Abstract
The trfA gene, encoding the essential replication initiation protein of the broad-host-range plasmid RK2, possesses an in-frame overlapping arrangement. This results in the production of TrfA proteins of 33 and 44 kDa, respectively. Utilizing deletion and site-specific mutagenesis to alter the trfA operon, we compared the replication of an RK2-origin plasmid in several distantly related gram-negative bacteria when supported by both TrfA-44 and TrfA-33, TrfA-33 alone, or TrfA-44/98L (a mutant form of the TrfA-44 protein) alone. TrfA-44/98L is identical to wild-type TrfA-44 with the exception of a single conservative amino acid alteration from methionine to leucine at codon 98; this alteration removes the translational start codon for the TrfA-33 protein. Copy number and stability were virtually identical for plasmids containing both TrfA-44 and TrfA-33 proteins or TrfA-44/98L alone in Pseudomonas aeruginosa and Agrobacterium tumefaciens, two unrelated bacteria in which TrfA-33 is poorly functional. This, along with recent in vitro studies comparing TrfA-44, TrfA-33, and TrfA-44/98L, suggests that the functional activity of TrfA-44 is not significantly affected by the 98L mutation. Analysis of minimal RK2 derivatives in certain gram-negative bacterial hosts suggests a role of the overlapping arrangement of trfA in facilitating the broad host range of RK2. RK2 derivatives encoding TrfA-44/98L alone demonstrated decreased copy number and stability in Escherichia coli and Azotobacter vinelandii when compared with derivatives specifying both TrfA-44 and TrfA-33. A strategy employing the trfA-44/98L mutant gene and in vivo homologous recombination was used to eliminate the internal translational start codon of trfA in the intact RK2 plasmid. The mutant intact RK2 plasmid produced only TrfA-44/98L. A small reduction in copy number and beta-lactamase expression resulted in E. coli, suggesting that overlapping trfA genes also enhance the efficiency of replication of the intact RK2 plasmid.
Full text
PDF
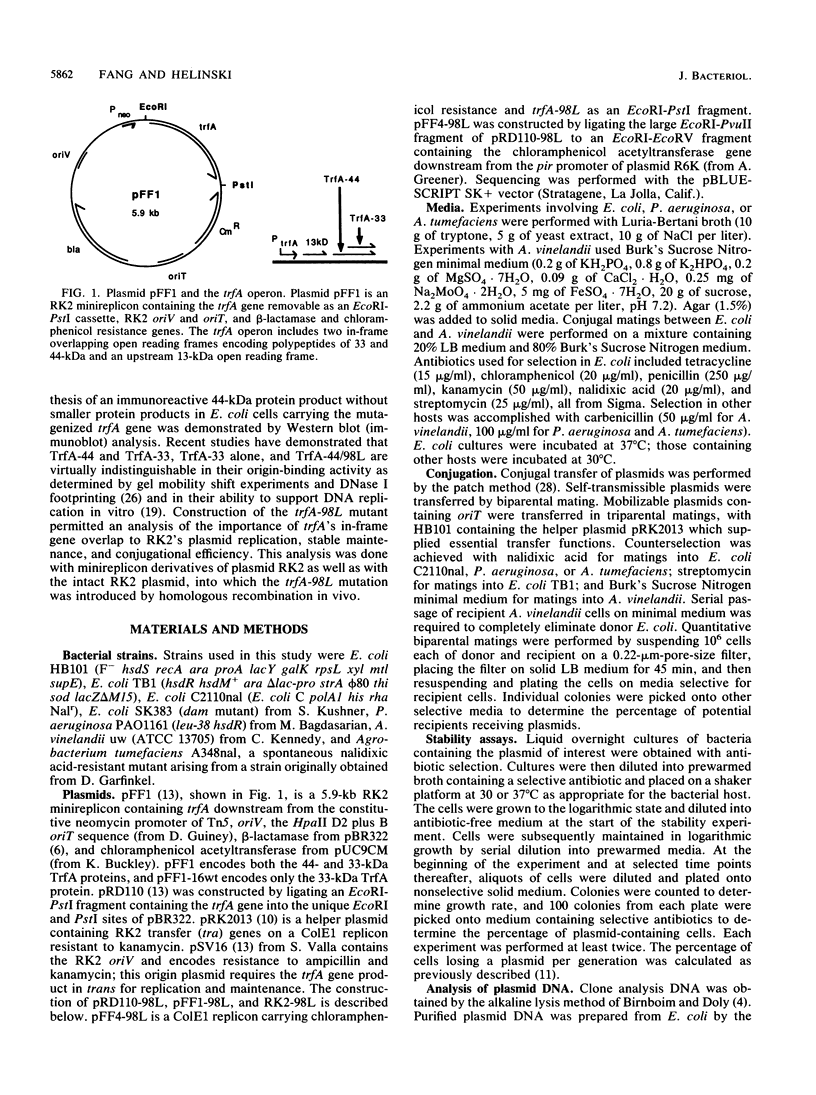




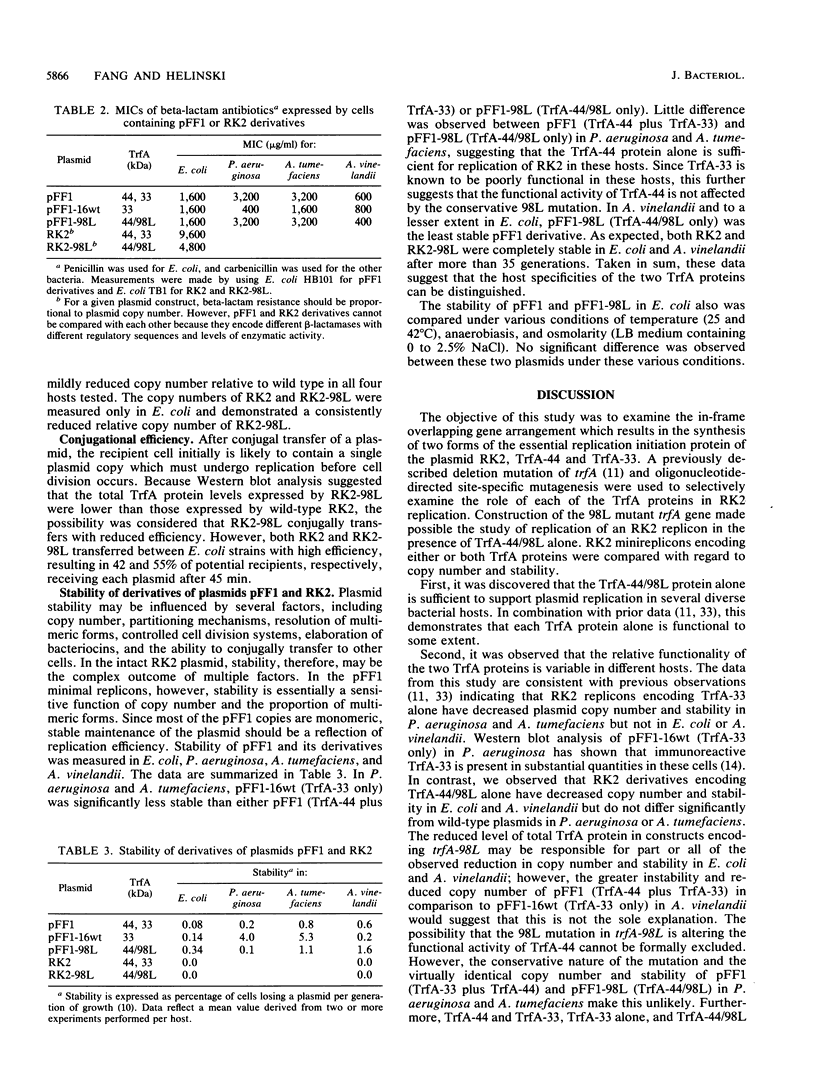


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Barth P. T., Ellis K., Bechhofer D. H., Figurski D. H. Involvement of kil and kor genes in the phenotype of a host-range mutant of RP4. Mol Gen Genet. 1984;197(2):236–243. doi: 10.1007/BF00330969. [DOI] [PubMed] [Google Scholar]
- Bernstein J. A., Richardson C. C. A 7-kDa region of the bacteriophage T7 gene 4 protein is required for primase but not for helicase activity. Proc Natl Acad Sci U S A. 1988 Jan;85(2):396–400. doi: 10.1073/pnas.85.2.396. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Betlach M. C., Boyer H. W. Construction and characterization of new cloning vehicles. I. Ampicillin-resistant derivatives of the plasmid pMB9. Gene. 1977;2(2):75–93. doi: 10.1016/0378-1119(77)90074-9. [DOI] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Greene P. J., Betlach M. C., Heyneker H. L., Boyer H. W., Crosa J. H., Falkow S. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene. 1977;2(2):95–113. [PubMed] [Google Scholar]
- Breton A. M., Jaoua S., Guespin-Michel J. Transfer of plasmid RP4 to Myxococcus xanthus and evidence for its integration into the chromosome. J Bacteriol. 1985 Feb;161(2):523–528. doi: 10.1128/jb.161.2.523-528.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen N. Y., Paulus H. Mechanism of expression of the overlapping genes of Bacillus subtilis aspartokinase II. J Biol Chem. 1988 Jul 5;263(19):9526–9532. [PubMed] [Google Scholar]
- Cohen S. N., Chang A. C., Hsu L. Nonchromosomal antibiotic resistance in bacteria: genetic transformation of Escherichia coli by R-factor DNA. Proc Natl Acad Sci U S A. 1972 Aug;69(8):2110–2114. doi: 10.1073/pnas.69.8.2110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ditta G., Stanfield S., Corbin D., Helinski D. R. Broad host range DNA cloning system for gram-negative bacteria: construction of a gene bank of Rhizobium meliloti. Proc Natl Acad Sci U S A. 1980 Dec;77(12):7347–7351. doi: 10.1073/pnas.77.12.7347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Durland R. H., Helinski D. R. Replication of the broad-host-range plasmid RK2: direct measurement of intracellular concentrations of the essential TrfA replication proteins and their effect on plasmid copy number. J Bacteriol. 1990 Jul;172(7):3849–3858. doi: 10.1128/jb.172.7.3849-3858.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Durland R. H., Helinski D. R. The sequence encoding the 43-kilodalton trfA protein is required for efficient replication or maintenance of minimal RK2 replicons in Pseudomonas aeruginosa. Plasmid. 1987 Sep;18(2):164–169. doi: 10.1016/0147-619x(87)90044-8. [DOI] [PubMed] [Google Scholar]
- Feinberg A. P., Vogelstein B. A technique for radiolabeling DNA restriction endonuclease fragments to high specific activity. Anal Biochem. 1983 Jul 1;132(1):6–13. doi: 10.1016/0003-2697(83)90418-9. [DOI] [PubMed] [Google Scholar]
- Guiney D. G., Hasegawa P., Davis C. E. Plasmid transfer from Escherichia coli to Bacteroides fragilis: differential expression of antibiotic resistance phenotypes. Proc Natl Acad Sci U S A. 1984 Nov;81(22):7203–7206. doi: 10.1073/pnas.81.22.7203. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heinemann J. A., Sprague G. F., Jr Bacterial conjugative plasmids mobilize DNA transfer between bacteria and yeast. Nature. 1989 Jul 20;340(6230):205–209. doi: 10.1038/340205a0. [DOI] [PubMed] [Google Scholar]
- Isberg R. R., Lazaar A. L., Syvanen M. Regulation of Tn5 by the right-repeat proteins: control at the level of the transposition reaction? Cell. 1982 Oct;30(3):883–892. doi: 10.1016/0092-8674(82)90293-8. [DOI] [PubMed] [Google Scholar]
- Kittell B. L., Helinski D. R. Iteron inhibition of plasmid RK2 replication in vitro: evidence for intermolecular coupling of replication origins as a mechanism for RK2 replication control. Proc Natl Acad Sci U S A. 1991 Feb 15;88(4):1389–1393. doi: 10.1073/pnas.88.4.1389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kornacki J. A., West A. H., Firshein W. Proteins encoded by the trans-acting replication and maintenance regions of broad host range plasmid RK2. Plasmid. 1984 Jan;11(1):48–57. doi: 10.1016/0147-619x(84)90006-4. [DOI] [PubMed] [Google Scholar]
- Kozak M. Bifunctional messenger RNAs in eukaryotes. Cell. 1986 Nov 21;47(4):481–483. doi: 10.1016/0092-8674(86)90609-4. [DOI] [PubMed] [Google Scholar]
- Lanka E., Lurz R., Fürste J. P. Molecular cloning and mapping of SphI restriction fragments of plasmid RP4. Plasmid. 1983 Nov;10(3):303–307. doi: 10.1016/0147-619x(83)90047-1. [DOI] [PubMed] [Google Scholar]
- Linney E., Hayashi M. Two proteins of gene A of psiX174. Nat New Biol. 1973 Sep 5;245(140):6–8. doi: 10.1038/newbio245006a0. [DOI] [PubMed] [Google Scholar]
- Meyer R., Figurski D., Helinski D. R. Physical and genetic studies with restriction endonucleases on the broad host-range plasmid RK2. Mol Gen Genet. 1977 Apr 29;152(3):129–135. doi: 10.1007/BF00268809. [DOI] [PubMed] [Google Scholar]
- Perri S., Helinski D. R., Toukdarian A. Interactions of plasmid-encoded replication initiation proteins with the origin of DNA replication in the broad host range plasmid RK2. J Biol Chem. 1991 Jul 5;266(19):12536–12543. [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmidhauser T. J., Helinski D. R. Regions of broad-host-range plasmid RK2 involved in replication and stable maintenance in nine species of gram-negative bacteria. J Bacteriol. 1985 Oct;164(1):446–455. doi: 10.1128/jb.164.1.446-455.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shaw J. E., Murialdo H. Morphogenetic genes C and Nu3 overlap in bacteriophage lambda. Nature. 1980 Jan 3;283(5742):30–35. doi: 10.1038/283030a0. [DOI] [PubMed] [Google Scholar]
- Shen B. F., Tai P. C., Pritchard A. E., Vasil M. L. Nucleotide sequences and expression in Escherichia coli of the in-phase overlapping Pseudomonas aeruginosa plcR genes. J Bacteriol. 1987 Oct;169(10):4602–4607. doi: 10.1128/jb.169.10.4602-4607.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shingler V., Thomas C. M. Analysis of nonpolar insertion mutations in the trfA gene of IncP plasmid RK2 which affect its broad-host-range property. Biochim Biophys Acta. 1989 Apr 12;1007(3):301–308. doi: 10.1016/0167-4781(89)90152-8. [DOI] [PubMed] [Google Scholar]
- Shingler V., Thomas C. M. Analysis of the trfA region of broad host-range plasmid RK2 by transposon mutagenesis and identification of polypeptide products. J Mol Biol. 1984 May 25;175(3):229–249. doi: 10.1016/0022-2836(84)90346-2. [DOI] [PubMed] [Google Scholar]
- Smith C. A., Thomas C. M. Nucleotide sequence of the trfA gene of broad host-range plasmid RK2. J Mol Biol. 1984 May 25;175(3):251–262. doi: 10.1016/0022-2836(84)90347-4. [DOI] [PubMed] [Google Scholar]
- Smith R. A., Parkinson J. S. Overlapping genes at the cheA locus of Escherichia coli. Proc Natl Acad Sci U S A. 1980 Sep;77(9):5370–5374. doi: 10.1073/pnas.77.9.5370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tabor S., Richardson C. C. DNA sequence analysis with a modified bacteriophage T7 DNA polymerase. Proc Natl Acad Sci U S A. 1987 Jul;84(14):4767–4771. doi: 10.1073/pnas.84.14.4767. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thomas C. M. Recent studies on the control of plasmid replication. Biochim Biophys Acta. 1988 Mar 31;949(3):253–263. doi: 10.1016/0167-4781(88)90150-9. [DOI] [PubMed] [Google Scholar]
- Thomas C. M., Smith C. A. Incompatibility group P plasmids: genetics, evolution, and use in genetic manipulation. Annu Rev Microbiol. 1987;41:77–101. doi: 10.1146/annurev.mi.41.100187.000453. [DOI] [PubMed] [Google Scholar]
- Thomas C. M., Smith C. A. The trfB region of broad host range plasmid RK2: the nucleotide sequence reveals incC and key regulatory gene trfB/korA/korD as overlapping genes. Nucleic Acids Res. 1986 Jun 11;14(11):4453–4469. doi: 10.1093/nar/14.11.4453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uhlin B. E., Nordström K. R plasmid gene dosage effects in Escherichia coli K-12: copy mutants of the R plasmic R1drd-19. Plasmid. 1977 Nov;1(1):1–7. doi: 10.1016/0147-619x(77)90003-8. [DOI] [PubMed] [Google Scholar]
- Yen T. S., Webster R. E. Bacteriophage f1 gene II and X proteins. Isolation and characterization of the products of two overlapping genes. J Biol Chem. 1981 Nov 10;256(21):11259–11265. [PubMed] [Google Scholar]
- Zoller M. J., Smith M. Oligonucleotide-directed mutagenesis of DNA fragments cloned into M13 vectors. Methods Enzymol. 1983;100:468–500. doi: 10.1016/0076-6879(83)00074-9. [DOI] [PubMed] [Google Scholar]



