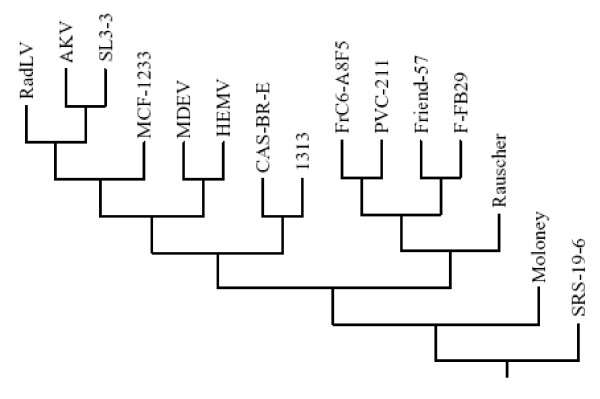
Figure 4.
Phylogenetic analyses of full Length MuLV genomes. Full-length nucleotide sequences as well as deduced amino acid sequences of 15 MuLV strains including MuLV-1313 were aligned and all gaps were stripped from the alignments before the phylogenetic trees were constructed and bootstrapping was set at 1000. Phylogenetic analyses were performed using PHYLIP [66, 71]. PHYLIP packages SEQBOOT, PROTDIST, DNADIST, NEIGHBOR, CONSENSE, and DRAWGRAM. The original data set was first analyzed by SEQBOOT which produced 100 bootstrapped data sets. The distance matrices on these data sets were achieved using PROTDIST for amino acid sequences and DNADIST for nucleotide sequences. The distance matrices were joined using NEIGHBOR. The tree files from NEIGHBOR were then applied with CONSENSE and the consensus tree was drawn using DRAWGRAM. Multiple sequence alignment were made using Vector NTI (Invitrogen, Carlsbad, California) with default gap opening penalty of 15 and default gap extension penalty of 6.66. Full length genomes used in the construction of the dendrogram included; AKV MuLV (J01998), MuLV 1313 (AF411814), Cas-Br-E MuLV (X57540), Friend-57 MuLV (X02794), Friend FB29 MuLV (Z11128), Friend PVC211 MuLV (M93134), Friend (FrC6-A8F5 D88386), mink cell focus-forming virus 1233 (MCF1233, U13766), Moloney MuLV (J02255), radiation leukemia virus (RadLV, K03363), Rauscher MuLV (Rauscher, U94692), SL3-3 MuLV (AF169256), solid-type reticulum cell sarcoma 19-6 MuLV (SRS 19-6, AF019230), HEMV (AY818896) and MDEV (AF053745). Note that both the amphotropic MuLV-1313 and ecotropic Cas-Br-E MuLV of the Southern California feral mice arise from a separate node of the phylogenetic tree indicating their evolutionary relationship.