Abstract
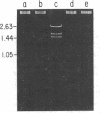
The methylations of adenine in the sequence —GATC— and of the second cytosine in the sequence — [Formula: see text] — were studied in Salmonella typhimurium and in Salmonella typhi. The study was carried out by using endonucleases which restrict the plasmid pBR322 by cleavage at the sequences —GATC— (DpnI and MboI) and — [Formula: see text] — (EcoRII). The restriction patterns obtained for this plasmid isolated from transformed S. typhimurium and S. typhi were compared with those of pBR322 isolated from Escherichia coli K-12. In E. coli K-12, adenines at the sequence —GATC— and the second cytosines at — [Formula: see text] — are met hylated by enzymes coded for by the genes dam and dem, respectively. From comparison of the restriction patterns obtained, it is concluded that S. typhimurium and S. typhi contain genes responsible for deoxyribonucleic acid methylation equivalent to E. coli K-12 genes dam and dcm.
Full text
PDF




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alfaro G., Martuscelli J., Mendoza-Hernández P. Antibiotic resistance and phage-types of Salmonella typhi strains isolated in Mexico City. Rev Latinoam Microbiol. 1978 Jan-Mar;20(1):5–11. [PubMed] [Google Scholar]
- Bigger C. H., Murray K., Murray N. E. Recognition sequence of a restriction enzyme. Nat New Biol. 1973 Jul 4;244(131):7–10. doi: 10.1038/newbio244007a0. [DOI] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Betlach M. C., Boyer H. W. Construction and characterization of new cloning vehicles. I. Ampicillin-resistant derivatives of the plasmid pMB9. Gene. 1977;2(2):75–93. doi: 10.1016/0378-1119(77)90074-9. [DOI] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Greene P. J., Betlach M. C., Heyneker H. L., Boyer H. W., Crosa J. H., Falkow S. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene. 1977;2(2):95–113. [PubMed] [Google Scholar]
- Brockes J. P., Brown P. R., Murray K. Nucleotide sequences at the sites of action of the deoxyribonucleic acid modification enzyme of bacteriophage P1. J Mol Biol. 1974 Sep 15;88(2):437–443. doi: 10.1016/0022-2836(74)90493-8. [DOI] [PubMed] [Google Scholar]
- Clewell D. B. Nature of Col E 1 plasmid replication in Escherichia coli in the presence of the chloramphenicol. J Bacteriol. 1972 May;110(2):667–676. doi: 10.1128/jb.110.2.667-676.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cohen S. N., Chang A. C., Hsu L. Nonchromosomal antibiotic resistance in bacteria: genetic transformation of Escherichia coli by R-factor DNA. Proc Natl Acad Sci U S A. 1972 Aug;69(8):2110–2114. doi: 10.1073/pnas.69.8.2110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- DUNN D. B., SMITH J. D. The occurrence of 6-methylaminopurine in deoxyribonucleic acids. Biochem J. 1958 Apr;68(4):627–636. doi: 10.1042/bj0680627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujimoto D., Srinivasan P. R., Borek E. On the nature of the deoxyribonucleic acid methylases. Biological evidence for the multiple nature of the enzymes. Biochemistry. 1965 Dec;4(12):2849–2855. doi: 10.1021/bi00888a041. [DOI] [PubMed] [Google Scholar]
- GOLD M., HURWITZ J. THE ENZYMATIC METHYLATION OF RIBONUCLEIC ACID AND DEOXYRIBONUCLEIC ACID. V. PURIFICATION AND PROPERTIES OF THE DEOXYRIBONUCLEIC ACID-METHYLATING ACTIVITY OF ESCHERICHIA COLI. J Biol Chem. 1964 Nov;239:3858–3865. [PubMed] [Google Scholar]
- Gelinas R. E., Myers P. A., Roberts R. J. Two sequence-specific endonucleases from Moraxella bovis. J Mol Biol. 1977 Jul;114(1):169–179. doi: 10.1016/0022-2836(77)90290-x. [DOI] [PubMed] [Google Scholar]
- Guerry P., LeBlanc D. J., Falkow S. General method for the isolation of plasmid deoxyribonucleic acid. J Bacteriol. 1973 Nov;116(2):1064–1066. doi: 10.1128/jb.116.2.1064-1066.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hattman S., Brooks J. E., Masurekar M. Sequence specificity of the P1 modification methylase (M.Eco P1) and the DNA methylase (M.Eco dam) controlled by the Escherichia coli dam gene. J Mol Biol. 1978 Dec 15;126(3):367–380. doi: 10.1016/0022-2836(78)90046-3. [DOI] [PubMed] [Google Scholar]
- Hattman S., Gold E., Plotnik A. Methylation of cytosine residues in DNA controlled by a drug resistance factor (host-induced modification-R factors-N 6 -methyladenine-5-methylcytosine). Proc Natl Acad Sci U S A. 1972 Jan;69(1):187–190. doi: 10.1073/pnas.69.1.187. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hattman S., Schlagman S., Goldstein L., Frohlich M. Salmonella typhimurium SA host specificity system is based on deoxyribonucleic acid-adenine methylation. J Bacteriol. 1976 Jul;127(1):211–217. doi: 10.1128/jb.127.1.211-217.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hattman S. Variation of 6-methylaminopurine content in bacteriophage P22 deoxyribonucleic acid as a function of host specificity. J Virol. 1971 May;7(5):690–691. doi: 10.1128/jvi.7.5.690-691.1971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hattman S., van Ormondt H., de Waard A. Sequence specificity of the wild-type dam+) and mutant (damh) forms of bacteriophage T2 DNA adenine methylase. J Mol Biol. 1978 Mar 5;119(3):361–376. doi: 10.1016/0022-2836(78)90219-x. [DOI] [PubMed] [Google Scholar]
- Krüger D. H., Schroeder C., Hansen S., Rosenthal H. A. Active protection by bacteriophages T3 and T7 against E. coli B- and K-specific restriction of their DNA. Mol Gen Genet. 1977 May 20;153(1):99–106. doi: 10.1007/BF01036001. [DOI] [PubMed] [Google Scholar]
- LENNOX E. S. Transduction of linked genetic characters of the host by bacteriophage P1. Virology. 1955 Jul;1(2):190–206. doi: 10.1016/0042-6822(55)90016-7. [DOI] [PubMed] [Google Scholar]
- Lacks S., Greenberg B. Complementary specificity of restriction endonucleases of Diplococcus pneumoniae with respect to DNA methylation. J Mol Biol. 1977 Jul;114(1):153–168. doi: 10.1016/0022-2836(77)90289-3. [DOI] [PubMed] [Google Scholar]
- Lederberg E. M., Cohen S. N. Transformation of Salmonella typhimurium by plasmid deoxyribonucleic acid. J Bacteriol. 1974 Sep;119(3):1072–1074. doi: 10.1128/jb.119.3.1072-1074.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mamelak L., Boyer H. W. Genetic control of the secondary modification of deoxyribonucleic acid in Escherichia coli. J Bacteriol. 1970 Oct;104(1):57–62. doi: 10.1128/jb.104.1.57-62.1970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marinus M. G., Morris N. R. Biological function for 6-methyladenine residues in the DNA of Escherichia coli K12. J Mol Biol. 1974 May 15;85(2):309–322. doi: 10.1016/0022-2836(74)90366-0. [DOI] [PubMed] [Google Scholar]
- Marinus M. G., Morris N. R. Isolation of deoxyribonucleic acid methylase mutants of Escherichia coli K-12. J Bacteriol. 1973 Jun;114(3):1143–1150. doi: 10.1128/jb.114.3.1143-1150.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sanger F., Coulson A. R., Friedmann T., Air G. M., Barrell B. G., Brown N. L., Fiddes J. C., Hutchison C. A., 3rd, Slocombe P. M., Smith M. The nucleotide sequence of bacteriophage phiX174. J Mol Biol. 1978 Oct 25;125(2):225–246. doi: 10.1016/0022-2836(78)90346-7. [DOI] [PubMed] [Google Scholar]
- Schlagman S., Hattman S., May M. S., Berger L. In vivo methylation by Escherichia coli K-12 mec+ deoxyribonucleic acid-cytosine methylase protects against in vitro cleavage by the RII restriction endonuclease (R. Eco RII). J Bacteriol. 1976 May;126(2):990–996. doi: 10.1128/jb.126.2.990-996.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Studier F. W. Gene 0.3 of bacteriophage T7 acts to overcome the DNA restriction system of the host. J Mol Biol. 1975 May 15;94(2):283–295. doi: 10.1016/0022-2836(75)90083-2. [DOI] [PubMed] [Google Scholar]
- Studier F. W., Movva N. R. SAMase gene of bacteriophage T3 is responsible for overcoming host restriction. J Virol. 1976 Jul;19(1):136–145. doi: 10.1128/jvi.19.1.136-145.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vanyushin B. F., Belozersky A. N., Kokurina N. A., Kadirova D. X. 5-methylcytosine and 6-methylamino-purine in bacterial DNA. Nature. 1968 Jun 15;218(5146):1066–1067. doi: 10.1038/2181066a0. [DOI] [PubMed] [Google Scholar]
- Vovis G. F., Lacks S. Complementary action of restriction enzymes endo R-DpnI and Endo R-DpnII on bacteriophage f1 DNA. J Mol Biol. 1977 Sep 25;115(3):525–538. doi: 10.1016/0022-2836(77)90169-3. [DOI] [PubMed] [Google Scholar]
- Watanabe T., Takano T., Arai T., Nishida H., Sato S. Episome-mediated Transfer of Drug Resistance in Enterobacteriaceae X. Restriction and Modification of Phages by fi R Factors. J Bacteriol. 1966 Aug;92(2):477–486. doi: 10.1128/jb.92.2.477-486.1966. [DOI] [PMC free article] [PubMed] [Google Scholar]