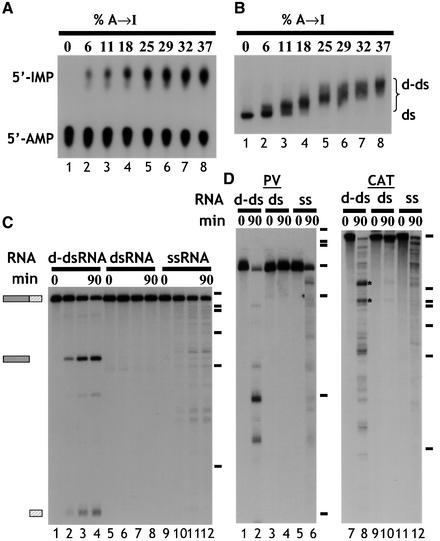
Fig. 1. (A) Quantitation of deamination. ΔKP dsRNA was deaminated to varying degrees using ADAR2. Digestion of the modified RNA using RNase P1, followed by TLC, enabled quantitation of the amount of A to I conversion. The RNAs used in the assays shown typically contained ∼40% A to I conversion (lane 8). (B) Deamination of ΔKP results in a homogenous population of RNA. As the level of deamination of ΔKP increased, there was a corresponding reduction in the mobility of the RNA on a native gel (lanes 1–8). For dsRNAs with ∼40% A to I conversion (lane 8), the RNA migrated as a relatively compact band (‘d-ds’), indicating that all RNAs in the population were modified. (C) Deaminated dsRNA is specifically cleaved. Incubation of ΔKP d-dsRNA in Xenopus oocyte extract gave rise to two discrete cleavage products (lanes 1–4). In contrast, ΔKP dsRNA was stable (lanes 5–8), and ΔKP ssRNA was degraded slowly yielding numerous products (lanes 9–12). The time course used in each assay was 0, 15, 60 and 90 min. Positions of DNA molecular weight markers are shown to the right of the figure (φX174 HaeIII, 310, 281, 271, 234, 194, 118, and 72 nt). (D) Incubation of both PV and CAT d-dsRNAs for 90 min in Xenopus oocyte extracts also gave rise to discrete cleavage products (lanes 1–6 and 7–12, respectively). The equivalent dsRNAs were stable (compare lanes 2 and 4, and lanes 8 and 10), while degradation of the ssRNAs gave more numerous products (lanes 6 and 12).

An official website of the United States government
Here's how you know
Official websites use .gov
A
.gov website belongs to an official
government organization in the United States.
Secure .gov websites use HTTPS
A lock (
) or https:// means you've safely
connected to the .gov website. Share sensitive
information only on official, secure websites.