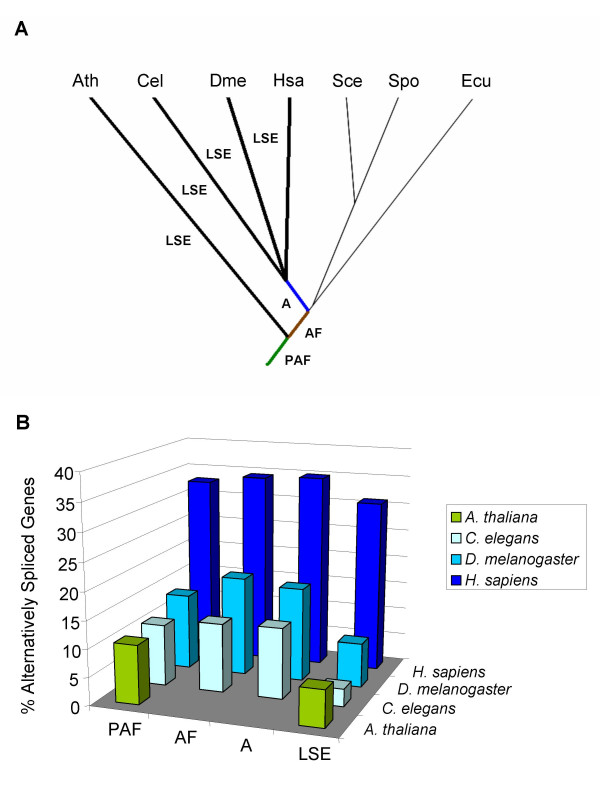
Figure 2.
Evolutionary origin of alternatively spliced genes. A: Phylogenetic tree showing the relations between the seven species included in the KOG database and used in this study. PAF (green line) corresponds to the group of KOG's likely appeared before the split of animals, plants and fungi; AF (brown line), KOG's likely appeared in the fungamal ancestor; A (blue line), clusters of orthologous likely appeared in the ancestors of nematodes, insects and vertebrates; LSE's (four wide black lines) correspond to lineage specific expansions of plants, nematodes, insects and vertebrates. B: Percentage of AS for genes according to phylogenetic origin. PAF: ancestral to plants, animals and fungi. AF: ancestor of animals and fungi. A: animals. LSE: lineage specific expansions. Hsa (Homo sapiens), Cel (Caenorhabditis elegans), Dme (Drosophila melanogaster), Ath (Arabidopsis thaliana), Sce (Saccharomyces cerevisae), Spo (Schizosaccharomyces pombe), Ecu (Encephalitozoon cuniculi). Note that in A. thaliana genes can only group into PAF or LSE.