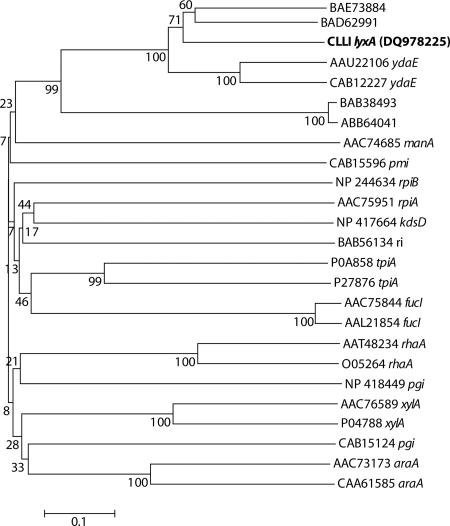
FIG. 5.
Phylogenetic tree of CLLI and other sugar isomerases. Bootstrap values are indicated at the branch points. The bar indicates a branch length equivalent to 0.1 change per amino acid. All sequences were obtained from GenBank. BAE73884, S. glossinidius conserved hypothetical protein; BAD62991, B. clausii conserved hypothetical protein; CLLI, C. laevoribosii d-lyxose isomerase (NCBI protein database accession no. DQ978225); AAU22106 ydaE, B. licheniformis ydaE-encoded protein; CAB12227 ydaE, B. subtilis ydaE-encoded protein; BAB38493, E. coli hypothetical protein; ABB64041, Shigella dysenteriae conserved hypothetical protein; AAC74685 manA, E. coli mannose-6-phosphate isomerase; CAB15596 pmi, B. subtilis phosphomannose isomerase; NP 244634 rpiB, Bacillus halodurans ribose-5-phosphate isomerase; AAC75951 rpiA, E. coli ribose-5-phosphate isomerase; NP 417664 kdsD, E. coli putative isomerase; BAB56134 ri, Acinetobacter sp. strain DL-28 l-ribose isomerase; P0A858 tpiA, E. coli triose phosphate isomerase; P27876 tpiA, B. subtilis triose phosphate isomerase; AAC75844 fucI, E. coli fucose isomerase; AAL21854 fucI, Salmonella enterica serovar Typhimurium LT2 fucose isomerase; AAT48234 rhaA, E. coli l-rhamnose isomerase; O05264 rhaA, B. subtilis l-rhamnose isomerase; NP 418449 pgi, E. coli glucose-6-phosphate isomerase; AAC76589 xylA, E. coli xylose isomerase; P04788 xylA, B. subtilis xylose isomerase; CAB15124 pgi, B. subtilis glucose-6-phosphate isomerase; AAC73173 araA, E. coli l-arabinose isomerase; CAA61585 araA, B. subtilis l-arabinose isomerase. Amino acid sequences were aligned with the Vector NTI AlignX software (Suite 9.0.0; Invitrogen, Carlsbad, CA). Phylogenetic trees were constructed by the neighbor-joining method (33), using the MEGA software, version 3.0 (25). The p-distance correction substitution model was used in a tree-building analysis. Bootstrap values were calculated based on 1,000 replicates of the data (11).