Abstract
Small bacteriophage D3112 transposable elements deleted for most of the phage-lytic functions while retaining the sites required for transposition and packaging were constructed to facilitate genetic studies in Pseudomonas aeruginosa. These mini-D derivatives were constructed with the terminal 1.85 kilobases (kb) of the phage left end and 1.4 kb of the phage right end and either the Tn5 kanamycin resistance or the pSC101 (pBR322) tetracycline resistance determinant. Thermally induced lysates of strains lysogenic for both a mini-D element and D3112 cts (temperature-sensitive repressor) transduced P. aeruginosa PAO recipients to drug resistance at frequencies of between 10(-4) and 10(-5)/PFU of the helper phage. As for the parent plaque-forming D3112 phage, the mini-D171 element could insert itself into many different sites in the chromosome but the frequency of insertion into particular genes varied widely. Among 1,000 insertions, none resulted in auxotrophy but 10 resulted in pigment production. Insertions were also selected in a cloning plasmid with a transduction scheme. At least eight different insertion sites were found to have been used among 10 individual insertions. Transductants harboring these mini-D elements were immune to infection by D3112, since they contained the D3112 repressor gene in the left 1.85-kb terminal fragment. Chromosomal genes were transduced in a generalized fashion 100 to 1,000 times more frequently by the mini-D-D3112 cts lysates than by the D3112 cts phage alone. Mini-D171-D3112 cts lysates also yielded some transductants that retained the drug resistance marker of the mini-D element and which were unstable for the chromosomal transduced marker. This is consistent with the miniduction properties of Mu whereby transduced genes are flanked by two mini-D elements in the same orientation.
Full text
PDF






Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Akhverdian V. Z., Khrenova E. A., Bogush V. G., Gerasimova T. V., Kirsanov N. B. Shirokaia rasprostranennost' transpozonnykh fagov v prirodnykh populiatsiiakh Pseudomonas aeruginosa. Genetika. 1984 Oct;20(10):1612–1619. [PubMed] [Google Scholar]
- BRAMMAR W. J., CLARKE P. H. INDUCTION AND REPRESSION OF PSEUDOMONAS AERUGINOSA AMIDASE. J Gen Microbiol. 1964 Dec;37:307–319. doi: 10.1099/00221287-37-3-307. [DOI] [PubMed] [Google Scholar]
- Bagdasarian M. M., Amann E., Lurz R., Rückert B., Bagdasarian M. Activity of the hybrid trp-lac (tac) promoter of Escherichia coli in Pseudomonas putida. Construction of broad-host-range, controlled-expression vectors. Gene. 1983 Dec;26(2-3):273–282. doi: 10.1016/0378-1119(83)90197-x. [DOI] [PubMed] [Google Scholar]
- Barth P. T., Grinter N. J. Comparison of the deoxyribonucleic acid molecular weights and homologies of plasmids conferring linked resistance to streptomycin and sulfonamides. J Bacteriol. 1974 Nov;120(2):618–630. doi: 10.1128/jb.120.2.618-630.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Beck E., Ludwig G., Auerswald E. A., Reiss B., Schaller H. Nucleotide sequence and exact localization of the neomycin phosphotransferase gene from transposon Tn5. Gene. 1982 Oct;19(3):327–336. doi: 10.1016/0378-1119(82)90023-3. [DOI] [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Castilho B. A., Olfson P., Casadaban M. J. Plasmid insertion mutagenesis and lac gene fusion with mini-mu bacteriophage transposons. J Bacteriol. 1984 May;158(2):488–495. doi: 10.1128/jb.158.2.488-495.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Craigie R., Mizuuchi M., Mizuuchi K. Site-specific recognition of the bacteriophage Mu ends by the Mu A protein. Cell. 1984 Dec;39(2 Pt 1):387–394. doi: 10.1016/0092-8674(84)90017-5. [DOI] [PubMed] [Google Scholar]
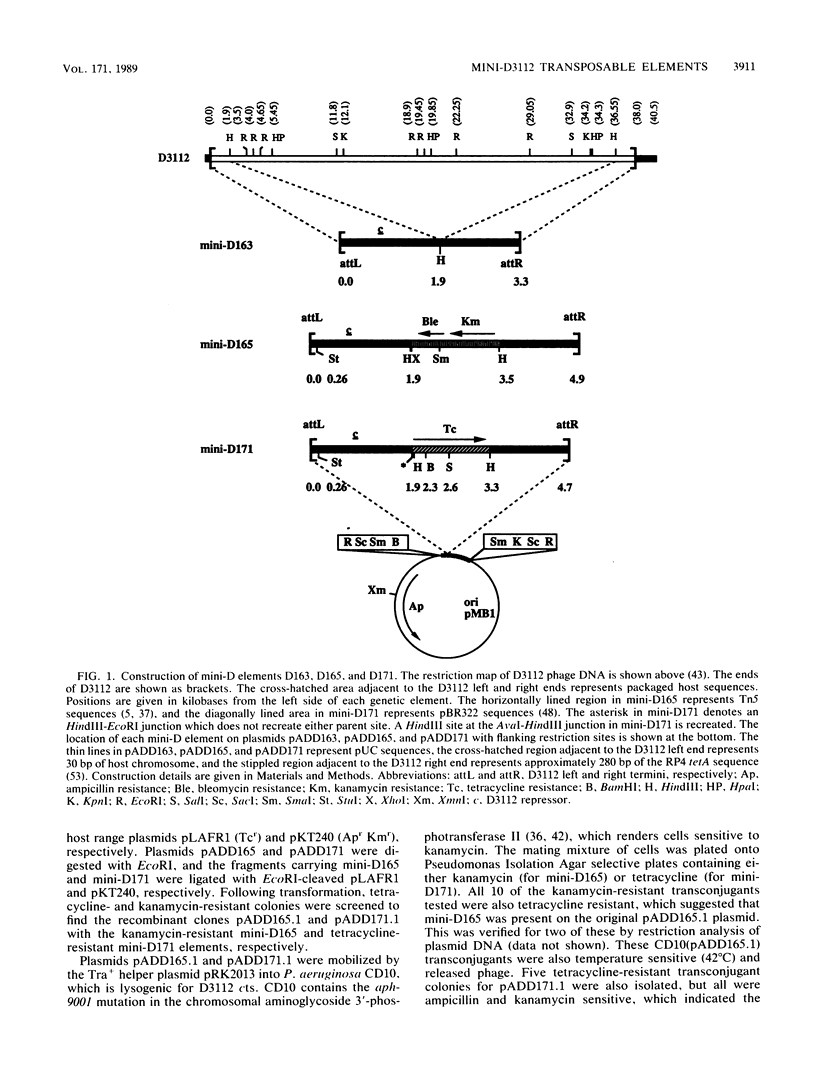
- Darzins A., Casadaban M. J. In vivo cloning of Pseudomonas aeruginosa genes with mini-D3112 transposable bacteriophage. J Bacteriol. 1989 Jul;171(7):3917–3925. doi: 10.1128/jb.171.7.3917-3925.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Darzins A., Wang S. K., Vanags R. I., Chakrabarty A. M. Clustering of mutations affecting alginic acid biosynthesis in mucoid Pseudomonas aeruginosa. J Bacteriol. 1985 Nov;164(2):516–524. doi: 10.1128/jb.164.2.516-524.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ditta G., Stanfield S., Corbin D., Helinski D. R. Broad host range DNA cloning system for gram-negative bacteria: construction of a gene bank of Rhizobium meliloti. Proc Natl Acad Sci U S A. 1980 Dec;77(12):7347–7351. doi: 10.1073/pnas.77.12.7347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Faelen M., Huisman O., Toussaint A. Involvement of phage Mu-1 early functions in Mu-mediated chromosomal rearrangements. Nature. 1978 Feb 9;271(5645):580–582. doi: 10.1038/271580a0. [DOI] [PubMed] [Google Scholar]
- Faelen M., Resibois A., Toussaint A. Mini-mu: an insertion element derived from temperate phage mu-1. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 2):1169–1177. doi: 10.1101/sqb.1979.043.01.132. [DOI] [PubMed] [Google Scholar]
- Faelen M., Toussaint A., Resibois A. Mini-muduction: a new mode of gene transfer mediated by mini-mu. Mol Gen Genet. 1979 Oct 3;176(2):191–197. doi: 10.1007/BF00273213. [DOI] [PubMed] [Google Scholar]
- Figurski D. H., Helinski D. R. Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc Natl Acad Sci U S A. 1979 Apr;76(4):1648–1652. doi: 10.1073/pnas.76.4.1648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Friedman A. M., Long S. R., Brown S. E., Buikema W. J., Ausubel F. M. Construction of a broad host range cosmid cloning vector and its use in the genetic analysis of Rhizobium mutants. Gene. 1982 Jun;18(3):289–296. doi: 10.1016/0378-1119(82)90167-6. [DOI] [PubMed] [Google Scholar]
- Grinter N. J. A broad-host-range cloning vector transposable to various replicons. Gene. 1983 Jan-Feb;21(1-2):133–143. doi: 10.1016/0378-1119(83)90155-5. [DOI] [PubMed] [Google Scholar]
- Groenen M. A., Timmers E., van de Putte P. DNA sequences at the ends of the genome of bacteriophage Mu essential for transposition. Proc Natl Acad Sci U S A. 1985 Apr;82(7):2087–2091. doi: 10.1073/pnas.82.7.2087. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Guiney D. G., Yakobson E. Location and nucleotide sequence of the transfer origin of the broad host range plasmid RK2. Proc Natl Acad Sci U S A. 1983 Jun;80(12):3595–3598. doi: 10.1073/pnas.80.12.3595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- HOLLOWAY B. W. Genetic recombination in Pseudomonas aeruginosa. J Gen Microbiol. 1955 Dec;13(3):572–581. doi: 10.1099/00221287-13-3-572. [DOI] [PubMed] [Google Scholar]
- Haas D. Genetic aspects of biodegradation by pseudomonads. Experientia. 1983 Nov 15;39(11):1199–1213. doi: 10.1007/BF01990357. [DOI] [PubMed] [Google Scholar]
- Holmes D. S., Quigley M. A rapid boiling method for the preparation of bacterial plasmids. Anal Biochem. 1981 Jun;114(1):193–197. doi: 10.1016/0003-2697(81)90473-5. [DOI] [PubMed] [Google Scholar]
- Ianenko A. S., Bogush V. G., Kirsanov N. B., Liapin M. N., Krylov V. N. Ispol'zovanie deletsionnykh mutantov plazmidy RP::4D3112 dlia geneticheskogo analiza bakteriofaga D3112 Pseudomonas aeruginosa. Genetika. 1983 Nov;19(11):1760–1768. [PubMed] [Google Scholar]
- Ianenko A. S., Kirsanov N. B., Gerasimova T. V., Bekkarevich A. O., Krylov V. N. Mini-genomy faga-transpozona D3112 Pseudomonas aeruginosa i ikh svoistva. Genetika. 1988 May;24(5):956–959. [PubMed] [Google Scholar]
- Kokjohn T. A., Miller R. V. Characterization of the Pseudomonas aeruginosa recA analog and its protein product: rec-102 is a mutant allele of the P. aeruginosa PAO recA gene. J Bacteriol. 1987 Apr;169(4):1499–1508. doi: 10.1128/jb.169.4.1499-1508.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Krishnapillai V. A novel transducing phage. Its role in recognition of a possible new host-controlled modification system in Pseudomonas aeruginosa. Mol Gen Genet. 1972;114(2):134–143. doi: 10.1007/BF00332784. [DOI] [PubMed] [Google Scholar]
- Krylov V. N., Bogush V. G., Ianenko A. S., Kirsanov N. B. Bakteriofagi Pseudomonas aeruginosa, struktura DNA kotorykh skhodna so strukturoi DNK faga Mu1. Soobshchenie II. Dokazatel'stvo rodstva bakteriofagov D3112, B3 i B39: analiz rasshchepleniia DNK éndonukleazami restriktsii, vydelenie rekombinanta fagov D3112 i B3. Genetika. 1980;16(6):975–984. [PubMed] [Google Scholar]
- Krylov V. N., Plotnikova T. G., Kulakov L. A., Fedorova T. V., Eremenko E. N. Integratsiia genoma Mu-podobnogo bakteriofaga D3112 Pseudomonas aeruginosa v plazmidu RP4 i vvedeneie v sostave gibridnoi plazmidy v bakterii Pseudomonas putida i Escherichia coli C600. Genetika. 1982;18(1):5–12. [PubMed] [Google Scholar]
- Kupersztoch Y. M., Helinski D. R. A catenated DNA molecule as an intermediate in the replication of the resistance transfer factor R6K in Escherichia coli. Biochem Biophys Res Commun. 1973 Oct 15;54(4):1451–1459. doi: 10.1016/0006-291x(73)91149-2. [DOI] [PubMed] [Google Scholar]
- Mandel M., Higa A. Calcium-dependent bacteriophage DNA infection. J Mol Biol. 1970 Oct 14;53(1):159–162. doi: 10.1016/0022-2836(70)90051-3. [DOI] [PubMed] [Google Scholar]
- Matsuhashi Y., Yagisawa M., Kondo S., Takeuchi T., Umezawa H. Aminoglycoside 3'-phosphotransferases I and II in Pseudomonas aeruginosa. J Antibiot (Tokyo) 1975 Jun;28(6):442–447. doi: 10.7164/antibiotics.28.442. [DOI] [PubMed] [Google Scholar]
- Mazodier P., Cossart P., Giraud E., Gasser F. Completion of the nucleotide sequence of the central region of Tn5 confirms the presence of three resistance genes. Nucleic Acids Res. 1985 Jan 11;13(1):195–205. doi: 10.1093/nar/13.1.195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Morrison D. A. Transformation and preservation of competent bacterial cells by freezing. Methods Enzymol. 1979;68:326–331. doi: 10.1016/0076-6879(79)68023-0. [DOI] [PubMed] [Google Scholar]
- Murooka Y., Takizawa N., Harada T. Introduction of bacteriophage Mu into bacteria of various genera and intergeneric gene transfer by RP4::Mu. J Bacteriol. 1981 Jan;145(1):358–368. doi: 10.1128/jb.145.1.358-368.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- O'Hoy K., Krishnapillai V. Recalibration of the Pseudomonas aeruginosa strain PAO chromosome map in time units using high-frequency-of-recombination donors. Genetics. 1987 Apr;115(4):611–618. doi: 10.1093/genetics/115.4.611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okii M., Iyobe S., Mitsuhashi S. Mapping of the gene specifying aminoglycoside 3'-phosphotransferase II on the Pseudomonas aeruginosa chromosome. J Bacteriol. 1983 Aug;155(2):643–649. doi: 10.1128/jb.155.2.643-649.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rehmat S., Shapiro J. A. Insertion and replication of the Pseudomonas aeruginosa mutator phage D3112. Mol Gen Genet. 1983;192(3):416–423. doi: 10.1007/BF00392184. [DOI] [PubMed] [Google Scholar]
- Royle P. L., Matsumoto H., Holloway B. W. Genetic circularity of the Pseudomonas aeruginosa PAO chromosome. J Bacteriol. 1981 Jan;145(1):145–155. doi: 10.1128/jb.145.1.145-155.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith C. A., Thomas C. M. Nucleotide sequence of the trfA gene of broad host-range plasmid RK2. J Mol Biol. 1984 May 25;175(3):251–262. doi: 10.1016/0022-2836(84)90347-4. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Stalker D. M., Thomas C. M., Helinski D. R. Nucleotide sequence of the region of the origin of replication of the broad host range plasmid RK2. Mol Gen Genet. 1981;181(1):8–12. doi: 10.1007/BF00338997. [DOI] [PubMed] [Google Scholar]
- Sutcliffe J. G. Complete nucleotide sequence of the Escherichia coli plasmid pBR322. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 1):77–90. doi: 10.1101/sqb.1979.043.01.013. [DOI] [PubMed] [Google Scholar]
- TAYLOR A. L. BACTERIOPHAGE-INDUCED MUTATION IN ESCHERICHIA COLI. Proc Natl Acad Sci U S A. 1963 Dec;50:1043–1051. doi: 10.1073/pnas.50.6.1043. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vieira J., Messing J. The pUC plasmids, an M13mp7-derived system for insertion mutagenesis and sequencing with synthetic universal primers. Gene. 1982 Oct;19(3):259–268. doi: 10.1016/0378-1119(82)90015-4. [DOI] [PubMed] [Google Scholar]
- Vogelstein B., Gillespie D. Preparative and analytical purification of DNA from agarose. Proc Natl Acad Sci U S A. 1979 Feb;76(2):615–619. doi: 10.1073/pnas.76.2.615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Waters S. H., Rogowsky P., Grinsted J., Altenbuchner J., Schmitt R. The tetracycline resistance determinants of RP1 and Tn1721: nucleotide sequence analysis. Nucleic Acids Res. 1983 Sep 10;11(17):6089–6105. doi: 10.1093/nar/11.17.6089. [DOI] [PMC free article] [PubMed] [Google Scholar]