Figure 5.
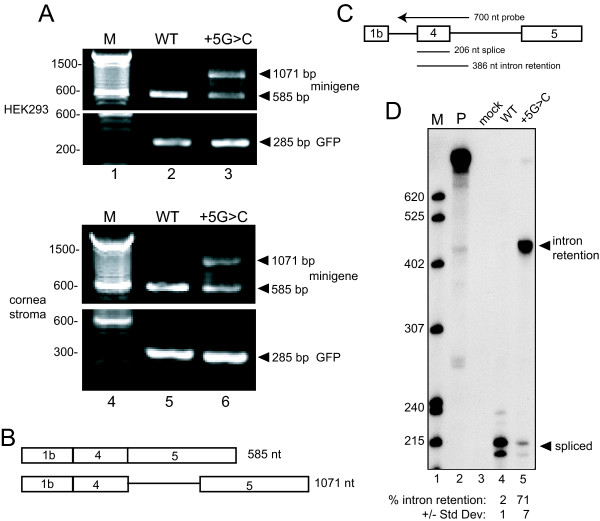
The IVS4+5G>C mutation causes intron retention. A. RNA from HEK293 cells (lanes 2 and 3) or human cornea stromal cells (lanes 5 and 6) transfected with WT or IVS4+5G>C minigenes and GFP was subjected to RT-PCR and products were resolved on an agarose gel. The positions of correctly spliced (585 bp), intron retention (1071 bp), and GFP products are shown at the right. M, 100 bp size standards with selected sizes shown. B. Diagram depicting the RNA structure of RT-PCR products from (A) that were cloned and sequenced. At the top is correctly spliced RNA, the bottom is RNA that retains the intron downstream of exon 4. C. Schematic of the RNase protection probe and protected products observed in D. The sizes of the probe and protected products are indicated. D. RNase protection assay (RPA). HEK293 RNA was subjected to RPA, protected products were resolved on a 6%–8 M urea polyacrylamide gel, and a phosphorimage is presented. Correct splicing generates a ~206 nt band and intron retention gives a ~386 nt band. M, 32P-labeled pBR322/MspI markers with sizes indicate on the left; P, undigested probe; mock, RPA using RNA from untransfected cells; WT, RNA from WT minigene-transfected cells; +5G>C, RNA from mutant IVS4+5G>C minigene-transfected cells. The positions of correctly spliced and intron retention protected products are shown at the right. The doublet bands for spliced product are likely due to incomplete RNase digestion. Below the lanes is quantitation of % intron retention (+/- standard deviation) as assessed by PhosphorImager analysis (n = 17).