Figure 3.
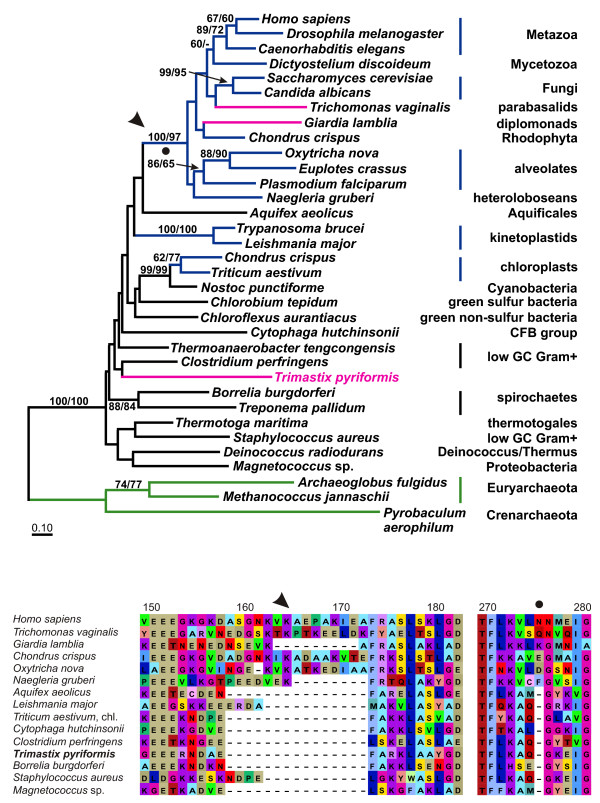
A – ML tree of PGK protein sequences. Maximum likelihood tree from PGK sequences of eukaryotes, eubacteria and archaebacteria (green labelling) based on 277 aligned aa positions. Color coding and labelling is as in Figures 1A and 2A. Possible chloroplast isoforms (Chondrus and Triticus) have not been directly localized, but their branching with cyanobacteria receives high bootstrap support. The presence of a surface loop (see arrowhead) and a 1 aa insertion (see black dot) unites all eukaryotes with the exception of kinetoplastids, Trimastix and the chloroplast homologs (see also Fig. 3B). For display purposes the tree has been rooted with archaebacteria. B – Alignment of deduced PGK amino acid sequences. Two sections of a PGK alignment are shown from eukaryotes and eubacteria. The first section (position 150–183) shows an insertion which produces a typical eukaryotic surface loop in the PGK protein (see also Fig. 3A) whereas the second section shows a 1 aa insertion at position 276. Both these features are not present in Trimastix, the kinetoplastids and the chloroplast homologs excluding these from the well supported remaining eukaryotes.