Figure 5.
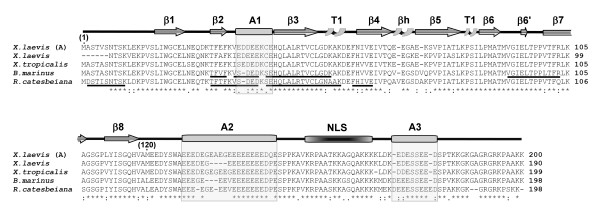
Protein sequence alignment of amphibian nucleoplasmins. The primary structures of nucleoplasmin from X. laevis (A) [60] [GenBank: X04766], X. laevis (Burglin et al., 1987) [GenBank: CAA68363], X. tropicalis [GenBank: NP_001016938], B. marinus and R. catesbeiana are shown. Identical amino acids are denoted by an asterisk, highly similar residues by a colon, and less similar residues by a period, as determined by CLUSTAL W software. The partial protein sequences of B. marinus and R. catesbeiana determined by mass spectroscopy peptide sequencing are underlined. The highly structured N-terminal protein core spans amino acids 1–120 and has β sheets (β1–8), two type 1 turns (T1) and a β hairpin (βh) [8]. The other boxes represent the A1, A2, A3 polyglutamic tracts and the bipartite nuclear localization signal (NLS), as indicated.
