TABLE 2.
Quantitative RT-PCR analysis of mRNA levels in tlc1 through tlc5
| Gene | Reaction efficiencya
|
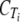 b b
|
Avg ΔCT from that of tlc1c | Expression relative to that of tlc1d | ||
|---|---|---|---|---|---|---|
| Sample 1 | Sample 2 | Sample 1 | Sample 2 | |||
| tlc1 | 3.1 | 3.6 | 25.5 | 25.7 | 1.0 | |
| tlc2 | 2.9 | 3.3 | 27.1 | 27.9 | −1.9 | 0.3 |
| tlc3 | 3.5 | 4.0 | 27.5 | 27.3 | −1.8 | 0.3 |
| tlc4 | 3.4 | 3.9 | 23.5 | 22.2 | 2.8 | 6.7 |
| tlc5 | 3.1 | 3.9 | 25.8 | 25.3 | 0.1 | 1.1 |
As described in Materials and Methods, quantitative PCR was performed in duplicate on serial dilutions of the tlc-specific cDNA, and the CT was plotted against the log template concentration from two biological replicates. Linear regression analysis determined the reaction efficiency, where a theoretical slope of 3.3 is expected.
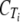 , the ordinate intercept of the linear regression.
, the ordinate intercept of the linear regression.
The 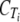 values from the two samples were averaged and used to calculate the difference in CT (ΔCT) from that of tlc1.
values from the two samples were averaged and used to calculate the difference in CT (ΔCT) from that of tlc1.
The ΔCT values were used to determine expression relative to that of tlc1 where change (n = fold) was 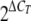 .
.
