Figure 9.
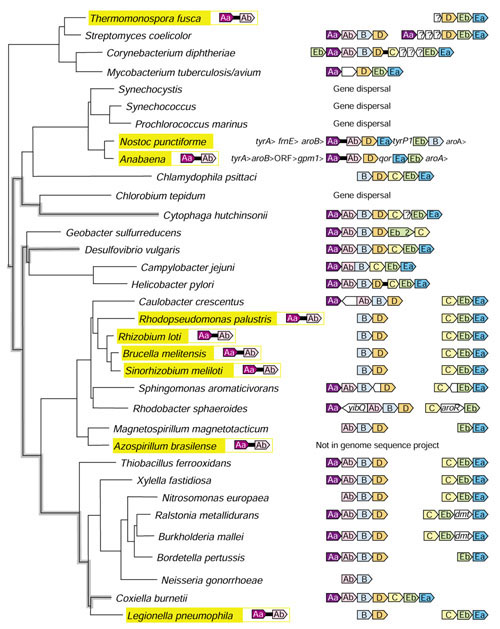
16S rRNA tree showing the phylogenetic distribution (highlighted in yellow) of trpAa•trpAb fusions. The gene fusions unlinked to any other trp genes are shown to the right of the highlighted name. The remaining trp-operon gene organizations are shown at the right. The white arrows indicate gene insertions that encode the following: Thermomonospora, integral membrane protein; Streptomyces, three membrane proteins: Corynebacterium, membrane protein, pantoate 3-alanine ligase (panC), and 3-methyl-2-oxobutanoate hydroxymethyl transferase (panB); Mycobacterium, conserved hypothetical protein; Cytophaga, conserved hypothetical protein; Sphingomonas, conserved hypothetical protein and outer-membrane protein; Rhodobacter, and acetyltransferase yibQ; Ralstonia, DNA methyltransferase (dmt); Burkholdaria, DNA methyltransferase (dmt). In addition aroR in R. sphaeroides is a putative regulatory gene [58]. The lineage relationships of three organisms that have maintained the putative ancestral trp operon are shown with heavy, gray lines.
