Abstract
Age is a critical determinant of an adult female mosquito's ability to transmit a range of human pathogens. Despite its central importance, relatively few methods exist with which to accurately determine chronological age of field-caught mosquitoes. This fact is a major constraint on our ability to fully understand the relative importance of vector longevity to disease transmission in different ecological contexts. It also limits our ability to evaluate novel disease control strategies that specifically target mosquito longevity. We report the development of a transcriptional profiling approach to determine age of adult female Aedes aegypti under field conditions. We demonstrate that this approach surpasses current cuticular hydrocarbon methods for both accuracy of predicted age as well as the upper limits at which age can be reliably predicted. The method is based on genes that display age-dependent expression in a range of dipteran insects and, as such, is likely to be broadly applicable to other disease vectors.
Keywords: Aedes aegypti, age-grading, gene expression, cuticular hydrocarbons, multivariate calibration
Mosquito-transmitted diseases, such as malaria and dengue, continue to be major causes of human morbidity and mortality throughout the world. The pathogens that cause these diseases require a period to replicate and disseminate in the mosquito's body before transmission to naïve hosts can occur. The period between when a mosquito blood feeds on an infected host and when it is able to transmit the infection is termed the extrinsic incubation period (EIP). The EIP is estimated to be between 10 and 14 days for dengue virus (1, 2) and 9 and 14 days for malaria (3). Not all female mosquitoes start feeding immediately after they eclose, therefore, a vector must live longer than the duration of the initial nonfeeding period (usually <2 days) plus the EIP of the pathogen to contribute to disease transmission. This combined period (>12 days) is quite long relative to adult mosquito lifespan and, consequently, mosquito population age structure is a critical determinant of a vector population's capacity for pathogen transmission (4).
Despite the central importance of insect population age structure to the epidemiology of these diseases, relatively few methods have been developed that can directly and accurately measure insect age in the field (5–9). Existing methods for age-grading insects, such as scoring modifications to ovarian anatomy, are laborious and discriminate only very young adult age classes. Pteridine concentration has been shown to predict age in various dipteran species (10–12), and although its initial application to predicting mosquito age showed promise (13), it has proven to be generally unreliable for wild mosquitoes because pteridines occur in limited quantities and their concentration fluctuates with blood feeding (9). In the case of the primary dengue vector, Aedes aegypti, the most recently developed age-grading method utilizes changes in proportions of cuticular hydrocarbons (CHC) from legs to predict adult age (6, 14). This method has been shown to be capable of estimating adult Ae. aegypti age up to 15 days in field evaluations (6). Given that many individuals live >15 days in the field and that dengue transmission primarily occurs in mosquitoes 12 days and older, this approach has limited utility in epidemiological investigations. Considering that a number of research groups are proposing new control strategies for these diseases that explicitly target mosquito lifespan (15–17), there is a growing need for tools to reliably measure mosquito age under field conditions.
Recent gene expression studies in Drosophila (18–21), anopheline mosquitoes (22), nematodes (23), mice (24), and monkeys (25) have demonstrated consistent age-dependent changes in global gene transcription profiles. Drosophila melanogaster microarray studies have shown that expression of 2–9% of the genome changes significantly with age (19–21). Results from similar studies in Anopheles gambiae estimate that transcription of ≈5% of the genome varies with age (22). These changes in transcription have the potential to be used as markers for determining insect age in the field. Using D. melanogaster transcriptional profiles (18), we identified a set of genes showing significant changes in expression across adult female life stages. Orthologues of these genes were isolated from Ae. aegypti and a quantitative reverse transcriptase PCR (qRT-PCR) assay developed to determine chronological age of female mosquitoes under field conditions. Comparison of this approach to the existing CHC method indicates that it provides greater accuracy and precision over a broader range of adult age classes.
Results and Discussion
Transcriptional profiling studies in D. melanogaster have demonstrated that expression levels of a number of genes change significantly with adult age (18, 20, 21). On examination of these data sets, we selected a set of genes for analysis in mosquitoes based on the following criteria: (i) displayed large variation in gene expression across adult age classes, (ii) expression was unlikely to be influenced by digestive or reproductive status, and (iii) orthologues were known to exist in Aedes mosquitoes. Once a suitable set of candidate genes was identified, PCR primers for Ae. aegypti orthologues of these genes were designed for qRT-PCR assays (Table 1).
Table 1.
D. melanogaster genes and their orthologues in Ae. aegypti that showed age-related changes in transcription
| Drosophila gene | Putative function | Ae. aegypti orthologue | Forward primer (5′–3′) | Reverse primer (5′–3′) | Amplicon size, bp |
|---|---|---|---|---|---|
| RpS17 (CG3922) | Structural ribosomal protein | Ae-RpS17(AY927787) | CACTCCCAGGTCCGTGGTAT | GGACACTTCCGGCACGTAGT | 81 |
| fizzy (CG4274) | Cell cycle, cell physiology | Ae-4274(TC6602) | GGACGCTTAGCGGGAAGAC | TTGGCGTTTGGGATTTACCT | 81 |
| CG4679 | — | Ae-4679(TC66515) | GGAGGCTATAACCATCCGAGAA | GGCCAAGGACGGTTCGAT | 81 |
| me@31B (CG4916) | Nucleic acid metabolism | Ae-4916(TC54107) | ACATTCCCACTGAGCGTGAAG | GGTGAGCTCCTCCATCAGGTT | 81 |
| CG6639 | — | Ae-6639(AY431255) | CCACGCGAATATCGATGATG | AGGGAACCTGCGTCCCATAC | 81 |
| Rpd3 (CG7471) | Histone methylation | Ae-7471(TC62395) | CGAGACGATCTCCCGTTAGG | TTTCGAAGGTGATGGAGACCTT | 81 |
| CG8505 | Structural component of cuticle | Ae-8505(AY432732) | CTCCCTGTGGTTGGAATCCA | TAACCTACACCGCCCCAGAT | 81 |
| CG12750 | Protein biosynthesis | Ae-12750(TC63489) | ACCCATCGACTGGACACGAA | CCATCCGATAGAGTCGGTGAA | 81 |
| Scp1 (CG15848) | Calcium-binding protein | Ae-15848(TC59614) | TGACGGCATCCTTGAACTCTT | AGGCTCTGTGGGATGAGATCTC | 84 |
Gene function was inferred from Drosophila gene ontologies. Ae. aegypti orthologues were found by two-way BLAST searches, and numbers in parentheses indicate either GenBank or TIGR Gene Indice sequence accession numbers. Primer sequences for the genes used for transcriptional age-grading of adult female mosquitoes are shown. Ae. aegypti orthologues in bold text were found to be most informative for predicting mosquito age.
Initial evaluations of the suitability of these genes for age grading was made on laboratory-reared female mosquitoes. Adult female mosquitoes aged up to 29 days were used to determine whether transcription of the selected orthologues was consistent with that seen for D. melanogaster (18). Transcriptional profiles were quantified from the head and thorax of individual mosquitoes. The abdomen was removed to limit inclusion of gut and reproductive tissues because blood feeding is known to induce transcriptional changes in these tissues (26, 27). To identify genes with age-dependent changes in expression, a reference gene was used to normalize samples. A requirement for this gene was that its transcription displayed minimal variation with age. Three Ae. aegypti genes, β-tubulin, actin-1, and ribosomal protein S17 (Ae-RpS17), were screened for this purpose by using qRT-PCR. Of the three, Ae-RpS17 was most suitable because it displayed minimal variation in expression across adult female age classes (Fig. 3, which is published as supporting information on the PNAS web site). Using Ae-RpS17 for normalization, relative changes in transcript representation of each gene was determined as a log contrast of cycle threshold (Ct) values. All eight Ae. aegypti orthologues tested showed age-related transcriptional profiles (Fig. 4, which is published as supporting information on the PNAS web site) that were comparable with results from D. melanogaster microarray studies (18). Of these eight genes, two showed large decreases in transcript abundance with age (Ae-8505 and Ae-15848) (Fig. 4). This decrease is represented as an increase in log contrasts of Ct values. The other six genes (Fig. 4) showed smaller changes across adult female mosquito age classes.
Having identified eight genes with age-dependent expression profiles from laboratory-reared Ae. aegypti, we investigated their transcription in mosquitoes reared and maintained under field conditions. This field study aimed to predict mosquito age from these transcript profiles and validate these estimates by direct comparisons with age estimates made from analysis of CHCs from the same specimens (6). Mosquitoes were reared under ambient conditions and offered a human blood meal every 4 days, and adults were held in large field cages in typical habitat in suburban Cairns, Australia. Mosquitoes of known age were sampled from these cages periodically up to 21 days, at which point cages were depleted of mosquitoes. Legs were removed for quantification of cuticular hydrocarbons, whereas the head and thorax were used to determine transcript abundance. Relative changes in expression of the eight genes of interest were comparable to results from the initial laboratory study (Fig. 4), indicating that fluctuating environmental conditions and blood feeding had little impact on expression of the selected genes. Transcriptional profiles were analyzed by using a multivariate calibration approach to determine whether adult mosquito age could be predicted reliably from these indirect measures.
Multivariate calibration problems involve situations in which particular traits cannot be measured, so other indirect traits that can be measured are used to derive estimates of the trait of interest. The relationship between the immeasurable traits and indirect measures was determined by using a training data set containing measures of both values. In this study, mosquito age was the unknown parameter, and we used multiple gene expression (GE) profiles or CHC measurements to establish a calibration model with age. In particular, this was a controlled calibration problem where we selected the values of the unknown trait (mosquito age) from which the training data were constructed (either GE or CHC measures) (28). This defined mosquito age as the explanatory variable and a vector of either GE or CHC measures as the response variable. Consequently, the calibration model was constructed from the regression of GE or CHC measures on age. Previous work (6, 14) has used multivariate regression by swapping the positions of the explanatory and response axes to determine age predictions, an approach that has been shown to be inappropriate for calibration problems (see ref. 28 for discussion), and, hence, age predictions require inverse regression of the calibration model.
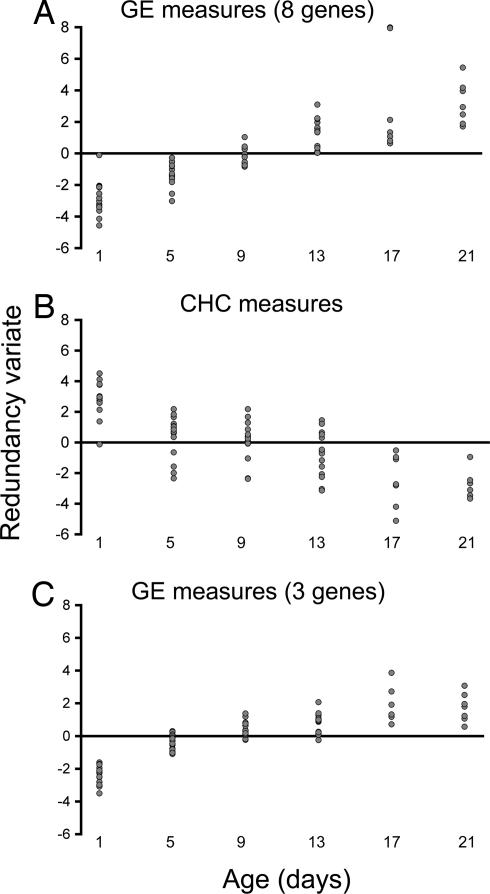
Canonical redundancy analysis (29) was used to reduce the dimensionality of the multivariate GE or CHC measures by calculating the linear combination of GE or CHC measures that maximized the correlation with mosquito age (termed a redundancy variate). Our analysis of transcript profiles from field-reared mosquitoes showed that 87.03% of the variance in gene expression was explained by age. Importantly, the first redundancy variate extracted from this analysis showed a linear association with mosquito age (Fig. 1A, n = 73, R2 = 0.718; P < 0.001), showing that chronological age can be estimated from training data established under field conditions. In contrast, redundancy analysis on CHC measures taken from the same individual mosquitoes showed that 68.94% of the variance in CHC abundance was explained by age. The first redundancy variate showed a negative linear association with age (Fig. 1B, n = 73, R2 = 0.608; P < 0.001). It is evident from the amount of variance explained in the two models that changes in gene expression are more closely associated with adult age than changes in CHC composition. When all GE and CHC measures are analyzed together, 89.66% of the variance in these measures is explained by age. The subsequent redundancy variate from this analysis has a positive linear association with age (n = 73, R2 = 0.77; P < 0.001) (data not shown).
Fig. 1.
Calibration data from individual mosquitoes. Calibration models were developed from adult female Ae. aegypti of known age. Log-contrast normalized measures of GE (A and C) and CHC (B) abundance were entered into separate canonical redundancy analyses. Each analysis produced a redundancy variate that is the linear combination that maximizes correlation with mosquito age. (A and C) The calibration model is based on expression data from eight genes (Ae-4274, Ae-4679, Ae-4916, Ae-6639, Ae-7471, Ae-8505, Ae-12750, and Ae-15848) (A), and the calibration model restricted to the three most informative genes (Ae-4274, Ae-8505, and Ae-15848) (C).
Having established a calibration model from the training data, we predicted ages by using blind samples. Blind samples were collected from field cages at a 2-day interval, and both transcript and CHC abundance were quantified for each mosquito, allowing direct comparisons of the age predictions derived from these two different measures. Canonical redundancy analysis was used to calculate a redundancy variate for both transcriptional and CHC measures. Inverse regression of CHC or GE calibration models from this redundancy variate were used to estimate mosquito age. The accuracy of both transcriptional and CHC age-grading methods was assessed by comparing age predictions with the known age for each blind sample. Estimation of the precision of predicted age was not possible by using feducial limits (28). Consequently, a nonparametric bootstrapping procedure was developed to derive 95% confidence intervals for the predicted age of individual mosquitoes, based on both GE and CHC measures (see Table 2 and Supporting Text, which are published as supporting information on the PNAS web site).
Comparison of the two methods showed that transcriptional profiles (Fig. 2A) gave more accurate age predictions than CHC measures (Fig. 2B). The accuracy of age prediction can be established from the divergence of the age prediction from actual age (Fig. 2 A and B). In general, age estimates based on gene expression data fall within ±5 days of the actual age. This range increases to approximately ±10 days for CHC-based predictions of age (Fig. 2) as a consequence of the weaker association between CHCs and age. It should be noted that negative ages can be predicted because of error associated with both GE and CHC measures. Age predictions from combined GE and CHC measures were comparable to predictions from transcriptional profiles alone (data not shown).
Fig. 2.
Age predictions for individual mosquitoes. Age predictions were made for additional blind samples by using both GE (A and C) and CHC (B) measures. Log-contrast normalized GE and CHC measures were calculated from each mosquito and entered into a redundancy analysis. Age predictions were estimated by inverse regression of the respective calibration model shown in Fig. 1. Error bars indicate 95% CI for individual age predictions as determined by nonparametric bootstrap methods. The dashed line indicates where predicted age equals actual age. Age predictions derived from the calibration model indicated in Fig. 1C based on a reduced data set of the three most informative genes (C).
Any assay that aims to predict the age of field-caught insects has to be insensitive to fluctations in environmental conditions. By developing our calibration model from mosquitoes maintained under ambient environmental conditions and provided with human blood meals, variability in gene expression due to these factors is incorporated into the assay. It is yet to be determined how robust the calibration models derived from these individuals are across seasons or regions. However, the observation of comparable transcriptional trends between our laboratory and field studies (Fig. 4) suggest that the described calibration models may be resilient to environmental fluctuations.
We have demonstrated that age determination of adult Ae. aegypti can be accomplished with specimens reared under field conditions to a much higher degree of accuracy by using transcriptional profiles of a selection of mosquito genes than by previous methods. However, researchers undertaking large-scale studies could consider using a reduced set of genes for age determination. The three most informative genes were Ae-15848, Ae-8505, and Ae-4274, and analysis showed that 74.99% of the variance in their expression was explained by age. The first redundancy variate had a linear association with age (Fig. 1C, n = 73, R2 = 0.734; P < 0.001); however, bias in the calibration of the model resulted in a slight overestimation of age in most age classes (Fig. 2C). Although this is undesirable, the significant simplification in the assay may justify tolerating this bias. By excluding the five least informative genes, it may be possible to develop a multiplex assay that would allow high-throughput examination of mosquito samples. This approach, although not as accurate as the eight-gene model, is still superior to the existing CHC method under the experimental conditions used.
The GE age-grading approach presented here represents a substantial improvement over existing methods to estimate adult mosquito age. Its use will facilitate future empirical approaches to understand the contribution of mosquito age to dengue transmission dynamics, both temporally and geographically. It also provides a simple and accurate method to evaluate existing vector control methods that target adult mosquitoes and will be of particular relevance for evaluating the efficacy of proposed control strategies that target mosquito lifespan (15–17, 30). Considering that the two most informative genes used in our study also show strong age-related transcriptional trends in D. melanogaster (18) and An. gambiae (22), it is likely that this approach will be able to be generalized across species.
Materials and Methods
Rearing of Mosquitoes.
Initial experiments investigating age-related transcriptional trends of Ae. aegypti genes were conducted with mosquitoes reared under laboratory conditions. Subsequent experiments were conducted with material reared and held under field conditions. Mosquitoes for each of these sets of experiments were reared as follows:
Laboratory studies.
Ae. aegypti eggs were obtained from a colony established from eggs collected in Cairns (Queensland, Australia) during 2005. Eggs were hatched in deoxygenated water, and larvae were reared at ≈100 larvae per liter on dried chicken feed pellets at 26°C. Adult mosquitoes were maintained in 27-cm3 cages placed in plastic bags, with damp cotton wool to maintain elevated humidity (28°C, ≈80% relative humidity, 12:12 h light:dark). Laboratory-reared adult mosquitoes were not supplied with blood meals, only 10% sucrose solution. Adult female mosquitoes were collected at 1 (0–24 h), 5, 9, 13, 17, 21, 25, and 29 days after eclosion. Adults emerging during each 24-h period were considered a single age class. Females were frozen in liquid nitrogen and stored at −80°C.
Field studies.
Field studies were undertaken in Cairns, Queensland, Australia during the wet season from January to February 2005. Field-collected eggs (G0) were hatched in hay-infused water, and resulting adults held through several gonotrophic cycles to generate enough eggs to stock field cages. G1 eggs were hatched by using hay-infused water and reared at the field site on a diet of dry adult cat food (Friskies; North Ryde, New South Wales, Australia) under ambient conditions and low densities to ensure synchronous development. Wing lengths of reared mosquitoes were measured to ensure that mean mosquito size, and, hence, nutritional status were comparable from field caught mosquitoes (31). Female wing lengths from this study (mean ± SE = 2.66 ± 0.01 mm) were not appreciably different from that reported previously for wild female Ae. aegypti from Cairns (2.76 ± 0.03 mm) (32). Adult mosquitoes were housed in two large field cages (13 m3 dome tents, ≈4,000 mosquitoes per cage) positioned in two representative habitats: (i) a room underneath an elevated house and (ii) outside a house under an open-sided shelter. Mosquitoes were supplied with 10% sucrose solution and offered a human blood meal every 4 days (human ethics approval from James Cook University H2250). Adult resting sites (e.g., tires and boxes) and oviposition sites were included in field cages. Ambient temperature and humidity were measured at each cage by using data loggers (Fig. 5, which is published as supporting information on the PNAS web site). Mean temperatures for the indoor and outdoor cages were 27.6°C (minimum, maximum: 24.9, 30.3°C) and 27.4°C (23.2, 32.4°C), respectively. Mean relative humidity was 78.0% (minimum, maximum: 54.6, 93.3%) and 78.7% (46.3, 96.7%) for the indoor and outdoor cage, respectively. Mosquitoes were collected within 24 h after eclosion and every 4 days thereafter. Additional samples for blind experiments were collected every 2 days. Mosquitoes were processed on a chill table; legs were removed and stored at −80°C for GC-MS analysis, and individual mosquito heads and thoraces were dissected and then stored in 125 μl of RNAlater (Ambion) to stabilize RNA transcripts for qRT-PCR analysis. Samples in RNAlater were maintained at 4°C for 24 h before storage at −20°C.
Identification of Putative Mosquito Genes with Age-Dependent Expression.
Potential genes of interest were identified from data generated in prior microarray studies of gene expression in adult female D. melanogaster (18). Gene candidates were selected on the basis of showing strong age-related expression trends. Ae. aegypti gene orthologues were identified by two-way BLAST searches of nucleotide and protein sequences from GenBank (www.ncbi.nlm.nih.gov) and TIGR's Ae. aegypti gene indices (www.tigr.org/tdb/tgi). Gene-specific primer sets (Table 1) were designed by using PrimerExpress version 3 (Applied Biosystems).
Total RNA Isolation.
Individual female heads and thoraces were dissected, and total RNA was extracted by using TRIzol reagent (Invitrogen) by following standard protocols with minor modifications: insect tissues were frozen in liquid nitrogen and ground dry with a micropestle before the addition of TRIzol reagent, and an additional chloroform wash was done before total RNA was precipitated with 8 M LiCl at −30°C overnight. RNA was resuspended in 25 μl of RNase-free water.
Total RNA was quantified by using spectrophotometric absorbance readings. One microgram of total RNA was treated with 1 unit of DNase I (Roche) in 10× RQ1 buffer (Promega) for 30 min at 37°C, then the reaction was terminated with RQ1 Stop solution (Promega, Madison, WI). RT was performed by using 500 ng of DNase-treated total RNA and oligo(dT)15 (Promega) priming with SuperScript III RNase H− (Invitrogen) by following the manufacturer's protocols. RT reactions were treated with RNase H− (Roche) before being diluted 5-fold to remove PCR inhibitors. The remaining 500 ng of RNA was used as a negative RT control (no reverse transcriptase) and screened for genomic DNA contamination by using standard PCR with Ae-RpS17 primers (95°C for 3 min; 95°C for 15 s; 58°C for 15 s; 72°C for 30 s; 35 cycles; 72°C for 5 min). PCR products were separated by gel electrophoresis, stained with ethidium bromide, and visualized under UV illumination.
Selection of Reference Gene.
Gene expression data are commonly normalized to a reference gene to remove variation associated with variable amounts of starting tissue, RNA integrity, and experimental treatment. Previous studies (33, 34) have shown the need to validate reference genes to ensure that they are not influenced by experimental treatment. We designed primers for several potential reference genes from Ae. aegypti; actin-1 (Aaeact-1, U20287), β-tubulin (TIGR GI TC66155) and ribosomal protein S17 (Ae-RpS17; AY927787) and quantified transcript abundance across adult age in laboratory-reared mosquitoes. PCR products were amplified from Ae. aegypti cDNA, gel purified and cloned into pGEM-Teasy (Promega), and confirmed by sequencing. qRT-PCR was performed on a Lightcycler (Roche) by using Platinum SYBR Green I Supermix (Invitrogen) and the following cycling conditions: 50°C for 2 min; 95°C for 2 min, then 95°C for 5 s; 60°C for 5 s; 72°C for 10 s; for 50 cycles; with fluorescence acquisition at the end of the extension step. Product specificity was confirmed by melting curve analysis. Linearized plasmids containing inserts of each reference gene were amplified in triplicate and used to construct standard curves with 105-fold dynamic range. Absolute quantification of reference gene transcripts were made from their respective standard curve by using Lightcycler software (version 3.5; Roche).
Determining Transcript Abundance with qRT-PCR.
Transcript abundance of the eight candidate genes and Ae-RpS17 was determined for individual mosquitoes. qRT-PCRs were performed on the Lightcycler (Roche) by using Platinum SYBR Green I Supermix (Invitrogen) according to the manufacturer's instructions for glass capillary thermal cyclers. Cycling conditions for all templates were as reported above for the reference gene, and PCR product specificity was determined by melting curve analysis. Triplicate reactions were run for each primer set (Table 1) across all templates. The Ct was determined for each reaction by using the second-derivative maximum method (Roche). Mean Ct values were calculated, and relative changes in transcript abundances between individuals were normalized to Ae-RpS17 by log contrasts (35).
Mosquito Cuticular Hydrocarbon Extraction and Quantification.
CHCs were extracted from all legs of individual field-reared mosquitoes and quantified by GC-MS based on methods described in Hugo et al. (36). Peak areas of four CHC measures of interest (C25H52, C26H54, C27H56, and C28H58) and a reference measure (C29H60) were quantified. Relative changes in peak area were normalized to C29H60 by calculating log contrasts.
Multivariate Calibration and Indirect Regression Methodologies.
A multivariate calibration framework was developed to estimate mosquito age from GE and CHC measures. A training data set was established, for both GE and CHC measures, by quantifying each phenotypic measure from mosquitoes of known age. Relative changes in GE and CHC measures from each individual were calculated as log contrasts (35). This approach was used to remove the unit sum constraint associated with compositional data. Log contrasts were calculated as follows:
where Xi equals the mean Ct value for a particular gene (or area of a CHC peak), Xref equals the mean Ct values of the reference gene (or peak area for reference CHC), and Xtotal equals the sum of Ct values for all genes from an individual (or total of all CHC peak areas).
Because some CHC measures were absent for certain individuals, log contrasts were calculated with the addition of 1 to eliminate missing values from the CHC data set. Training data were entered into a canonical redundancy analysis to reduce the dimensionality of the regressions between multiple GE or CHC measures and age. Linear regression of the first redundancy variate on age was used for each calibration model. Age estimates then were derived from this calibration model by inverse regression of new GE or CHC measures.
Additional samples were processed and used to evaluate the reliability of both GE and CHC calibration models. A nonparametric bootstrapping method was used to assess sampling error in the calibration model procedure (see Supporting Text and Table 2). Data sets were standardized (N ≈ 0,1) before undertaking the bootstrapping procedure. Field cage samples were bootstrapped by resampling individuals within cages and age classes with replacement, thereby generating 1,000 pseudorandom data sets. A canonical redundancy analysis was performed on each data set to calculate a single redundancy variate that was a linear combination of phenotypic values that maximized the correlation with age. Each individual within each bootstrap replicate was scored for its value of the redundancy variate. The relationship between the redundancy variate and age was predicted by using simple linear regression, producing 1,000 regression equations. These models then were used to score mosquitoes that had been sampled from field cages as effective blind samples for age estimation. Phenotypes of these samples were also standardized (N ≈ 0,1) before analysis. Each individual was scored for 1,000 bootstrap redundancy variates and ages predicted according to the 1,000 regression equations from the corresponding redundancy variates. The top and bottom 2.5% of predicted age values represent the 95% confidence intervals for age. For all numerical procedures, we used SAS (version 9.1; SAS Institute, Cary, NC); see Supporting Text for SAS editor syntax.
Supplementary Material
Acknowledgments
We thank the staff at Cairns Tropical Population Health Unit, especially Sharron Long for her assistance with field work; Geoff Eaglesham and Neil Holling from Queensland Health Scientific Services for technical assistance with gas chromatography mass spectroscopy analysis; and Dr. Jeremy C. Brownlie for his constructive comments on drafts of the manuscript. This research was funded by Australian Research Council Grant LP0455732 and grants from the Foundation for the National Institutes of Health through the Grand Challenges in Global Health Initiative and the Queensland Government's Smart State Ph.D. funding initiative.
Abbreviations
- CHC
cuticular hydrocarbons
- Ct
cycle threshold
- GE
gene expression
- qRT-PCR
quantitative reverse transcriptase PCR.
Footnotes
The authors declare no conflict of interest.
This article is a PNAS direct submission.
References
- 1.Siler JF, Hall MW, Hitchens AP. Philipp J Sci. 1926;29:1–304. [Google Scholar]
- 2.Watts DM, Burke DS, Harrison BA, Whitmire RE, Nisalak A. Am J Trop Med Hyg. 1987;36:143–152. doi: 10.4269/ajtmh.1987.36.143. [DOI] [PubMed] [Google Scholar]
- 3.Gilles HM, Warrell DA. Essential Malariology. London: Arnold; 2002. [Google Scholar]
- 4.Dye C. Annu Rev Entomol. 1992;37:1–19. doi: 10.1146/annurev.en.37.010192.000245. [DOI] [PubMed] [Google Scholar]
- 5.Detinova TS. WHO Monograph No. 47. Geneva: World Health Org; 1962. [PubMed] [Google Scholar]
- 6.Gerade BB, Lee SH, Scott TW, Edman JD, Harrington LC, Kitthawee S, Jones JW, Clark JM. J Med Entomol. 2004;41:231–238. doi: 10.1603/0022-2585-41.2.231. [DOI] [PubMed] [Google Scholar]
- 7.Hayes EJ, Wall R. Physiol Entomol. 1999;24:1–10. [Google Scholar]
- 8.Tyndale-Biscoe M. Bull Entomol Res. 1984;74:341–377. doi: 10.1017/S0007485300042358. [DOI] [PubMed] [Google Scholar]
- 9.Penilla RP, Rodriguez MH, Lopez AD, Viader-Salvado JM, Sanchez CN. Med Vet Entomol. 2002;16:225–234. doi: 10.1046/j.1365-2915.2002.00364.x. [DOI] [PubMed] [Google Scholar]
- 10.Hayes EJ, Wall R. Physiol Entomol. 1999;24:1–10. [Google Scholar]
- 11.Lehane MJ. Parasitol Today. 1985;1:81–85. doi: 10.1016/0169-4758(85)90048-1. [DOI] [PubMed] [Google Scholar]
- 12.Robson SK, Vickers M, Blows MW, Crozier RH. J Exp Biol. 2006;209:3155–3163. doi: 10.1242/jeb.02318. [DOI] [PubMed] [Google Scholar]
- 13.Wu D, Lehane MJ. Med Vet Entomol. 1999;13:48–52. doi: 10.1046/j.1365-2915.1999.00144.x. [DOI] [PubMed] [Google Scholar]
- 14.Desena ML, Edman JD, Clark JM, Symington SB, Scott TW. J Med Entomol. 1999;36:824–830. doi: 10.1093/jmedent/36.6.824. [DOI] [PubMed] [Google Scholar]
- 15.Blanford S, Chan BH, Jenkins N, Sim D, Turner RJ, Read AF, Thomas MB. Science. 2005;308:1638–1641. doi: 10.1126/science.1108423. [DOI] [PubMed] [Google Scholar]
- 16.Brownstein JS, Hett E, O'Neill SL. J Invertebr Pathol. 2003;84:24–29. doi: 10.1016/s0022-2011(03)00082-x. [DOI] [PubMed] [Google Scholar]
- 17.Rasgon JL, Styer LM, Scott TW. J Med Entomol. 2003;40:125–132. doi: 10.1603/0022-2585-40.2.125. [DOI] [PubMed] [Google Scholar]
- 18.Arbeitman MN, Furlong EEM, Imam F, Johnson E, Null BH, Baker BS, Krasnow MA, Scott MP, Davis RW, White KP. Science. 2002;297:2270–2275. doi: 10.1126/science.1072152. [DOI] [PubMed] [Google Scholar]
- 19.Landis GN, Abdueva D, Skvortsov D, Yang J, Rabin BE, Carrick J, Tavare S, Tower J. Proc Natl Acad Sci USA. 2004;101:7663–7668. doi: 10.1073/pnas.0307605101. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Pletcher SD, Macdonald SJ, Marguerie R, Certa U, Stearns SC, Goldstein DB, Partridge L. Curr Biol. 2002;12:712–723. doi: 10.1016/s0960-9822(02)00808-4. [DOI] [PubMed] [Google Scholar]
- 21.Zou S, Meadows S, Sharp L, Jan LY, Jan YN. Proc Natl Acad Sci USA. 2000;97:13726–13731. doi: 10.1073/pnas.260496697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Marinotti O, Calvo E, Nguyen QK, Dissanayake S, Ribeiro JM, James AA. Insect Mol Biol. 2006;15:1–12. doi: 10.1111/j.1365-2583.2006.00610.x. [DOI] [PubMed] [Google Scholar]
- 23.Lund J, Tedesco P, Duke K, Wang J, Kim SK, Johnson TE. Curr Biol. 2002;12:1566–1573. doi: 10.1016/s0960-9822(02)01146-6. [DOI] [PubMed] [Google Scholar]
- 24.Lee CK, Klopp RG, Weindruch R, Prolla TA. Science. 1999;285:1390–1393. doi: 10.1126/science.285.5432.1390. [DOI] [PubMed] [Google Scholar]
- 25.Kayo T, Allison DB, Weindruch R, Prolla TA. Proc Natl Acad Sci USA. 2001;98:5093–5098. doi: 10.1073/pnas.081061898. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Holt RA, Subramanian GM, Halpern A, Sutton GG, Charlab R, Nusskern DR, Wincker P, Clark AG, Ribeiro JMC, Wides R, et al. Science. 2002;298:129–149. [Google Scholar]
- 27.Sanders HR, Evans AM, Ross LS, Gill SS. Insect Biochem Mol Biol. 2003;33:1105–1122. doi: 10.1016/s0965-1748(03)00124-3. [DOI] [PubMed] [Google Scholar]
- 28.Sundberg R. Scand J Stat. 1999;26:161–191. [Google Scholar]
- 29.Rencher AC. Methods of Multivariate Analysis. New York: Wiley; 2001. [Google Scholar]
- 30.Sinkins SP, Gould F. Nat Rev Genet. 2006;7:427–435. doi: 10.1038/nrg1870. [DOI] [PubMed] [Google Scholar]
- 31.Nasci RS. J Am Mosq Control Assoc. 1986;2:61–62. [PubMed] [Google Scholar]
- 32.Russell RC, Webb CE, Williams CR, Ritchie SA. Med Vet Entomol. 2005;19:451–457. doi: 10.1111/j.1365-2915.2005.00589.x. [DOI] [PubMed] [Google Scholar]
- 33.Thellin O, Zorzi W, Lakaye B, De Borman B, Coumans B, Hennen G, Grisar T, Igout A, Heinen E. J Biotechnol. 1999;75:291–295. doi: 10.1016/s0168-1656(99)00163-7. [DOI] [PubMed] [Google Scholar]
- 34.Vandesompele J, De Preter K, Pattyn F, Poppe B, Van Roy N, De Paepe A, Speleman F. Genome Biol. 2002;3:research0034.1–0034.11. doi: 10.1186/gb-2002-3-7-research0034. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Aitchison J. The Statistical Analysis of Compositional Data. Caldwell, NJ: Blackburn; 2003. [Google Scholar]
- 36.Hugo LE, Kay BH, Eaglesham GK, Holling N, Ryan PA. Am J Trop Med Hyg. 2006;74:462–474. [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.