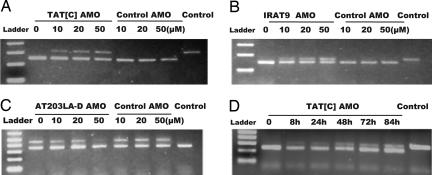
Fig. 2.
RT-PCR to demonstrate restoration of ATM mRNA splicing in A-T cell lines after AMO treatment. For A–C, cells were treated with different concentrations of the cognate AMO and a control AMO for 24 h, RNA was isolated, and RT-PCR was performed. Concentrations of AMOs are indicated above the lanes. RNA from normal cells was used as the control for correct splicing (last lanes). As a specificity control, IRAT9-AMO was used as the control for TAT[C] and AT203LA cells; TAT[C]-AMO was used as the control for IRAT9 cells. (A) TAT[C]-AMO was used to treat TAT[C] cells. For RT-PCR, the forward primer was placed at the junction of exon 54 and exon 55; the reverse primer one was in exon 57. The PCR product sizes of the wild-type (corrected) and mutant transcripts were 309 bp and 245 bp, respectively. (B) IRAT9-AMO was used to treat IRAT9 cells. Forward and reverse primers were located in exons 7 and 8, respectively. The PCR product sizes of the wild-type (corrected) and mutant transcripts were 170 bp and 148 bp, respectively. (C) AT203LA-D was used to treat AT203LA cells. Forward and reverse primers were placed in exons 28 and 29, respectively. The PCR product sizes of the wild-type and mutant transcripts were 344 bp and 456 bp, respectively. (D) Time–effect response of TAT[C]-AMO on TAT[C] cells. Cells were incubated with 30 μM TAT[C] AMO. RNA samples were collected at 8, 24, 48, 72, and 84 h and analyzed by RT-PCR.