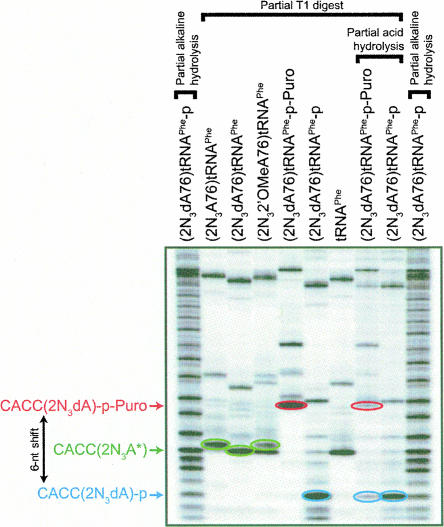
FIGURE 2.
Analysis of modified photoreactive tRNA ligands. (2N3A76)tRNAPhe, (2N3dA76)tRNAPhe, (2N32′OMeA76)tRNAPhe, (2N3dA76)tRNAPhe-p-Puro, (2N3dA76)tRNAPhe-p, and unmodified tRNAPhe, labeled with 32P as described in Materials and Methods, were subjected to digestion with RNase T1 under partial conditions to avoid overdigestion. In addition, (2N3dA76)]tRNAPhe-p-Puro and (2N3dA76)tRNAPhe-p were subjected to partial acid hydrolysis (60 min in 1% trifluoroacetic acid at 23°C) after T1 digestion. Partial alkaline digestion of (2N3dA76)tRNAPhe-p was used to generate a reference ladder. The samples were resolved by denaturing 25% PAGE and visualized by autoradiography. Labeled products released from the 3′ ends of the photoreactive tRNA ligands are CACC(2N3dA), CACC(2N32′OMeA), and CACC(2N3A) (collectively designated as CACC[2N3A*] in the figure), CACC(2N3dA)-p, and CACC(2N3dA)-p-Puro. A characteristic mobility shift of about 6 nt in the 3′ T1 RNase product is observed when (2N3dA76)tRNAPhe-p is condensed with puromycin. The observed 6-nt shift in (2N3dA76)tRNAPhe-p-Puro is reversed upon treatment with acid, consistent with hydrolysis of phosphoramidate linkage between (2N3dA76)tRNAPhe-p and puromycin (Benkovic and Sampson 1971; Welch et al. 1995).