Abstract
An EST database has been generated for coffee based on sequences from approximately 47,000 cDNA clones derived from five different stages/tissues, with a special focus on developing seeds. When computationally assembled, these sequences correspond to 13,175 unigenes, which were analyzed with respect to functional annotation, expression profile and evolution. Compared with Arabidopsis, the coffee unigenes encode a higher proportion of proteins related to protein modification/turnover and metabolism—an observation that may explain the high diversity of metabolites found in coffee and related species. Several gene families were found to be either expanded or unique to coffee when compared with Arabidopsis. A high proportion of these families encode proteins assigned to functions related to disease resistance. Such families may have expanded and evolved rapidly under the intense pathogen pressure experienced by a tropical, perennial species like coffee. Finally, the coffee gene repertoire was compared with that of Arabidopsis and Solanaceous species (e.g. tomato). Unlike Arabidopsis, tomato has a nearly perfect gene-for-gene match with coffee. These results are consistent with the facts that coffee and tomato have a similar genome size, chromosome karyotype (tomato, n=12; coffee n=11) and chromosome architecture. Moreover, both belong to the Asterid I clade of dicot plant families. Thus, the biology of coffee (family Rubiacaeae) and tomato (family Solanaceae) may be united into one common network of shared discoveries, resources and information.
Keywords: Coffea canephora, Rubiaceae, Solanaceae, Seed development, Comparative genomics
Introduction
Coffee is an important international commodity, ranking among the five most valuable agricultural exports from developing countries (Food and Agriculture Organization, http://apps.fao.org). Moreover, production and processing of coffee employs more than 25 million people worldwide (O’Brien and Kinnaird 2003). Despite its economic importance, coffee has received little attention with respect to molecular genetics and genomics research. As of December 2004, only 1,570 nucleotide and 115 protein sequences from coffee had been deposited in GenBank with the majority of those sequences derived from leaf ESTs. Many of the remaining sequences correspond to enzymes in the caffeine biosynthesis pathway—the most extensively studied pathway in coffee (Moisyadi et al. 1998; Ogawa et al. 2001; Mizuno et al. 2003; Uefuji et al. 2003).
Commercial coffee production relies mainly on two closely related species: Coffea arabica and Coffea canephora, accounting for approximately 70 and 30% of worldwide coffee production, respectively (Herrera et al. 2002). Although C. canephora accounts for a lower total proportion of the coffee market than does C. arabica, it is the main source for soluble coffee, which is consumed widely throughout the world. C. canephora is a diploid (2n=2x=22), outcrossing and highly polymorphic species native to central Africa, but which has expanded, through cultivation, especially to western Africa, Indonesia and Vietnam (Wrigley 1988). In contrast, C. arabica is believed to be a recently derived tetraploid (2n=4x=44) native to a small region of what is now Ethiopia. C. arabica is now grown widely throughout the world.
The goal of the current project was to increase the genetic and molecular knowledge of coffee through the generation and annotation of an EST database using high throughput single-pass 5′ sequencing of cDNAs derived from leaf, pericarp and seed tissues from a set of C. canephora varieties. Special emphasis was given to sequencing cDNAs from different stages of seed development, both to shed light on this important, but not well understood aspect of plant development and to capture as many genes as possible involved in determining the final chemical composition of seeds which constitute the commercial product. As a result, the EST database reported herein is, to our knowledge, the largest public database of seed-derived ESTs (White et al. 2000; Suh et al. 2003).
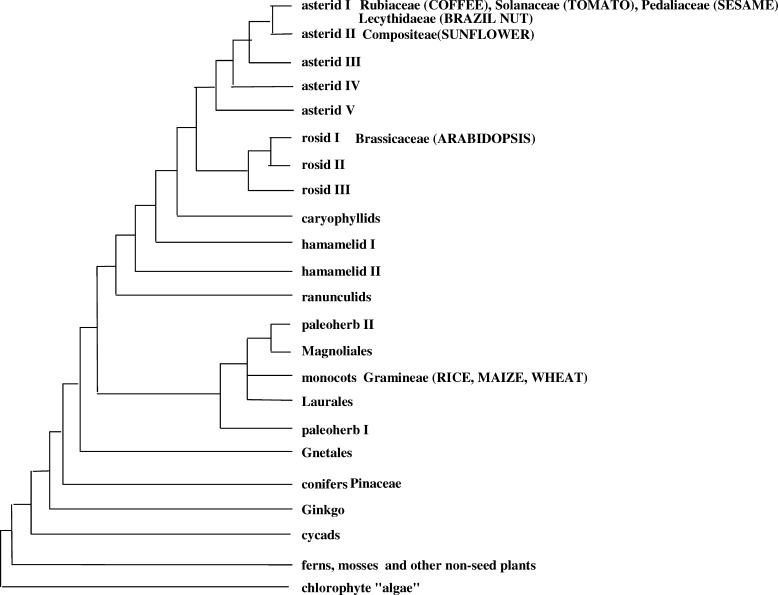
The EST database was used to derive a coffee unigene build, which was subsequently subjected to functional annotation. The coffee unigene build was compared with the model species Arabidopsis. Arabidopsis is the only fully sequenced dicot genome and hence provides a complete set of predicted genes against which to compare the EST-derived unigene set of coffee. However, coffee and Arabidopsis belong to different plant families (Rubiaceae and Brassicaceae, respectively) which are distantly related phylogenetically and which diverged from their last common ancestor approximately 94 million years ago (MYA) (Gandolfo et al. 1998; Crepet et al. 2004) (Fig. 1).
Fig. 1.
Dendrogram depicting phylogenetic relationships of coffee to other higher plant taxa (based on Chase et al. 1993)
The plant family most closely related to coffee in which extensive sequencing has been conducted is Solanaceae (Fig. 1). In this family, comprehensive EST databases have been developed for tomato, potato, pepper, eggplant and petunia (http://www.sgn.cornell.edu/) (Hoeven et al. 2002; Ronning and Stegalkina 2003; Lee et al. 2004). Both Rubiaceae and Solanaceae belong to the Asterid I clade of dicots, and based on existing fossil evidence, are thought to have diverged from one another approximately 50 MYA (Gandolfo et al. 1998; Crepet et al. 2004) (Crepet personal communication) (Fig. 1). The closer taxonomic affinities of coffee and Solanaceae (e.g. tomato) are paralleled by a number of striking botanic and genetic similarities, including the production of fleshy berries, a similar genome content (C=950 and 640 Mb for tomato and coffee, respectively) (Hoeven et al. 2002), similar basic chromosome number (x=12 for tomato and most other Solanaceae; x=11 for coffee) and similar chromosome architecture with highly condensed pericentric heterochromatin and decondensed euchromatin at the pachytene stage of meiosis (Rick 1971; Pinto-Maglio and Cruz 1998). For these reasons, the coffee unigene set was also compared against a series of Solanaceae EST-derived unigene sets.
Materials and methods
Library construction
Source of tissues
C. canephora was farm-grown in east Java. Plant tissue was frozen in liquid nitrogen, transported on dry ice and stored at –80°C until RNA extraction. The specified tissues (leaf, pericarp, young cherries with seed and pericarp mixed, middle stage seed and late stage seed) were collected from five different varieties Table 1). The maturation period of the C. canephora varieties under study is approximately 9–11 months, from pollination to ripening. The early stage cherry, middle stage seed and late stage seed were collected between 18 to 22 weeks, 30 weeks and 42 to 46 weeks after pollination, respectively.
Table 1.
Characteristics of the 5 cDNA libraries used to develop the coffee EST database
| Library name | Tissue | Varieties | Average insert size, kb | Good quality ESTs |
|---|---|---|---|---|
| Leaf | Leaves, young | BP409 | 1.5±0.6 | 8,942 |
| Pericarp | Pericarp, all developmental stages | BP358, BP409, BP42, BP961, Q121 | 1.4±0.5 | 8,956 |
| Early stage cherry | Whole cherries, 18 and 22 week after pollination | BP358, BP409, BP42, Q121 | 1.4±0.3 | 9,843 |
| Middle stage seed | Endosperm and perisperm of seeds, 30 week after pollination | BP409, BP961, Q121 | 1.4±0.3 | 10,077 |
| Late stage seed | Endosperm and perisperm of seeds, 42 and 46 week after pollination | BP358, BP409, BP42, BP961, Q121 | 1.4±0.3 | 9,096 |
RNA and mRNA isolation
Total RNA was extracted using phenol/chloroform (Rogers et al. 1999) and further treated with DNase I (RNase-free) and purified using an RNeasy Kit (Qiagen, Valencia, CA 91355). Messenger RNA was extracted from total RNA with PolyTrack mRNA Isolation Systems (Promega, Madison, WI 53711).
cDNA libraries
Directional cDNA libraries were constructed with 3–5 μg of mRNA with the ZAP-cDNA Gigapack III Gold Cloning Kit (Stratagene, La Jolla, CA 92037). The average insert length was estimated by PCR in 36 randomly selected clones from each library and ranged 1.2–1.5 kb (Table 1).
Sequencing
Bacteria, containing coffee cDNAs, were cultured in 384-well plates and cDNA inserts subjected to 5′ end sequencing at the BioResource Center at Cornell University (http://www.brc.cornell.edu). The average size of quality reads was 613 bp with a maximum of 1,037 bp.
Sequence quality processing
EST sequences were base-called and screened for vector sequences using PHRED software (Ewing et al. 1998). The longest stretch of overall high quality (PHRED score over 15 which corresponds to over 98% confidence) of each sequence was identified. PolyA repeats were trimmed to at most 20 bp and any sequence past the PolyA (mostly low quality sequence) was discarded. After the trimming, the sequences were screened against the E.coli K12 genome to remove any bacteria contamination. The remaining sequences were screened for minimum length (150 bp) and maximum allowed ambiguity (4%) and low complexity (60% of the sequence are of the same nucleotide, or 80% of same two nucleotides, which indicate error in sequencing).
Unigene assembly
Unigene sets were built by combining the sequences from all five coffee cDNA libraries. Clustering was performed using a program developed at the Sol Genomics Network (SGN at http://www.sgn.cornell.edu), which relied on a custom pre-clustering algorithm, and on the CAP3 program for contig generation (Huang and Madan 1999). The preclustering algorithm clustered sequences using a Smith Waterman type algorithm with initial word matching. The command line settings for CAP3 were as follows: -e 5000 -p 90 -d 10000 -b 60. The -e, -d and -b options are set such that the assembler disregards them or minimizes their effect. The -p option increases the sequence identity necessary on overlaps to 90 from a default of 75, which were found to be not stringent enough. Sequences were also checked for length, complexity and contamination. The builds were uploaded to the database, where each unigene was assigned a unique unigene ID.
Annotation
Protein prediction
The most likely coding frame and the corresponding transcribed peptide for each unigene were generated using ESTScan, a program that detects coding regions in EST sequences and corrects some sequencing errors (nucleotide substitution, deletion/insertion, erroneous stop codon, etc) (Iseli et al. 1999). As a hidden Markov Model (HMM) based program, ESTScan requires a training set of high quality coding sequences to generate a model. However, very few high quality coffee coding sequences are publicly available (less than 50 full length in GenBank). Therefore, we used as a training set of 483 nuclear genes from the closely related species, tomato (http://www.ebi.ac.uk/embl). Table 2 compares the coffee unigene assembly and translation (using ESTScan) with that of tomato, using the same training tomato set and parameters. The fact that the two give such similar results, gives confidence that the coffee EST-derived unigenes are being efficiently translated via the ESTScan software using the tomato training set (Table 2).
Table 2.
Comparison of the coffee and tomato EST databases derived from use of ESTScan calibrated with the same tomato training set (see Materials and methods for details)
| Tomato | Coffee | |
|---|---|---|
| Total unigenes | 30,576 | 13,175 |
| Average unigene length, bp | 774 | 678 |
| Unigenes with coding regions | 96% | 95% |
| Average length (bp) of predicated peptides | 569 | 556 |
| Average ESTScan score | 409 | 346 |
BLAST matches between coffee unigenes and other sequence databases
Sequence matches for coffee unigenes against other sequence databases were obtained using BLAST (Altschul et al. 1997) (see Results and discussion section for details of each BLAST analysis). The target databases were:
The GenBank non-redundant (NR) protein and dbest dataset (NCBI, http://www.ncbi.nlm.nih.gov).
The predicted Arabidopsis proteome and cDNA databases which were downloaded from the Arabidopsis Information Resource (TAIR, http://www.arabidopsis.org and ftp://ftp.arabidopsis.org/home/tair/).
The Solanaceae EST-derived unigene sets, including tomato (Solanum. lycopersicum) (184,860 ESTs, 30,576 unigenes), potato (Solanum. tuberosum) (97,425 ESTs, 24,932 unigenes), pepper (Capsicum annuum) (20,738 ESTs, 9,554 unigenes), petunia (Petunia hybrida) (3,181 ESTs, 1,841 unigenes) and eggplant (Solanum melongena) (11,479 ESTs, 5,135 unigenes), all of which can be accessed at SGN (http://www.sgn.cornell.edu).
To estimate ribosomal, chloroplast and mitochondrial contamination, the coffee unigene set was also screened against the Arabidopsis complete mitochondrial genome (NC_001284), tobacco ribosomal rRNA gene (5.8s AJ012367, 18s AJ236016 and 25s S52185) and the complete tobacco chloroplast genome (Z00044) sequences from GenBank.
Functional annotation based on predicted peptides
ESTScan-predicted coffee peptides were subjected to InterPro Scan annotation, which integrates the most commonly used protein signature databases (PROSITE, PRINTS, Pfam, ProDom, etc.) together with their associated scanning methods for protein domain analysis (Apweiler et al. 2001; Zdobnov and Apweiler 2001). Based on the domain annotation, GO accession of the unigenes were assigned using interpro2go conversion file from the GO consortium (http://www.geneontology.org, also available at http://www.ebi.ac.uk/interpro).
Functional categorization based on gene ontology
GO annotations were formatted for input into the GOSlim program and the output parsed to count the occurrence of each GO category. GOSlims are ‘slimmed down’ versions of the ontologies that allow a high-level view of gene functions. The GOSlim file and program were obtained from the Gene Ontology Consortium at http://www.geneontology.org.
Gene family analysis
The predicted protein sequences for the coffee unigene set and the Arabidopsis protein set were combined into a single file, formatted as a blast database using formatdb, and run with BLASTP (protein vs. protein sequence) against itself with option m8 for output. The resulting file was used as the input for the tribeML program. TribeML (Enright et al. 2002) formats the clusters, such that each cluster was given in a tab delimited file, one cluster per line. Simple scripts were used to parse the information to detect the largest gene families, coffee specific families, and families that showed large expansions in coffee.
Results and discussion
Generation of coffee EST database and unigene set
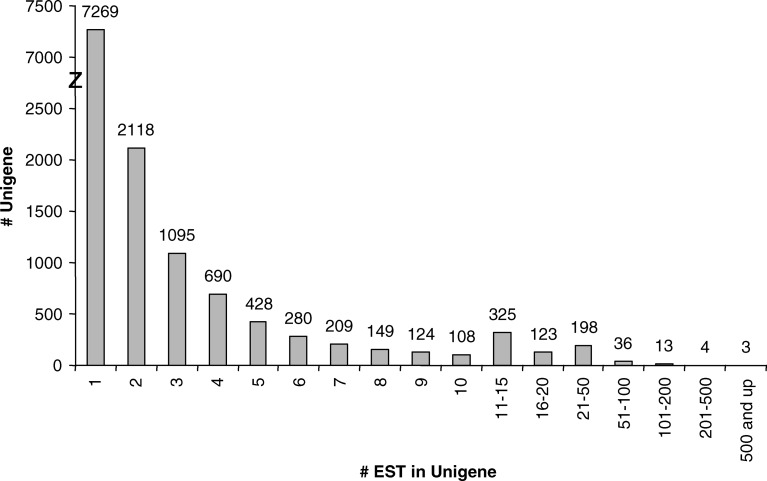
A total of 62,829 cDNA clones, derived from mRNA from five different tissues, were subjected to 5′ sequencing. After quality evaluation (see Materials and methods for details), the database was reduced to 46,914 high quality ESTs, averaging 613 bp in length. This corresponded to approximately 9,000 high quality ESTs from each of the five cDNA libraries (Table 1). These high quality ESTs were assembled into 13,175 unigenes with average length of 678 bp (ranging 150–2,714 bp), among which 7,272 (55%) were singletons and 5,903 (45%) were contigs (Fig. 2). Of the contigs, the majority (87%) was represented by two to ten ESTs while some (13%) were comprised of more than ten ESTs (Fig. 2). All trace files, sequences and derived unigenes/annotation can be found at the Solanaceae Genomics Network (SGN) website (http://www.sgn.cornell.edu) and will be submitted to GenBank at the time of publication.
Fig. 2.
Histogram depicting the distribution of EST content for all coffee unigenes. Numbers above bars equals the number of unigenes represented in each
Differentiating between paralogs and alleles
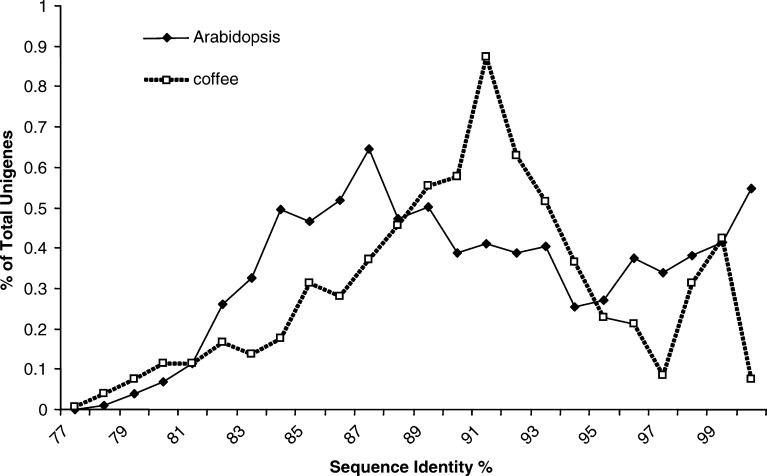
The coffee ESTs were derived from five C. canephora varieties (Table 1), however, unigene assembly for EST reads does not automatically distinguish between polymorphic alleles and highly similar paralogs. C. canephora is known to be a highly allogamous, heterozygous and polymorphic species, raising the possibility that some of the EST-derived unigenes might actually represent different allelic forms of the same gene. In order to evaluate how well paralogs were distinguished from allelic polymorphism in the unigene assembly, BLASTN (nucleotide vs. nucleotide sequences) was performed amongst all of the coffee unigenes and the results parsed to identify the best non-self matches. As a control, BLASTN was performed in a similar manner on a complete set of Arabidopsis predicted genes, including both coding and UTR regions obtained from TAIR (http://www.arabidopsis.org). Since the Arabidopsis database was derived from the complete genome sequence of a single, inbred stock (Columbia), the entire gene complement is present without the complication of allelic variation (initiative). For both coffee and Arabidopsis, pair-wise combinations for which the homology extended over more than 70% of the sequence were recorded. The sequence similarity for each of these combinations was plotted for both coffee and Arabidopsis (Fig. 3).
Fig. 3.
Plot depicting the sequence identify of the most similar match for each coffee unigene as compared with all other coffee unigenes. As a control, a similar analysis is shown for Arabidopsis genes (see Results for details)
The Arabidopsis plot showed two peaks, one with low identity (~87%) and the other with higher identity (over 99%) with 6.4 and 1.6% of genes falling into the low and high identity peaks, respectively. Like Arabidopsis, the coffee plot also showed two peaks, one with lower identity (around 91%) and the other with high identity (over 99%). These two peaks corresponded to 6.2 and 0.8% of the total unigenes, respectively. ESTs corresponding to ten pairs of coffee unigenes, from the >99% peak, were used as probes on genomic southern hybridizations to determine whether the matching pairs were truly duplicated in the coffee genome (paralogs) or rather allelic (single copy). For eight of the ten pairs, the paired ESTs hybridized the same single copy gene on southerns (data not show). Thus, a significant number (approximately 80%) of the unigenes in this peak are likely to be allelic. However, this category represented only a small portion of the coffee unigenes (0.8%). A similar experiment was performed with 11 ESTs from 11 pairs of unigenes in the second, lower homology peak (around 91% identity, see Fig. 3). In this case, the majority (8 out of 11) was determined to represent true paralogs (two or more copies in the genome) (data not shown). Thus for further discussions, it is assumed that the majority coffee EST-derived unigenes do in fact correspond to unique coffee genes.
Functional annotation of coffee EST-derived unigenes
Predicted coffee proteins
ESTScan (see Materials and methods for details) was able to identify protein-coding sequences in 12,534 coffee unigenes (95% of total unigenes), among which 1,515 (12%) were putatively full-length (starting with ATG and ending with a stop codon). Due to the cDNA library construction method, the unigenes were biased for the 3′ end—57% of the unigenes covered the 3′ end (ending with a stop codon) while only 36% covered the 5′ end (starting with ATG). Of the 5% of unigenes from which a protein sequence could not be predicted, 81% were singletons and the majority (97%) did not match to any Arabidopsis, GenBank non-redundant (NR) or Solanaceae unigene sequences, suggesting that they are not bona fide gene transcripts.
Protein domain annotation
Peptides translated from ESTScan predicted coding sequences were subjected to InterPro Scan (see Materials and methods for details) for protein domain analysis. A total of 1,678 distinct domains were identified in the database, which were present in 4,414 (35%) unigenes. Table 3 lists the 20 most represented InterPro domains found in the coffee genes, as well as comparative statistics for tomato and Arabidopsis genes. Coffee and tomato were very similar with regard to the most frequent protein domains; however, a number of striking differences were noted when comparing coffee and Arabidopsis. Most notably, the following five domains were much more common in coffee encoded proteins than in Arabidopsis: proline-rich regions, tyrosine protein kinases, glucose/ribitol dehydrogenases, ubiquitins and ubiquitin-conjugating enzymes. The latter two categories, involving ubiquitinization, suggest a more active and/or complex system for protein turnover in coffee versus Arabidopsis.
Table 3.
Twenty most abundant InterPro domains identified in coffee unigene set and comparative statistics for tomato and Arabidopsis genes
| InterPro accession | Description | % of unigenes (ranking) | ||
|---|---|---|---|---|
| Coffee | Tomato | Arabidopsis | ||
| IPR000719 | Protein kinase | 1.6 | 1.20 (1) | 3.0 (1) |
| IPR000694 | Proline-rich region | 1.3 | 0.91 (4) | 0.003 (1763) |
| IPR002290 | Serine/threonine protein kinase | 0.85 | 1.10 (2) | 0 |
| IPR001245 | Tyrosine protein kinase | 0.69 | 1.0 (3) | 0.15 (311) |
| IPR008271 | Serine/threonine protein kinase, active site | 0.61 | 0.68 (5) | 2.6 (2) |
| IPR000504 | RNA-binding region RNP-1 (RNA recognition motif) | 0.55 | 0.60 (6) | 0.59 (6) |
| IPR001680 | G-protein beta WD-40 repeat | 0.49 | 0.51 (8) | 0.51 (8) |
| IPR001611 | Leucine-rich repeat | 0.48 | 0.59 (7) | 0.59 (7) |
| IPR002048 | Calcium-binding EF-hand | 0.36 | 0.34 (13) | 0.34 (13) |
| IPR000379 | Esterase/lipase/thioesterase | 0.33 | 0.43 (10) | 0.43 (10) |
| IPR001806 | Ras GTPase superfamily | 0.32 | 0.26 (22) | 0.43 (70) |
| IPR003579 | Ras small GTPase, Rab type | 0.29 | 0.23 (27) | 0 |
| IPR005123 | 2OG-Fe(II) oxygenase superfamily | 0.27 | 0.26 (21) | 0.47 (52) |
| IPR000626 | Ubiquitin | 0.27 | 0.22 (32) | 0.40 (89) |
| IPR002401 | E-class P450, group I | 0.27 | 0.46 (8) | 0.77 (24) |
| IPR002347 | Glucose/ribitol dehydrogenase | 0.26 | 0.23 (28) | 0.33 (110) |
| IPR001005 | Myb DNA-binding domain | 0.26 | 0.34 (15) | 1.34 (8) |
| IPR005225 | Small GTP-binding protein domain | 0.26 | 0.24 (25) | 0.68 (27) |
| IPR000608 | Ubiquitin-conjugating enzymes | 0.26 | 0.21 (34) | 0.19 (221) |
| IPR007090 | Leucine-rich repeat, plant specific | 0.25 | 0.40 (12) | 1.07 (11) |
Gene ontology annotation
Gene ontology (GO) annotations were made for each coffee unigene based on InterPro domain annotation (see Materials and methods for details). A total of 3,248 unigenes (25% of total unigenes and 74% of unigenes with InterPro domain match) could be assigned a GO annotation. In order to obtain an overview of representation of biological processes by the coffee unigene set, the GO annotations were mapped to the plant GOSlim, a group of selected higher-level categories of the GO ontology (see Materials and methods for details). Figure 4 depicts the biological process GOSlim categories for each of which at least 1% of the coffee unigenes were assigned. Metabolism is the most abundant category, both in terms of number of subcategories (14 subgroups) and number of unigenes in the subcategories (2,541 unigenes and ~80% of the GO annotated unigenes). In addition, signal transduction, cell growth and/or maintenance, response to stress and response to endogenous stimulus were among the most abundant categories.
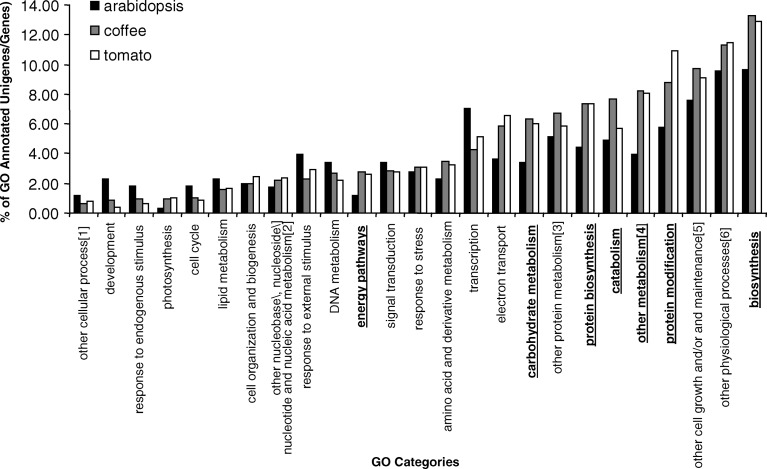
Fig. 4.
Comparison of the gene ontology-based gene annotation categories for the coffee EST-derived unigene set, tomato EST-derived unigene set and the Arabidopsis proteome. Figure contains only categories in which more than 1% of the coffee unigenes were assigned. Categories for which coffee differs most significantly from Arabidopsis are shown in underline bold. (1) Cellular processes other than signal transduction and cell growth and/or maintenance. (2) Nucleobase/nucleoside/nucleotide and nucleic acid metabolism other than DNA metabolism and transcription. (3) Protein metabolism other than protein biosynthesis and protein modification. (4) Metabolism other than amino acid and derivative metabolism, biosynthesis, carbohydrate metabolism, catabolism, electron transport, lipid metabolism, nucleobase/nucleoside/nucleotide and nucleic acid metabolism and protein metabolism. (5) Cell growth and/or maintenance other than cell cycle and cell organization and biogenesis. (6) Physiological processes other than photosynthesis, response to stress, response to endogenous stimulus, response to external stimulus and metabolism
A comparison was made between the GOSlim biological process of Arabidopsis, tomato and coffee (Fig. 4). For both the tomato and coffee unigene sets, the GO annotations were based on InterProScan results and approximately 25% of both unigene sets were assigned a GO annotation. In Arabidopsis, the genes are of full length, giving a higher chance of finding functional domains. Moreover, extensive experimental research and manual annotation has been carried out in Arabidopsis, resulting in a higher proportion of genes with assigned GO annotation. Therefore 83% of the Arabidopsis genes are assigned GO annotations. No significant differences were observed in the annotated categories for coffee versus tomato—possibly reflecting their close taxonomic affinity. However, for a number of categories, coffee had significantly different proportions of genes than Arabidopsis. The categories with the largest significant differences (P<0.001, based on Chi-square test) are: carbohydrate metabolism, other metabolism, biosynthesis, catabolism, protein biosynthesis, protein modification and energy pathways. In all cases, coffee had a significantly higher proportion of genes in these categories than Arabidopsis (Fig. 4). It is interesting to note that many of these categories center around the synthesis, breakdown or modification of compounds. One of the hallmarks of coffee is its high-level diversity of primary and secondary compounds, which contributes to the sensory quality of brewed coffee beans. The Rubiaceae family in general contains some of the most diverse species with regard to secondary metabolism and is an especially rich source of alkaloids—a number of which have pharmacological and/or psychotropic properties (Kutchan 1995; Facchini 2001). In fact, the most widely used psychotropic drug, caffeine, comes mainly from coffee. One can speculate that this metabolic diversity is reflected in the relatively high proportion of coffee genes with putative functions related to metabolism.
In silico analysis of unigene expression
Complexity and uniqueness of different stages/tissues
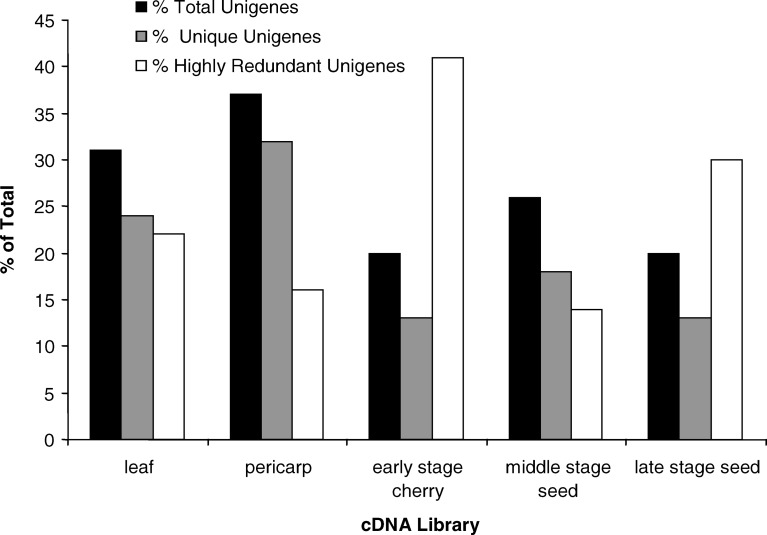
In an effort to determine which tissues gave rise to the more redundant or more complex pools of gene expression, three library/tissue parameters were calculated: (1) the percentage of total coffee unigenes represented by each library/tissue; (2) the percentage of more highly expressed unigenes (represented by ten or more ESTs from the library) in each library/tissue; (3) the percentage of unigenes from each tissue/stage which are unique to that library/tissue (not found in any other cDNA library). The results from these calculations are depicted in Fig. 5. The pericarp and leaf libraries represented more unigenes, contributed to more unique unigenes, and contained fewer highly abundant unigenes than did the early cherry, middle seed and late seed libraries—perhaps reflecting the overall lower complexity of these latter stages/tissues (Fig. 5). Among the seed stages, the middle stage of development gave more novel genes than did the early or late stages of seed development (Fig. 5).
Fig. 5.
Characteristics of each coffee cDNA library in comparison to the entire coffee EST-derived unigene set. The total unigene and highly expressed unigene categories sum to greater 100% since the same unigene may contain ESTs from more than one library
Differential expression of genes across stages/tissues
The coffee cDNA libraries were non-normalized and deeply sampled, and thus could be used to compare gene expression amongst the five cDNA libraries using statistical method (Audic and Claverie 1997). In all the pair-wise comparison between the cDNA libraries, only a small proportion of the overall unigenes (approximately 5%) showed significantly different (P<0.05) expression levels between two or more libraries (Table 4). The leaf and the early stage cherry libraries showed the highest divergence in gene expression (752 unigenes which is 5.7% of total unigenes, showed significant differential expression) while the leaf and the pericarp libraries had the lowest number of differentially expressed unigenes (384 unigenes and 2.9% of total). Interestingly, the three stages of seed development showed as much differential expression, relative to each other, as they did to non-seed tissues (leaf and pericarp). These results provide strong evidence that throughout the approximately 11 months from pollination to mature seed, a continuously changing cascade of molecular events is unfolding in the developing seed.
Table 4.
Number of coffee unigenes showing significantly (P<0.05) different expression in pairwise comparisons of cDNA libraries
| Library | Pericarp | Early stage cherry | Middle stage seed | Late stage seed |
|---|---|---|---|---|
| Leaf | 384 | 752 | 548 | 562 |
| Pericarp | 610 | 458 | 527 | |
| Early stage cherry | 602 | 728 | ||
| Middle stage seed | 585 |
Highly expressed genes
Table 5 lists the blast annotation, library composition and the best Arabidopsis/Solanaceae gene match for the 20 most highly expressed unigenes from the coffee EST database. Some of these highly expressed genes are common to and highly expressed in all plant species—a good example of this is Unigene 122071, which encodes the small subunit of ribulose bisphosphate carboxylase—a key enzyme in photosynthesis and carbon fixation (Table 5). However, a number of these highly expressed genes are unique to coffee, and many have expression patterns confined to particular tissue/stages, providing a glimpse into some of the potential unique aspects of coffee biology. A brief description of some of these is given below.
Table 5.
The 20 most highly expressed coffee unigenes: functional annotation and most similar Arabidopsis and Solanaceae homologs
| Coffee unigene#: annotation | Best match (e value/score) | EST count | ||||||
|---|---|---|---|---|---|---|---|---|
| Arbidopsis | Solanaceae Unigene_species | Total ESTs | Leaf | Pericarp | Early stage cherry | Middle stage seed | Late stage seed | |
| 125230: putative 2s seed storage protein | ND | 243065_tomato (e-103/238) | 1,219 | 7 | 15 | 21 | 1,037 | 139 |
| 120912: 11s seed storage protein | At5g44120 (1e-88/324) | 228376_tomato (0/802) | 687 | 0 | 3 | 28 | 244 | 412 |
| 121707: unknown function | At1g29050 (1e-139/489) | 246695_potato (e-163/283) | 324 | 2 | 3 | 1 | 149 | 169 |
| 120118: unknown function | At5g59320 (2e-21/99.8) | 221585_tomato (e-134/475) | 292 | 0 | 0 | 3 | 58 | 231 |
| 124988: unknown function | ND | ND | 204 | 58 | 84 | 55 | 1 | 6 |
| 120685: chitinase | At5g24090 (2e-43/172) | 214596_tomato (1e-35/84.5) | 202 | 99 | 40 | 58 | 0 | 5 |
| 124158: photoassimilate-responsive protein | At3g54040 (2e-36/149) | 196924_pepper (2e-39/138) | 182 | 1 | 1 | 2 | 150 | 28 |
| 119890: unknown function | ND | 204426_pepper (5e-07/52.8) | 183 | 0 | 0 | 0 | 183 | 0 |
| 123265: ADP-ribosylation factor | At2g47170 (1e-99/359) | 238338_tomato (0/693) | 182 | 58 | 14 | 65 | 21 | 24 |
| 124083: secretory peroxidase | At4g21960 (e-153/537) | 196145_pepper (0/681) | 161 | 55 | 5 | 19 | 49 | 33 |
| 124911: metallothionein | At5g02380 (0.32/32.3) | 207464_petunia (2e-06/51.0) | 163 | 40 | 65 | 30 | 11 | 17 |
| 119817: chitinase | At3g12500 (e-103/373) | 248120_potato (e-148/521) | 148 | 0 | 22 | 0 | 0 | 126 |
| 124815: unknown function | At3g29240 (1e-87/320) | 227940_tomato (e-146/517) | 145 | 2 | 0 | 1 | 0 | 142 |
| 122206: SAM synthase | At2g36880 (0/711) | 270415_petunia (0/887) | 142 | 1 | 9 | 130 | 0 | 2 |
| 119460: WRKY4 transcription factor | At1g80840 (3e-75/279) | 237166_tomato (e-137/487) | 123 | 0 | 0 | 123 | 0 | 0 |
| 123045: unknown function | At3g16000 (0.69/31.2) | 218824_tomato (90.36/33.1) | 123 | 81 | 17 | 22 | 3 | 0 |
| 120481: AdoMet synthase | At4g01850 (0/723) | 243236_potato (0/886) | 108 | 19 | 27 | 31 | 20 | 11 |
| 121265: Mobl/phocein | At5g45550 (e-119/425) | 196814_pepper (e-146/513) | 113 | 1 | 0 | 112 | 0 | 0 |
| 124791: plasmodesmal receptor | At5g15140 (1e-99/360) | 203764_pepper (8e-86/314) | 105 | 0 | 0 | 3 | 26 | 76 |
| 122071: rubiso small subunit | At1g67090 (9e-70/260) | 207453_petunia (3e-89/297) | 99 | 76 | 8 | 9 | 6 | 0 |
BLAST match values are given in parentheses
Bold numbers indicate library for which the highest number of ESTs were observed for each gene. Italic numbers indicate library for which the number of EST is significantly lower (P<0.05) than the highest
Seed storage protein genes
Unigene 125230: a putative 2S seed storage protein
Unigene 125230 is the most highly expressed gene across the entire coffee unigene set (1219 ESTs) and was the dominant transcript in the middle stage seed library, accounting for 10% of the ESTs at this stage (Table 5). This gene shows high homology to a tomato unigene derived from a developing seed cDNA library, but has no detectable homolog in Arabidopsis (Table 5). Other than the match with tomato, weak homology was also detected for 2S seed storage proteins from sesame, sunflower, and Brazil nut (in decreasing order of similarity). It is interesting to note that coffee, tomato, sesame and sunflower are fairly closely related taxonomically. All belong to the Asterid I/II clade of Eudicots (Fig. 1). This close phylogenetic relationship may explain why Unigene 125230 has homologous matches only in these species. Moreover, since Unigene 125230 shows homology to the 2S seed storage proteins in these related species, we conjecture that unigene 125230, its tomato unigene match, the sesame, Brazil nut and sunflower 2S storage protein gene all encode orthologous 2S seed storage proteins. This is the first time that a 2S seed storage protein has been identified in coffee or any Solanaceae species. Finally, a BLAST search of Unigene 125230 against the coffee unigene set revealed additional putative copies of the 2S seed storage protein. However, on close examination, all appear to be splicing variants or low quality sequences. Moreover, southern hybridization with a 2S cDNA probe on genomic DNA confirmed that the 2S gene is single copy in the coffee genome (data not shown).
Unigene 120912: 11S seed storage protein
Unigene 120912 is the second most abundant unigene, containing 687 ESTs (Table 5). This gene is preferentially expressed during middle and late stage seed development and shares high similarity (over 98% identity) with a previously cloned C. arabica 11S seed storage protein (Marraccini et al. 1999; Rogers et al. 1999). This unigene also has a highly significant match to the Arabidopsis 12S storage protein and to a tomato unigene derived from seed ESTs (Pang et al. 1988) (Table 5). Given these results, we conclude that unigene 12912 is allelic with the previously described 11S seed storage protein gene from C. arabica and orthologous to 11S/12S seed storage proteins in both tomato and Arabidopsis. A BLAST search of Unigene 120912 against the coffee unigene set revealed additional putative copies of the 11S seed storage protein. However, like the 2s seed storage protein (Unigene 124230), all appears to be results of alternative splicing or low sequence quality. Moreover, southern hybridization with an 11S cDNA probe on genomic DNA confirmed that the 11S gene is single copy in the coffee genome (data not shown).
Other seed-specific genes
Early stage seed development
Unigenes 122206, 119460 and 121265 were all highly expressed and specific to the early cherry stage. The early cherry library was derived from RNA from both pericarp and seed tissue while the pericarp library was derived from RNA coming from all stages of pericarp development. Thus, if the above genes were highly expressed in the pericarp of the early cherry, they should be present in the pericarp library as well. The fact that these genes showed little or no expression outside the early cherry stage, suggests that they are probably specific to early developing seed tissues and not pericarp tissues (Table 5). Unigene 122206 showed high homology to an Arabidopsis gene annotated as encoding the enzyme S-adenosyl-l-methionine (SAM) synthetase (Table 5). This enzyme synthesizes S-adenosyl-l-methionine from l-methionine and ATP and is often represented by multiple isozymes in plant species (Schroder et al. 1997). Thus, Unigene 122206 appears to be an SAM synthetase specific to early seed development (Table 5).
Unigene 119460 shows high homology to the highly conserved WRKY transcription factor family. The WRKY transcription factor is a large gene family having more than 70 members in the Arabidopsis genome (Dong et al. 2003). Previous studies showed that it is related to wounding, stress, pathogen infection and senescence in many plant species. In some recent studies, the WRKY protein family was found to be involved in sugar signaling in barley and seed development in Arabidopsis (Johnson et al. 2002; Sun et al. 2003). However, the function of the best Arabidopsis match to Unigene 119460 (At1g80840) has not been determined. Hence, understanding the function of this highly expressed, WRKY-like coffee gene awaits further study.
Unigene 121265 is highly homologous to a gene in Arabidopsis annotated as encoding a Mob1/phocein protein. Mob1/phocein proteins are found in virtually all eukaryotes. While they are conjectured to be involved in cell cycle control, there is still little experimental evidence demonstrating biological function (Pon 2004). Thus it seems premature to conjecture what role Unigene 121265 might have that is specific to the early development of coffee seeds.
Middle stage seed development
As described earlier, Unigene 125230 is a putative 2S seed storage protein with peak expression during middle seed development. Also showing preferential expression during this same stage were Unigenes 121707, 124158 and 119890. Unigene 121707 is a gene of unknown function with high homology matches both in Arabidopsis and Solanaceae EST-derived unigenes (Table 5). Unigene 124158 is homologous to an Arabidopsis gene classified as a photoassimilate-responsive protein, which is related to pathogenesis (Herbers 1995). Finally, Unigene 119890, which is also specific to middle stage seed development, is apparently a gene unique to coffee, which will be discussed more in the following section.
Late stage seed development
As discussed previously, Unigene 120912 corresponds to the 11S seed storage protein, which is largely expressed late in seed development. Other genes with preferential expression in late stage seeds are Unigenes 120118, 119817 and 124791. Unigene 119817 likely encodes a chitinase and is further discussed in the next section. Unigene 120118 shows high homology to genes in both Arabidopsis and Solanaceae EST-derived unigene sets; however, none have known function. Unigene 124791 gives a strong match to an Arabidopsis gene annotated as a plasmodesmatal receptor.
Two highly expressed genes with homology to chitinase
Unigenes 120685 and 119817 show high sequence similarity to a number of genes classified as chitinases in other organisms. Chitinases are a large and diverse class of proteins, some of which have been implicated in resistance to fungi in various plant species, including coffee (Rojas-Herrera 2002; Chen et al. 2003). The two unigenes differ in that Unigene 120685 is expressed in leaves, pericarp and early stage cherries, but not in mid or late stage seed development (Table 5). Unigene 119817, on the other hand, was found to be exclusively expressed in late stage developing seeds and pericarp tissue. As previously mentioned, early stage cherries contained both pericarp and seed tissues. The fact that Unigene 120685 was not found in the middle and late stages of seed development suggests that this gene may not be expressed in seeds, but rather in the maternally derived pericarp and leaf tissues. Based on these results, one can speculate that these two putative chitinase genes may be involved in pathogen defense in developing CHERRIES, with Unigene 120685 being expressed in early developing, post pollination pericarp and leaf tissues and Unigene 119817 being expressed primarily late in seed development, just prior to maturity.
Highly expressed genes unique to coffee
Unigene 124988
This highly expressed unigene had no significant matches in the Arabidopsis proteome, Solanaceae EST-derived databases, GenBank NR databases, or GenBank dbest. Moreover, the predicted protein encoded by Unigene 124988 has no recognizable domains, which might give clues to its function. ESTs for this unigene were detected in all five libraries, with highest expression being observed in the pericarp (Table 5).
Unigene 119890
Unigene 119890 also has no significant match in any of the tested databases, with the possible exception of a very weak match in the Solanaceae unigene sets (the best hit was from pepper with an e value of 5e-7, Table 5). Like Unigene 124988, its predicted protein has no recognizable domains. Unigene 119890 was highly and exclusively expressed in the middle stage of developing seeds (Table 5).
The fact that neither Unigene 124988 nor Unigene 119890 have counterparts in any other databases suggests that they may represent coffee-specific genes or genes that have been evolving at such a rapid rate that they no longer bear any recognizable homologies with proteins from other plants, including the closely related Solanaceous plants. We speculate that these genes may be related to chemical or morphological features unique to coffee.
Gene families unique or significantly expanded in coffee
The predicted protein sequences for the coffee unigene set and the Arabidopsis proteome were combined and organized into protein families using tribeML software (see Materials and methods for details). In most cases, the number of gene family members was higher in arabidopsis than in coffee (data not shown). However, this is to be expected since the entire Arabidopsis genome has been sequenced (thus revealing all genes), whereas in coffee the EST-derived unigene set is by nature incomplete and unlikely to contain all members of any given gene family. However, despite the incomplete nature of the coffee EST-derived unigene set, several cases was encountered in which coffee has more gene family members than does Arabidopsis. To further investigate this topic, a list was made of the gene families for which coffee has significantly more members than Arabidopsis (Table 6). We refer to these as coffee-expanded gene families. In addition, gene families found in coffee, but for which no counterparts exist in Arabidopsis were also compiled (Table 7). We refer to these as coffee-unique gene families. All gene families listed in both Tables 6 and 7 were subjected to manual examination to remove any members determined to be results of alternative splicing and/or poor sequence quality—both of which could inflate the number of putative gene family members in coffee. Hence, the differences in copy number reported are likely to be due to true differences in gene copy number and not artifacts of EST-based gene predictions.
Table 6.
Gene families expanded in coffee relative to Arabidopsis
| Family # | # Arabidopsis family member | # Coffee family member | Longest coffee member | Annotation |
|---|---|---|---|---|
| 266 | 1 | 21 | 122330 | Retrotransposon gag protein, class I |
| 180 | 5 | 14 | 124952 | Polygalacturonase isoenzyme 1 beta subunit with BURP domain |
| 632 | 1 | 12 | 123451 | Acidic endochitinase |
| 386 | 2 | 10 | 124158 | Photoassimilate-responsive protein |
| 382 | 4 | 8 | 119672 | Hypersensitive-induced protein, band 7 protein |
| 394 | 2 | 7 | 122791 | E-class P450 |
| 483 | 2 | 6 | 120054 | Bet v I allergen |
| 623 | 3 | 6 | 119581 | Root hair defective protein |
| 1,182 | 1 | 5 | 126674 | Unknown function |
| 695 | 2 | 5 | 126974 | Tyrosine decarboxylase |
| 783 | 2 | 5 | 122423 | Unknown function |
| 1,117 | 2 | 5 | 119449 | Trypsin inhibitor Kunitz |
Table 7.
Gene families unique to coffee in comparison to Arabidopsis
| Gene family # | # Family member | Longest member | Solanaceae hit | Annotation |
|---|---|---|---|---|
| 243 | 27 | 122956 | 258190 potato | Retrotransposon gag protein, class II |
| 687 | 11 | 120121 | 221585 tomato | Thaumatin, pathogenesis related |
| 965 | 10 | 119718 | 249401 potato | Zn-finger, CCHC type |
| 974 | 10 | 120244 | 2610402 potato | Disease resistance protein (TIR-NBS-LRR class) |
| 852 | 9 | 119638 | 225732 tomato | Retrotransposon gag protein, classs III |
| 360 | 8 | 121998 | 23671 tomato | Disease resistance protein |
| 1,019 | 7 | 124574 | 222350 tomato | Leucine-rich repeat, disease resistance protein |
| 1,607 | 7 | 122216 | none | Unknown function |
| 1,610 | 7 | 130519 | none | Unknown function |
| 1,676 | 7 | 126264 | 243065 tomato | Unknown function |
| 708 | 6 | 123769 | 236157 tomato | ABA/WDS induced protein |
| 1,852 | 5 | 120284 | 213688 tomato | Proline-rich region, extension-like protein |
| 2,362 | 5 | 122218 | 237314 tomato | Unknown function |
| 2,459 | 5 | 124466 | 267984 potato | Leucine-rich repeat, plant specific, receptor-related protein kinase |
Coffee-expanded gene families
The most expanded gene family in coffee corresponds to a retrotransposon gag protein (Table 6). This result has two implications. First, the retroelement encoding this gag protein occurs at a higher frequency in coffee compared with Arabidopsis, although we cannot determine whether this difference is due to a true expansion of this element in coffee subsequent to divergence from Arabidopsis, or rather a loss of the element in the Arabidopsis lineage. Second, the fact that this retrotransposon gag protein element was discovered in an EST-database indicates that this particular retroelement is being transcribed in the coffee genome, and hence may represent an active retrotransposon.
Another gene family for which coffee has significantly more members than Arabidopsis, encodes proteins annotated in Arabidopsis as acid endochitinases and photoassimilate-responsive proteins (Table 6). As noted earlier, chitinases are associated with fungal resistance and are among the most highly expressed genes in coffee. The fact that chitinases are both highly expressed and represented by an expanded gene family in coffee may reflect a greater need for fungal resistance engendered both by the perennial nature of coffee and the fact that it is a tropical species for which a multiplicity of fungal pathogens is common. The reasons for the putative expansions of the other gene families listed in Table 6 remain for future studies to determine.
Coffee-unique gene families
Table 7 lists the top gene families (based on copy number), which occur in coffee, but not in Arabidopsis. Of the 15 gene families listed, four are of unknown function. For those that could be functionally annotated, five (45%) have putative functions related to disease resistance, such as TIR-NBS-LRR disease resistance proteins, LRR proteins and thaumatin pathogenesis-related proteins (Table 7). These finding are consistent with rapid evolution of genes/gene families related to disease resistance, likely driven by selection pressure from continuously changing pathogens and/or pathogens unique to the particular environments of a species (Meyers 1998; Michelmore and Meyers 1998). Also included in this list of coffee-unique gene families are two, which encode retrotransposon gag-proteins (Table 7).
Comparison of the coffee gene repertoire with that of Arabidopsis and Solanaceae species
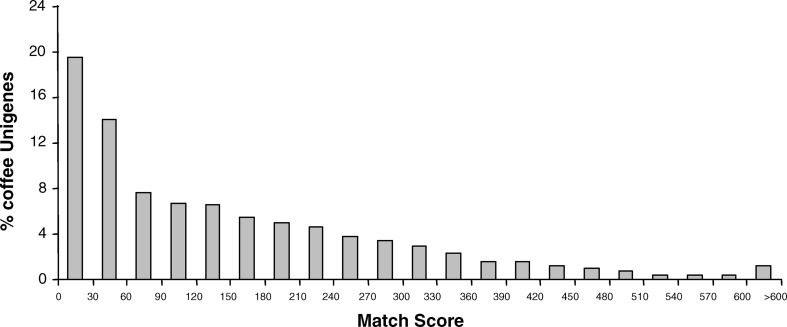
Each coffee unigene was screened against the Arabidopsis proteome using BLASTX, which compares all six translated frames of the query with the target. In total, 21% of the coffee unigenes had no clear match in Arabidopsis (match score 0–30) (Fig. 6). These coffee genes, which lack an Arabidopsis counterpart, may have evolved rapidly since coffee and Arabidopsis diverged from their last common ancestor, and hence their counterparts in each genome are no longer recognizable by standard string searches like BLAST. To investigate this topic further, the 2,853 unigenes with no clear match in Arabidopsis (match score<30, see Fig. 6) were screened against the Solanaceae EST-derived unigene sets using TBLASTX which compares all six translated frames of both query (coffee unigenes) and target (Solanaceae unigenes, see Materials and methods for details) sequences. The family Solanaceae is much more closely related to coffee than is Arabidopsis (Fig. 1), and hence coffee genes without a match in Arabidopsis may well have recognizable orthologs in the Solanaceae. The results show that a large portion (90%) of the coffee genes that lack a match in Arabidopsis do have a match in Solanaceae at the same stringency criteria (match score>30). For many of these cases, however, the coffee-Solanaceae matches are still quite diverged (low match scores) suggesting that the proteins encoded by these genes have been evolving at a rapid rate in dicots.
Fig. 6.
Histogram showing match scores for each coffee unigene as compared with its best match in the Arabidopsis proteome
There were, however, some exceptions to this trend represented by coffee genes having no match in Arabidopsis, yet a strong match in Solanaceae. These latter cases may represent genes that have been under differential selection pressure since the time that the Rosids (which includes Arabidopsis) diverged from the last common ancestor with the Asterids (which includes Solanaceae and coffee) (Fig. 1). Alternatively, these genes may have been lost in the lineage leading to Arabidopsis, but retained in the Asterids. In either case, such genes may have functional roles that are, in part, responsible for the morphological, developmental and chemical characteristics that differentiate Arabidopsis from Rosid species (e.g. coffee, Solanaceae). In an effort to shed more light on this issue, the 20 coffee unigenes having the highest match scores with Solanaceae, but lacking a counterpart in Arabidopsis were parsed and annotated (Table 8). Seven of these coffee genes have no match either in GenBank NR or dbest hence their function is unknown. Another unigene (Unigene 125230, the 2s seed storage protein, see above discussion) had weak match to ESTs of sesame, a species also belonging to the Asterid I clade (Fig. 1). Thus, these eight genes (40%) appear to be specific to the tomato/coffee/sesame Asterid I clade (Fig. 1, Table 8). Elucidating their function may shed light on the molecular processes, which diversified early in the evolution of dicot species, especially those species belonging to the Asterid clade of dicots.
Table 8.
Coffee genes not found in Arabidopsis, but with conserved counterparts in tomato or other Solanaceous species
| Coffee unigene | Solanaceae EST-derived unigene match | Score | GenBank (non-redundant and dbest) best match | Score | Annotation |
|---|---|---|---|---|---|
| 124978 | 240871 tomato | 454 | Unknown function | ||
| 121324 | 235756 tomato | 429 | gblCB686389.1 [Brassica napus] | 44 | Unknown function |
| 131820 | 213100 tomato | 426 | gil50252229.1 [Oryza sativa] | 73 | Unknown function |
| 121542 | 240321 tomato | 416 | refiNP_922676.1 [Oryza sativa] | 75 | Unknown function |
| 131934 | 219759 tomato | 377 | embiCAE05735.1 [Oryza sativa] | 297 | TFIIH basal transcription factor p52 subunit |
| 121140 | 236347 tomato | 320 | Unknown function | ||
| 131445 | 225435 tomato | 320 | Unknown function | ||
| 125230 | 243065 tomato | 238 | gil13183175 [Seasame indicum] | 45 | 2S albumin |
| 131030 | 246364 potato | 213 | refINP_524404.1 [Drosophila melanogaster] | 110 | Phospyhatidyl inositol transfer protein |
| 120120 | 237254 tomato | 202 | gbICF349465.1 [Rose] | 52 | Unknown function |
| 126635 | 237314 tomato | 185 | Unknown function | ||
| 126575 | 237314 tomato | 182 | Unknown function | ||
| 130675 | 209387 petunia | 177 | gbICK093976.1 [Populus tremula] | 438 | Unknown function |
| 128020 | 237314 tomato | 167 | Unknown function | ||
| 123615 | 249253 potato | 163 | gbIAAO73272.1 [Oryza sativa] | 140 | Unknown function |
| 126432 | 240551 tomato | 163 | gil34878866 [Rattus norvegicus] | 56 | Phosphatidylinositolglycan class N |
| 124384 | 197378 pepper | 156 | gblCA815435.1 [Vitis vinifera] | 1,009 | Unknown function |
| 122126 | 239632 tomato | 153 | Unknown function | ||
| 131601 | 232010 tomato | 145 | gbICK229938.1 [Macaca mulatta] | 74 | 40S Ribosomal protein S21 |
| 119644 | 237150 tomato | 143 | refINP_921250.1 [Oryza sativa] | 70 | Helicase |
The GenBanks Best match exclude those from Solanaceae, Coffea and Hedyotis (both members of the Rubiaceae family). Solanaceae EST-derived
Surprisingly, 8 (40%) of the 20 coffee genes having no match in Arabidopsis did have matches in species phylogenetically distant from both coffee and Arabidopsis, including two to non-plant species (Drosophila and rat) (Table 8). Five of these matches were to rice genes, which is a monocot and highly divergent from coffee, Arabidopsis and other dicot species (Table 8). The fact that all of these species diverged from Arabidopsis and coffee long before the latter diverged from each other suggests that these genes may have been present in the last common ancestor of Arabidopsis and coffee/Solanaceae, but subsequently lost in the Arabidopsis lineage.
Coffee genes share greater similarity to genes in tomato/Solanaceae than to Arabidopsis
As discussed earlier, coffee is much more closely related to the Solanaceae than to Arabidopsis (Fig. 1). Hence, Solanaceae species may be better models for coffee genomics than Arabidopsis. The results for fast-evolving genes, presented above, are consistent with this prediction. To further investigate this assertion, the degree to which each coffee unigene matched Arabidopsis versus Solanaceae was investigated. In doing this analysis, one has to keep in mind that the entire gene repertoire of Arabidopsis is known, whereas the EST-derived unigene sets for Solanaceae do not represent the entire gene repertoire of these species. We estimate that the combined EST-derived unigene sets of Solanaceae species represent as much as three-quarters of the Solanaceae gene content (Hoeven et al. 2002). Moreover, Arabidopsis genes are of full length, while Solanaceae EST-derived unigenes are not necessarily of full length.
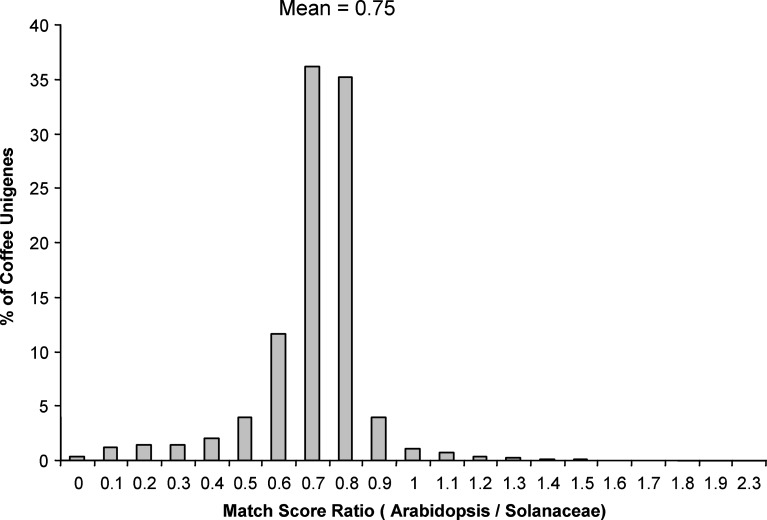
To mitigate against these factors, each coffee unigene was first screened, via TBLASTX, against the Solanaceae EST-derived unigene databases. Only those coffee unigenes matching a Solanaceae unigene with a match score greater than 100 were compared with the Arabidopsis proteome. Moreover, the match to Arabidopsis was confined to only the portion of the coffee unigene matching a Solanaceae EST-derived unigene. The ratio of the Arabidopsis match score divided by the Solanaceae match score was used as a measure of the degree to which the Solanaceae homolog shows greater similarity to the coffee unigene than to the closest Arabidopsis homolog. The results of this analysis are shown in Fig. 7. The ratio of the Arabidopsis/Solanaceae match score averaged 0.75, indicating a significantly better match in most cases with Solanaceae versus Arabidopsis. Moreover, in more than 95% of the cases, the ratio was less than one, indicating that, in most instances, coffee contains genes significantly more homologous to Solanaceae than to Arabidopsis. For those cases in which the ratio was greater than 1 (5%), we cannot rule out the possibility that the true ortholog was missing in the Solanaceae EST-derived unigene sets, hence the comparison was between paralogs. These results, combined with the results on fast evolving genes presented earlier, provide clear evidence that the Solanaceae provide a much better model than Arabidopsis for genomic and biological studies in coffee. This is especially relevant as sequencing of the tomato genome is currently underway (http://www.sgn.cornell.edu/help/about/tomato_sequencing.html).
Fig. 7.
Ratio of highest Arabidopsis match score to highest Solanaceae match score for individual coffee unigene. The analysis restricted to coffee unigenes with a Solanaceae match score >100
Conclusions
Herein, we describe the development and analysis of a large EST database for coffee. The resulting 47,000 ESTs correspond to 13,175 unique genes (unigenes), a large portion of which are expressed during seed development—a stage important to coffee as a crop and one for which our understanding of molecular development is still rudimentary. To our knowledge, this is the largest public database for seed-derived ESTs. Hence, this EST database represents a new public resource, which can facilitate a better understanding of seed development, as well as genomic, molecular and breeding research in coffee. By comparisons with Arabidopsis and Solanaceous species, we have identified the two major seed storage proteins of coffee (2S and 11S) and demonstrated that these proteins are expressed at different times during seed development. Through in silico gene expression analysis, we have identified a number of highly expressed genes that show high specificity for different stages of seed development as well as for the pericarp tissue that surrounds the seeds. Many of these highly expressed genes are unique to coffee and/or the Asterid clade of higher plants. While the functions of most of these highly expressed, tissue/stage specific genes remain to be determined, the fact that they have been identified points the way to promoters, which can potentially be used to drive gene expression in specific stages/tissues of the coffee plant. Many of these genes are specific to defined periods of seed and/or pericarp development—both critically important for insect/pathogen resistance and in determining the quality of the coffee bean with respect to commercial coffee products.
Coffee, as a member of the family Rubiaceae, is distantly related to the model species Arabidopsis. A computational comparison of the coffee EST-derived unigene set with the sequence databases for Arabidopsis and Solanaceous species (e.g. tomato, pepper), indicate that the latter are much better genomic models for coffee than is Arabidopsis. These results are consistent with the fact that coffee and solanaceous species share very similar chromosome architecture and are closely related, both belonging to the Asterid I clade of dicot plant family. Moreover, the ability to identify orthologous genes between coffee and tomato opens the door to eventually developing detailed comparative maps for these two species and to the sharing of genomic and biological tools/discoveries—an outcome that should expedite research in both taxa.
Acknowledgements
This work was supported by a grant from the Nestle Corporation to increase the knowledge base on coffee for the benefit of all stakeholders of the production chain, and funding from the National Science Foundation Plant Genome Program (no. 0116076). The authors wish to thank Dr. Ir Zaenudin Su, Dr. T. Wahyudi, Dr. Surip Mawardi and P. Priyono of the Indonesian Coffee and Cacao Research Institute (ICCRI) for supplying all the leaf and fruit samples used here from the ICCRI collection. We thank V. Caillet and Dr. Pierre Marraccini for their respective efforts in preparing the RNA samples used in the cDNA library construction. Thanks to Dr. Anne Frary for critical reading and editing of the manuscript.
References
- Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, Lipman D. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 1997;25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Apweiler R, Attwood TK, Bairoch A, Bateman A, Birney E, et al. The InterPro database, an intergrated documentation resource for protein families, domains and functional sites. Nucleic Acids Res. 2001;29:37–40. doi: 10.1093/nar/29.1.37. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Audic S, Claverie J-M. The significance of digital gene expression profiles. Genome Res. 1997;7:986–995. doi: 10.1101/gr.7.10.986. [DOI] [PubMed] [Google Scholar]
- Chase MW, Soltis DE, et al. Phylogenetics of seed plants—an analysis of nucleotide-sequences from the plastid gene rbcl. Ann Mo Bot Gard. 1993;80:528–580. doi: 10.2307/2399846. [DOI] [Google Scholar]
- Chen ZJ, Ribeiro R, Silva MC. Heat shock-induced susceptibility of green coffee leaves and berries to Colletotrichum gloeosporioides and its association to PR and hsp70 gene expression. Physiol Mol Plant P. 2003;63:181–190. doi: 10.1016/j.pmpp.2003.11.004. [DOI] [Google Scholar]
- Crepet WL, Nixon KC, Gandolfo MA. Fossil evidence and phylogeny: the age of major angiosperm clades based on mesofossil and macrofossil evidence from cretaceous deposits. Am J Bot. 2004;91:1666–1682. doi: 10.3732/ajb.91.10.1666. [DOI] [PubMed] [Google Scholar]
- Dong JX, Chen CH, Chen ZX. Expression profiles of the Arabidopsis WRKY gene superfamily during plant defense response. Plant Mol Biol. 2003;51:21–37. doi: 10.1023/A:1020780022549. [DOI] [PubMed] [Google Scholar]
- Enright AJ, Dongen SV, Ouzounis CA. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 2002;30:1575–1584. doi: 10.1093/nar/30.7.1575. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ewing B, Hillier L, Wendl MC, Green P. Base-calling of automated sequencer traces using Phred. I. Accuracy assessment. Genome Res. 1998;8:175–185. doi: 10.1101/gr.8.3.175. [DOI] [PubMed] [Google Scholar]
- Facchini PJ. Alkaloid biosynthesis in plants: biochemistry, cell biology, molecular regulation, and metabolic engineering applications. Annu Rev Plant Phys Plant Mol Biol. 2001;52:29–66. doi: 10.1146/annurev.arplant.52.1.29. [DOI] [PubMed] [Google Scholar]
- Gandolfo MA, Nixon KC, Crepet WL. A new fossil flower from the Turonian of New Jersey: Dressiantha bicarpellata gen. et sp. nov. (Capparales) Am J Bot. 1998;85:964–974. doi: 10.2307/2446363. [DOI] [PubMed] [Google Scholar]
- Herbers KMG, Badur R, Sonnewald U. A simplified procedure for the subtractive cDNA cloning of photoassimilate-responding genes: isolation of cDNAs encoding a new class of pathogenesis-related proteins. Plant Mol Biol. 1995;29:1027–1038. doi: 10.1007/BF00014975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herrera JC, Combes MC, Anthony F, Charrier A, Lashermes P. Introgression into the Allotetraploid Coffee (Coffea arabica L.): segregation and recombination of the C. canephora genome in the tetreploid interspecific hybrid (C. arabica×C. canephora) Theor Appl Genet. 2002;104:661–668. doi: 10.1007/s001220100747. [DOI] [PubMed] [Google Scholar]
- Hoeven RVd, Ronning C, Giovannoni J, Matin G, Tanksley S. Deductions about the number, organization and evolution of genes in the tomato genome based on analysis of a large expressed sequence tag collection and selective genomic sequencing. Plant Cell. 2002;14:1441–1456. doi: 10.1105/tpc.010478. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Huang X, Madan A. CAP3: a DNA sequence assembly program. Genome Res. 1999;9:868–877. doi: 10.1101/gr.9.9.868. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Initiative TAG. Analysis of the genome sequence of the flowering plant Arabidopsis thaliana. Nature. 2000;408:796–815. doi: 10.1038/35048692. [DOI] [PubMed] [Google Scholar]
- Iseli C, Jongeneel CV, Bucher P (1999) ESTScan: a program for detecting, evaluating, and reconstructing potential coding regions in EST sequences. American Association of Artificial Intelligence [PubMed]
- Johnson CS, Kolevski B, Smyth DR. TRANSPARENT TESTA GLABRA2, a trichome and seed coat development gene of Arabidopsis, encodes a WRKY transcription factor. Plant Cell. 2002;14:1359–1375. doi: 10.1105/tpc.001404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kutchan TM. Alkaloid biosynthesis—the basis for metabolic engineering of medicinal plants. Plant Cell. 1995;7:1059–1070. doi: 10.1105/tpc.7.7.1059. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee S, Kim SY, Chung E, Joung YH, Pai HS, Hur CG, Choi D. EST and microarray analyses of pathogen-responsive genes in hot pepper (Capsicum annuum L.) non-host resistance against soybean pustule pathogen (Xanthomonas axonopodis pv. glycines. Funct Integr Genomics. 2004;4:196–205. doi: 10.1007/s10142-003-0099-1. [DOI] [PubMed] [Google Scholar]
- Marraccini P, Deshayes A, Petiard V, Rogers WJ. Molecular cloning of the complete 11S seed storage protein gene of Coffea arabica and promoter analysis in transgenic tobacco plants. Plant Physiol Biochem. 1999;37:273–282. doi: 10.1016/S0981-9428(99)80025-4. [DOI] [Google Scholar]
- Meyers BCCD, Shen KA, Sivaramakrishnan S, Lavelle DO, Zhang Z, Michelmore RW. The major resistance gene cluster in lettuce is highly duplicated and spans several megabases. Plant Cell. 1998;10:1817–1832. doi: 10.1105/tpc.10.11.1817. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Michelmore RW, Meyers BB. Clusters of resistance genes in plants evolve by divergent selection and a birth-and-death process. Genome Res. 1998;8:1113–1130. doi: 10.1101/gr.8.11.1113. [DOI] [PubMed] [Google Scholar]
- Mizuno K, Kuda A, Kato M, Yoneyama N, Tanaka H, Ashihara H, Fujimura T. Isolation of a new dual-functional caffeine synthase gene encoding an enzyme for the conversion of 7-methylxanthine to caffeine from coffee (Coffeaarabica L.) FEBS Lett. 2003;534:75–81. doi: 10.1016/S0014-5793(02)03781-X. [DOI] [PubMed] [Google Scholar]
- Moisyadi S, Neupane KR, Stiles JI. Cloning and characterization of a cDNA encoding xanthosine-N7-methyltransferase from coffee (Coffea arabica) Acta Hort. 1998;461:367–377. doi: 10.17660/ActaHortic.1998.461.42. [DOI] [Google Scholar]
- O’Brien TG, Kinnaird MF. Caffeine and conservation. Science. 2003;300:587. doi: 10.1126/science.1082328. [DOI] [PubMed] [Google Scholar]
- Ogawa M, Herai Y, Koizumi N, Kusano T, Sano H. 7-methylxanthine methyltransferase of coffee plants gene isolation and enzyme properties. J Biol Chem. 2001;276:8213–8218. doi: 10.1074/jbc.M009480200. [DOI] [PubMed] [Google Scholar]
- Pang PP, Pruitt RR, Meyerowitz EM. Molecular cloning, genomic organization, expression and evolution of 12s seed storage protein genes of Arabidopsis thaliana. Plant Mol Biol. 1988;11:805–820. doi: 10.1007/BF00019521. [DOI] [PubMed] [Google Scholar]
- Pinto-Maglio CAF, Cruz ND. Pachytene chromosome morphology in Coffea L. II. C. arabica L. complement. Caryologia. 1998;51:19–35. doi: 10.1080/00087114.1998.10589117. [DOI] [Google Scholar]
- Ponchon LDC, Kajava AV, Fesquet D, Padilla A. NMR solution structure of Mob1, a mitotic exit network protein and its interaction with an NDR kinase peptide. J Mol Biol. 2004;337:167–182. doi: 10.1016/j.jmb.2004.01.010. [DOI] [PubMed] [Google Scholar]
- Rick CM. Some cytogenetic features of the genome in diploid species. Stadler Sym. 1971;1:153–174. [Google Scholar]
- Rogers WJ, Bezard G, Deshayes A, Meyer I, Petiard V, Marraccini P. Biochemical and molecular characterization and expression of the 11S-type storage protein from Coffea arabica endosperm. Plant Physiol Biochem. 1999;37:261–272. doi: 10.1016/S0981-9428(99)80024-2. [DOI] [Google Scholar]
- Rojas-Herrera RL-VV. Induction of a class III acidic chitinase in foliar explants of Coffea arabica L. during somatic embryogenesis and wounding. Plant Sci. 2002;163:705–711. doi: 10.1016/S0168-9452(02)00156-5. [DOI] [Google Scholar]
- Ronning CM, Stegalkina SSAR, Bougri O, Hart AL, Utterbach TR, Vanaken SE, Riedmuller SB, White JA, Cho J, Pertea GM, Lee Y, Karamycheva S, Sultana R, Tsai J, Quackenbush J, Griffiths HM, Restrepo S, Smart CD, Fry WE, van der Hoeven R, Tanksley S, Zhang PF, Jin HL, Yamamoto ML, Baker BJ, Buell CR. Comparative analyses of potato expressed sequence tag libraries. Plant Physiol. 2003;131:419–429. doi: 10.1104/pp.013581. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schroder G, Eichel J, Breinig S, Schroder J. Three differentially expressed S-adenosylmethionine synthetases from Catharanthus roseus: molecular and functional characterization. Plant Mol Biol. 1997;33:211–222. doi: 10.1023/A:1005711720930. [DOI] [PubMed] [Google Scholar]
- Suh MCKM, Hur CG, Bae JM, Park YI, Chung CH, Kang CW, Ohlrogge JB. Comparative analysis of expressed sequence tags from Sesamum indicum and Arabidopsis thaliana developing seeds. Plant Mol Biol. 2003;52:1107–1123. doi: 10.1023/B:PLAN.0000004304.22770.e9. [DOI] [PubMed] [Google Scholar]
- Sun CXPS, Olsson H, Boren M, Ahlandsberg S, Jansson C. A novel WRKY transcription factor, SUSIBA2, participates in sugar signaling in barley by binding to the sugar-responsive elements of the iso1 promoter. Plant Cell. 2003;15:2076–2092. doi: 10.1105/tpc.014597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uefuji H, Ogita S, Yamaguchi Y, Koizumi N, Sano H. Molecular cloning and functional characterization of three distinct N-methyltransferases involved in the caffeine biosynthetic pathway in coffee plants. Plant Physiol. 2003;132:372–380. doi: 10.1104/pp.102.019679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- White JATT, Newman T, et al. A new set of Arabidopsis expressed sequence tags from developing seeds. The metabolic pathway from carbohydrates to seed oil. Plant Physiol. 2000;124:1582–1594. doi: 10.1104/pp.124.4.1582. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wrigley G (1988) Coffee. (Logman Scientific & Technical)
- Zdobnov EM, Apweiler R. InterProScan—an intergration for the signature-recognition methods in InterPro. Bioinformatics. 2001;17:847–848. doi: 10.1093/bioinformatics/17.9.847. [DOI] [PubMed] [Google Scholar]