Figure 1.
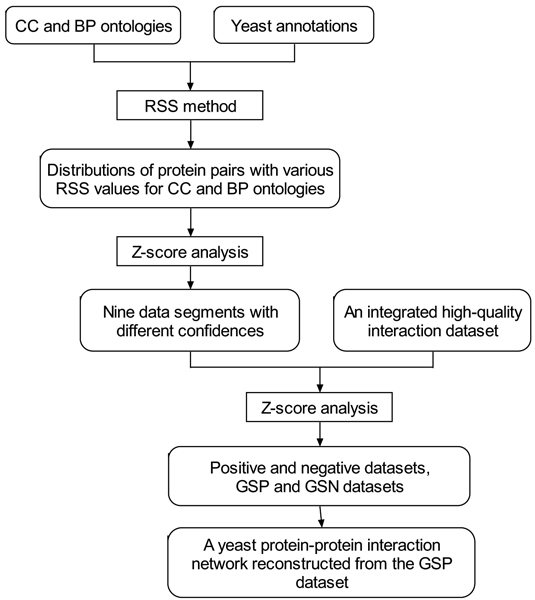
Schematic representation of the GO-based prediction of protein-protein interaction network. It includes four steps: (1) Using both CC and BP ontologies and their respective annotations, the distributions of protein pairs with various RSS values for the two ontologies were obtained. (2) Then a Z-score analysis was applied to draw the statistical significance of the quality scoring system and the nine data segments with different combinations of confidences were obtained. (3) To evaluate the RSS method, an integrated high-quality interaction dataset was applied. Based on a Z-score analysis, a positive dataset and a negative dataset were selected. Moreover, a gold standard positive (GSP) dataset with the highest level of confidence and a gold standard negative (GSN) dataset with the lowest level of confidence were derived. (4) Finally, a yeast protein-protein interaction network was reconstructed from the GSP dataset. The RSS method, a new metric for semantic similarity to score the degree of the functional association between two different proteins by comparing the relative specificity of pairs of GO terms assigned to them in similarity within a GO DAG [21].
