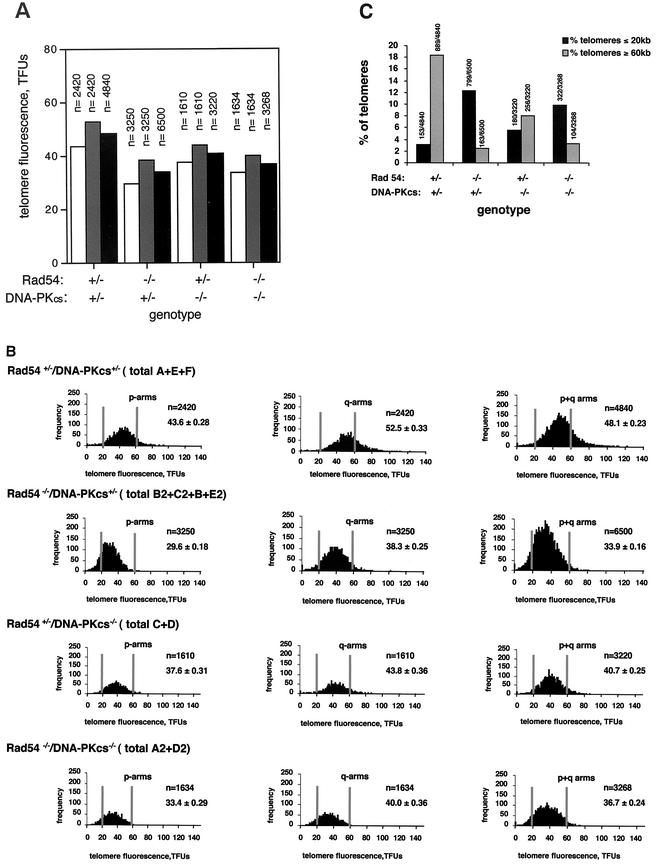
FIG. 1.
Telomere length analysis by Q-FISH. (A) Average telomere fluorescence and standard error of q-telomeres, p-telomeres, or all (p+q) telomeres in primary MEFs grouped by genotype. Fluorescence is expressed in telomere fluorescence units (TFUs), where 1 TFU corresponds to 1 kb of TTAGGG repeats (30). Each value represents the mean of 10 metaphases and of the indicated number (n) of individual telomere values. n refers to the total number of telomere values used for calculation of the average telomere fluorescence for each chromosome arm, as well as for the sum of both arms. Bars: ▪, average of q- and p-telomeres; , q-telomeres; □, p-telomeres. Despite the wide heterogeneity in individual telomere fluorescence intensity values (see, for example, Fig. 1B), the standard errors of the mean are very small due to the large number of datum points (see “n” values). As a result, the error bars are not visible in the graphs (see Table 1 for standard error values). (B) Histograms showing the telomere length frequencies for p-arms, q-arms, or the sum of p+q-arms of primary MEFs grouped by genotype. The letters indicate the individual MEFs of each genotype used for the analysis. n is the total number of p-telomeres, q-telomeres, or the sum of p+q-telomeres per genotype that are represented in each histogram. One TFU corresponds to 1 kb of TTAGGG repeats (30). To facilitate visualization of the telomere length values, two vertical lines indicate the position of the 20- and 60-kb telomeres for each histogram. The telomere length frequency distribution in each histogram is an indication of the standard deviation of telomere length values and not of the standard error. (C) Percentage of telomeres of the total number of telomeres analyzed for each genotype that are ≤20 kb or ≥60 kb, as indicated. The absolute numbers of telomeres used for the analysis are also shown on top of the bars.