2000). Further support for a small d value comes from noise analysis of single molecule traces (Ralf Seidel, Joost GP Bloom, Cees Dekker and Mark D Szczelkun, manuscript in preparation).
Principal motor contacts occur via the 3′–5′ strand. For every substrate examined here, discontinuities in and modifications on the 3′–5′ strand had a more significant effect on translocation than those on the 5′–3′ strand (Figures 3, 5 and 6). The simplest explanation of our data is that HsdR steps along dsDNA via contacts to the 3′–5′ strand (Figure 7A). Accordingly, 3′–5′ gaps are more likely to result in stalling (Figure 3C), as the leading edge of the motor disconnects from the tracked strand (Figure 7B, upper panel). Stalling will be rarer at 5′–3′ gaps as the motor can continue unimpeded on the intact 3′–5′ track (Figure 7B, lower panel). This model necessitates that EcoR124I can translocate both ds- and ssDNA. If ssDNA translocation were >10-fold slower than dsDNA translocation (∼580 bp/s at 25°C; Seidel et al, 2004), then this would have been resolved in the tweezers assay as a reduced translocation rate during gap bypass and in the triplex assay as distance-dependent changes in ΔTlag5′–3′. That neither were observed suggests that translocation is about as fast over short ssDNA stretches as on dsDNA. The significantly slower ΔTlag5′–3′ values on longer gaps (Figure 5D) could be due to a slower translocation rate over long distances, but could also be due to conformational anisotropy in the track (e.g., hairpin structures).
Supplementary ‘Clamp' contacts occur via the 5′–3′ strand. We observed that while stalling events at 5′–3′ gaps were infrequent, the probability of dissociation was elevated (Figure 3C). We interpret these data as showing that EcoR124I makes ‘clamp' contacts to the 5′–3′ strand that do not have a direct role in DNA transport, but which ensure that the motor remains in tight contact with the tracked 3′–5′ strand (Figure 7A). Accordingly, during bypass of a 5′–3′ gap via ssDNA translocation of the 3′–5′ strand, the loss of available clamp contacts will increase the overall dissociation rate (Figure 7B, lower panel). Modifications to the 5′–3′ strand also had a lesser effect on translocation bypass than those on the tracked strand (Figure 6).
Contacts to both backbone and bases play a role in translocation. We observed that both backbone and base modifications affected translocation, with the greatest effects seen on the 3′–5′ strand (Figure 6). We therefore suggest that, like other related dsDNA motors (Dürr et al, 2005), EcoR124I utilises the entire dsDNA structure for its motor activity.
A model for the bypass of gapped DNA via ssDNA looping
For some DNA motors, translocation can be completely blocked by discontinuities in the tracked strand (Bertram et al, 2002; Whitehouse et al, 2003; Kawaoka et al, 2004). For others, as here, modifications can be overcome relatively efficiently (Amaratunga and Lohman, 1993; Bianco and Kowalczykowski, 2000; Tackett et al, 2001; Eoff et al, 2005). One explanation for bypass is that if a motor has a large step size, its leading protein domain can ‘step over' an adduct, grabbing the downstream DNA (Amaratunga and Lohman, 1993; Lohman and Bjornson, 1996). It has also been suggested that some helicases move on DNA using two coordinated but distinct step sizes; the quantum inchworm model of Bianco and Kowalczykowski (2000). In this mechanism, a leading domain makes a large forward step (≫1 bp) and stabilises the motor on the downstream dsDNA while a secondary domain unwinds the captured upstream DNA segment in smaller substeps (<5 bp). However, it is difficult to reconcile either of these bypass mechanisms with translocation by EcoR124I over gaps of ∼300 nt, particularly as the maximum step that this enzyme makes on DNA is ⩽2 bp.
Given that ssDNA gaps are intrinsically flexible (Mills et al, 1994, 1999; Rippe, 2001; Mills and Hagerman, 2004; Du et al, 2005), an alternative explanation for bypass is that during the lifetime of a stall event at a gap, looping of the ssDNA linker allows the broken ends of the translocated strand to juxtapose and form a pseudo-continuous track (Figure 7B, upper panel). As long as the protein complex can accommodate the ssDNA loop, a motor with a relatively small step size could bypass a large gap by stepping across a distance similar to a nick or relatively short gap. It has been suggested that RNA polymerases, motors that explicitly make steps of only 1 nt, can bypass gaps in their template strands in just such a manner (e.g., Zhou et al, 1995). Moreover, equivalent ssDNA looping events allows T4 DNA ligase to ligate across gaps without recourse to any protein stepping at all (e.g., Nilsson and Magnusson, 1982). We further suggest that DNA rearrangements can explain bypass of other flexible adducts; for example, by looping of both ssDNA and PEG.
In a looping model, bypass will be sensitive to linker flexibility, and efficiency will peak at a length which is a compromise between energetic suppression of bending at short distances and entropic suppression of looping at long distances (Rippe, 2001). The persistence length of ssDNA (pss) has been estimated as anywhere between 2–20 nt dependent upon conditions and nucleotide sequence (e.g., Mills et al, 1999; Murphy et al, 2004; Du et al, 2005). Therefore, the probability of end juxtaposition will almost certainly be reduced for gaps <10 nt, as in Figure 5B, due to energetically unfavourable ssDNA bending. We also observed that small changes in the backbone or bases had significant effects (Figures 3 and 5). The DNA around a de-phosphorylated nick is likely to have the same rigid properties as intact or nicked DNA (Mills et al, 1994, 1999; Mills and Hagerman, 2004), so EcoR124I may be lacking key interactions with the missing phosphate. Once a 1 nt gap is generated, the ssDNA is more flexible (Mills et al, 1994, 1999; Mills and Hagerman, 2004), but may be anisotropically bent (Guo and Tullius, 2003). This could reduce bypass efficiency in addition to bend stiffness. For gaps ≫30 nt, the probability of end juxtaposition should decrease, as in Figure 5D, due to the lower relative concentration of the ends (although hairpin formation might also play a compensatory role by shortening the apparent end-to-end length of the ssDNA linker).
If increasing gap length above pss increases flexibility, why does Pbypass3′–5′ decrease in the tweezers experiments? The critical factor is the DNA tension necessary for single molecule measurements (1.6 pN in Figure 3). At this force, looping will be suppressed at least 1000-fold, as estimated from calculations on dsDNA (Yan et al, 2005). As the ssDNA loop geometry may be critical to allow EcoR124I to re-engage the leading DNA (e.g., the dsDNA ends might need to approach to within ∼0.4 nm), force could suppress gap bypass to an even greater degree (Blumberg et al, 2005; Sankararaman and Marko, 2005; Yan et al, 2005). The force dependence observed here (Figure 4), coupled with an ability to efficiently bypass very large gaps at zero force (Figure 5C), are simply not consistent with a model in which EcoR124I makes large steps over supposedly straight ssDNA regions (see also Supplementary Section). Clearly, EcoR124I is sufficiently stable during translocation (Seidel et al, 2004), which potentially slow and complex DNA rearrangements can occur. Our work also highlights a very important consideration when analysing DNA or RNA motors—where single molecule and bulk solutions studies disagree, the disparity may reflect real differences (i.e., only one of the studies is actually correct) or a conditional difference (i.e., measurement conditions, enzyme preparations, etc.) If the measurements require the application of tension, then the consequences of the applied force on the motor kinetics must be carefully considered both in terms of the effects on the enzyme and on the polynucleotide.
A clear result from our analysis of EcoR124I is that dsDNA translocation requires close contacts with both strands of intact duplex DNA (Figure 7). This interpretation is obviously different from that expected for a classical DNA helicase, which will translocate along just one single strand (Lohman and Bjornson, 1996; Soultanas and Wigley, 2001). Instead EcoR124I can be said to be a prototypical member of a new class of dsDNA translocases, which also includes enzymes involved in chromatin remodelling (Whitehouse et al, 2003; Dürr et al, 2005; Lia et al, 2006) and DNA recombination (Singleton et al, 2001). The relationship to orthodox helicases is simply the presence of ATP binding motifs that couple nucleotide binding energy to protein domain motions (Singleton and Wigley, 2002). It is clear that the current tendency to classify helicases as ‘unwindases until proven otherwise' is unsatisfactory, a new motor taxonomy is required with a more transparent nomenclature.
Materials and methods
Proteins
EcoR124I was purified, reconstituted and tested as previously described (Seidel et al, 2005). BbvCI and its nicking mutants were supplied by Geoff Wilson (New England Biolabs). E. coli RecBCD was supplied by Mark Dillingham (University of Bristol).
DNA constructs
Gapped and modified DNA were generated from suitable plasmid substrates using modifications of the methods described by Wang and Hays (2001). For a full description of these DNA constructs, as well as those used for the crosslinking studies, please refer to the Supplementary data.
DNA crosslinking
HTMP (Sigma) was dissolved in absolute ethanol to a final concentration of 710 μg/ml. DNA substrates (18–26 μM dT equivalents) were incubated with either <0.033 HTMP/dT or >1 HTMP/dT (as indicated in the figure legends) in 10 mM Tris–Cl, 20 mM NaCl, 1 mM EDTA (pH 8.0), and then irradiated at 4°C with 0–76 kJ m−2 of 365 nm light. Irradiation conditions were chosen dependent on the HTMP concentration and the number of crosslinks required, and were determined empirically (Supplementary Methods). Irradiated samples were dialysed extensively against ddH2O using nitrocellulose membranes (0.025 μm, Millipore).
Magnetic tweezers measurements
Magnetic tweezers experiments were carried out as described previously (Seidel et al, 2004, 2005) in reaction Buffer R (50 mM Tris–HCl pH 8.0, 10 mM MgCl2, 1 mM dithiothreitol) supplemented with 4 mM ATP. All measurements were performed at 25°C.
Triplex displacement measurements
Triplex displacement measurements were carried out at 20°C as described (Firman and Szczelkun, 2000; McClelland et al, 2005; Seidel et al, 2005) with the final reactions conditions being either 1 nM DNA (0.5 nM tetramethylrhodamine triplex) for the stopped flow assays (McClelland et al, 2005) or 5 nM DNA (2.5 nM 32P-labelled triplex) for the gel assays (Firman and Szczelkun, 2000), and, in both cases, 30 nM MTase, 300 nM HsdR, 1.5 U/ml creatine phosphokinase, 14.5 mM phosphocreatine, 100 μM S-adenosyl methionine and 25 μM ATP in Buffer R.
DNA unwinding and cleavage measurements
Conditions for DNA unwinding by RecBCD or DNA cleavage by EcoR124I are described in the Supplementary Methods.
Data analysis
Previously described methods were used for the analysis of triplex displacement profiles (McClelland et al, 2005) and time traces from the magnetic tweezers (Seidel et al, 2004, 2005). Bypass, stalling and dissociation probabilities were calculated as described in Supplementary Methods.
Supplementary Material
Supplementary Methods
Acknowledgments
We thank G Wilson, M Dillingham and P Janscak for supplying materials. This work was supported by grants from the Wellcome Trust (071432 and 067439, MDS), the European Commission through the Mol Switch project (IST-2001-38036, CD), the Nederlandse Organisatie voor Wetenschappelijk Onderzoek (NWO, CD) and the Dutch Foundation for Fundamental Research on Matter (FOM, NHD).
References
- Amaratunga M, Lohman TM (1993) Escherichia coli rep helicase unwinds DNA by an active mechanism. Biochemistry 32: 6815–6820 [DOI] [PubMed] [Google Scholar]
- Bertram RD, Hayes CJ, Soultanas P (2002) Vinylphosphonate internucleotide linkages inhibit the activity of PcrA DNA helicase. Biochemistry 41: 7725–7731 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bianco PR, Kowalczykowski SC (2000) Translocation step size and mechanism of the RecBC DNA helicase. Nature 405: 368–372 [DOI] [PubMed] [Google Scholar]
- Blumberg S, Tkachenko AV, Meiners JC (2005) Disruption of protein-mediated DNA looping by tension in the substrate DNA. Biophys J 88: 1692–1701 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dillingham MS, Wigley DB, Webb MR (2000) Demonstration of unidirectional single-stranded DNA translocation by PcrA helicase: measurement of step size and translocation speed. Biochemistry 39: 205–212 [DOI] [PubMed] [Google Scholar]
- Du Q, Vologodskaia M, Kuhn H, Frank-Kamenetskii M, Vologodskii A (2005) Gapped DNA and cyclization of short DNA fragments. Biophys J 88: 4137–4145 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dürr H, Körner C, Müller M, Hickmann V, Hopfner KP (2005) X-ray structures of the Sulfolobus solfataricus SWI2/SNF2 ATPase core and its complex with DNA. Cell 121: 363–373 [DOI] [PubMed] [Google Scholar]
- Endlich B, Linn S (1985) The DNA restriction endonuclease of Escherichia coli B. I. Studies of the DNA translocation and the ATPase activities. J Biol Chem 260: 5720–5728 [PubMed] [Google Scholar]
- Eoff RL, Spurling TL, Raney KD (2005) Chemically modified DNA substrates implicate the importance of electrostatic interactions for DNA unwinding by Dda helicase. Biochemistry 44: 666–674 [DOI] [PubMed] [Google Scholar]
- Firman K, Szczelkun MD (2000) Measuring motion on DNA by the type I restriction endonuclease EcoR124I using triplex displacement. EMBO J 19: 2094–2102 [DOI] [PMC free article] [PubMed] [Google Scholar]
- George H, Kuraoka I, Nauman DA, Kobertz WR, Wood RD, West SC (2000) RuvAB-mediated branch migration does not involve extensive DNA opening within the RuvB hexamer. Curr Biol 10: 103–106 [DOI] [PubMed] [Google Scholar]
- Guo H, Tullius TD (2003) Gapped DNA is anisotropically bent. Proc Natl Acad Sci USA 100: 3743–3747 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Halford SE, Welsh AJ, Szczelkun MD (2004) Enzyme-mediated DNA looping. Annu Rev Biophys Biomol Struct 33: 1–24 [DOI] [PubMed] [Google Scholar]
- Hearst JE (1981) Psoralen photochemistry. Annu Rev Biophys Bioeng 10: 69–86 [DOI] [PubMed] [Google Scholar]
- Janscak P, MacWilliams MP, Sandmeier U, Nagaraja V, Bickle TA (1999) DNA translocation blockage, a general mechanism of cleavage site selection by type I restriction enzymes. EMBO J 18: 2638–2647 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Karu AE, Linn S (1972) Uncoupling of the recBC ATPase from DNase by DNA crosslinked with psoralen. Proc Natl Acad Sci USA 69: 2855–2859 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawaoka J, Jankowsky E, Pyle AM (2004) Backbone tracking by the SF2 helicase NPH-II. Nat Struct Mol Biol 11: 526–530 [DOI] [PubMed] [Google Scholar]
- Kim JL, Morgenstern KA, Griffith JP, Dwyer MD, Thomson JA, Murcko MA, Lin C, Caron PR (1998) Hepatitis C virus NS3 RNA helicase domain with a bound oligonucleotide: the crystal structure provides insights into the mode of unwinding. Structure 6: 89–100 [DOI] [PubMed] [Google Scholar]
- Kuhn B, Abdel-Monem M, Hoffmann-Berling H (1979) DNA helicases. Cold Spring Harb Symp Quant Biol 43: 63–67 [DOI] [PubMed] [Google Scholar]
- Lia G, Praly E, Ferreira H, Stockdale C, Tse-Dinh YC, Dunlap D, Croquette V, Bensimon D, Owen-Hughes T (2006) Direct observation of DNA distortion by the RSC complex. Mol Cell 21: 417–425 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lohman TM, Bjornson KP (1996) Mechanisms of helicase-catalyzed DNA unwinding. Annu Rev Biochem 65: 169–214 [DOI] [PubMed] [Google Scholar]
- Makovets S, Powell LM, Titheradge AJ, Blakely GW, Murray NE (2004) Is modification sufficient to protect a bacterial chromosome from a resident restriction endonuclease? Mol Microbiol 51: 135–147 [DOI] [PubMed] [Google Scholar]
- McClelland SE, Dryden DTF, Szczelkun MD (2005) Continuous assays for DNA translocation using fluorescent triplex dissociation: application to type I restriction endonucleases. J Mol Biol 348: 895–915 [DOI] [PubMed] [Google Scholar]
- McClelland SE, Szczelkun MD (2004) The type I and III Restriction endonucleases: structural elements in molecular motors that process DNA. In Nucleic Acids and Molecular Biology—Restriction Enzymes, Pingound A (ed), Vol. 14, pp 111–135. Germany: Springer Verlag [Google Scholar]
- Mills JB, Hagerman PJ (2004) Origin of the intrinsic rigidity of DNA. Nucleic Acids Res 32: 4055–4059 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mills JB, Cooper JP, Hagerman PJ (1994) Electrophoretic evidence that single-stranded regions of one or more nucleotides dramatically increase the flexibility of DNA. Biochemistry 33: 1797–1803 [DOI] [PubMed] [Google Scholar]
- Mills JB, Vacano E, Hagerman PJ (1999) Flexibility of single-stranded DNA: use of gapped duplex helices to determine the persistence lengths of poly(dT) and poly(dA). J Mol Biol 285: 245–257 [DOI] [PubMed] [Google Scholar]
- Murphy MC, Rasnik I, Cheng W, Lohman TM, Ha T (2004) Probing single-stranded DNA conformational flexibility using fluorescence spectroscopy. Biophys J 86: 2530–2537 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nilsson SV, Magnusson G (1982) Sealing of gaps in duplex DNA by T4 DNA ligase. Nucleic Acids Res 10: 1425–1437 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rippe K (2001) Making contacts on a nucleic acid polymer. Trends Biochem Sci 26: 733–740 [DOI] [PubMed] [Google Scholar]
- Sankararaman S, Marko JF (2005) Formation of loops in DNA under tension. Phys Rev E Stat Nonlinear Soft Matter Phys 71: 021911. [DOI] [PubMed] [Google Scholar]
- Seidel R, Bloom JG, van Noort J, Dutta CF, Dekker NH, Firman K, Szczelkun MD, Dekker C (2005) Dynamics of initiation, termination and reinitiation of DNA translocation by the motor protein EcoR124I. EMBO J 24: 4188–4197 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Seidel R, van Noort J, van der Scheer C, Bloom JG, Dekker NH, Dutta CF, Blundell A, Robinson T, Firman K, Dekker C (2004) Real-time observation of DNA translocation by the type I restriction modification enzyme EcoR124I. Nat Struct Mol Biol 11: 838–843 [DOI] [PubMed] [Google Scholar]
- Singleton MR, Scaife S, Wigley DB (2001) Structural analysis of DNA replication fork reversal by RecG. Cell 107: 79–89 [DOI] [PubMed] [Google Scholar]
- Singleton MR, Wigley DB (2002) Modularity and specialization in superfamily 1 and 2 helicases. J Bacteriol 184: 1819–1826 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soultanas P, Wigley DB (2001) Unwinding the ‘Gordian knot' of helicase action. Trends Biochem Sci 26: 47–54 [DOI] [PubMed] [Google Scholar]
- Szczelkun MD, Dillingham MS, Janscak P, Firman K, Halford SE (1996) Repercussions of DNA tracking by the type IC restriction endonuclease EcoR124I on linear, circular and catenated substrates. EMBO J 15: 6335–6347 [PMC free article] [PubMed] [Google Scholar]
- Tackett AJ, Wei L, Cameron CE, Raney KD (2001) Unwinding of nucleic acids by HCV NS3 helicase is sensitive to the structure of the duplex. Nucleic Acids Res 29: 565–572 [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Noort J, van der Heijden T, Dutta CF, Firman K, Dekker C (2004) Initiation of translocation by Type I restriction-modification enzymes is associated with a short DNA extrusion. Nucleic Acids Res 32: 6540–6547 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang H, Hays JB (2001) Simple and rapid preparation of gapped plasmid DNA for incorporation of oligomers containing specific DNA lesions. Mol Biotechnol 19: 133–140 [DOI] [PubMed] [Google Scholar]
- Whitehouse I, Stockdale C, Flaus A, Szczelkun MD, Owen-Hughes T (2003) Evidence for DNA translocation by the ISWI chromatin-remodeling enzyme. Mol Cell Biol 23: 1935–1945 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yan J, Kawamura R, Marko J (2005) Statistics of loop formation along double helix DNAs. Phys Rev E Stat Nonlinear Soft Matter Phys 71: 061905. [DOI] [PubMed] [Google Scholar]
- Zhou W, Reines D, Doetsch PW (1995) T7 RNA polymerase bypass of large gaps on the template strand reveals a critical role of the nontemplate strand in elongation. Cell 82: 577–585 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Supplementary Methods


