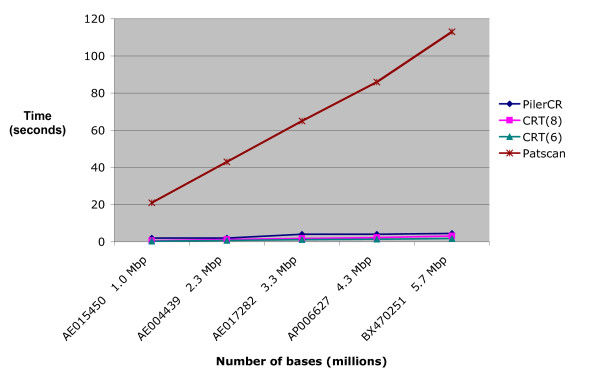
Figure 2.
Running time based on genome size, using repeat length 21–37 and spacer length 19–48. Running times for the three compared search tools, based on genome size (CRT is listed twice, once for windows size 6 and once for window size 8). The y-axis represents time in seconds. The x-axis lists the genome accession numbers, followed by their sizes in million base pairs (Mbp). As the size of the genomes increase, it can be seen that running times of the search tools increase at different rates. Below, the corresponding organism names are given. [IMG:AE015450] Mycoplasma gallisepticum (strain R(low)) [IMG:AE004439] Pasteurella multocida (strain Pm70) [IMG:AE017282] Methylococcus capsulatus (strain Bath/NCIMB 11132) [IMG:AP006627] Bacillus clausii (strain KSM-K16) [IMG:BX470251] Photorhabdus luminescens (subsp. laumondii, strain TT01).