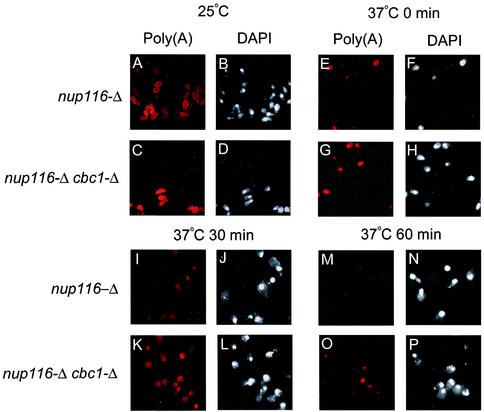
FIG. 2.
FISH analyses revealing that total poly(A)+ RNA in nup116-Δ and nup116-Δ cbc1-Δ strains are retained in the nucleus and that there is less degradation in the nup116-Δ cbc1-Δ strain. The isogenic pairs of strains were grown at 25°C to the mid-logarithmic phase of growth. Subsequently, one-half of each culture was transferred to the restrictive temperature of 37°C; the cultures were further incubated for one additional hour at both temperatures. Transcription of the cells in the culture shifted to 37°C was inhibited by the addition of thiolutin (4 μg/ml); the cells were harvested at various times after transcription block as indicated on top of each panel in the figure and subsequently fixed and processed for FISH and DAPI analysis as described in Materials and Methods. Left panels of mock shifted as well as each time point after transcription block show the localization and decay of the total poly(A)+ RNA as visualized using Cy3-labeled oligo(dT) are denoted as Poly(A), whereas right panels of respective time points show the nuclear DNA as visualized using DAPI staining denoted as DAPI. The time indicated at the top of each panel represents the time after transcription block to after shift to 37°C. See the Results (“Existence of DRN, a Cbc1p-dependent nuclear mRNA degradation pathway: cytological evidence”) for details of each panel.