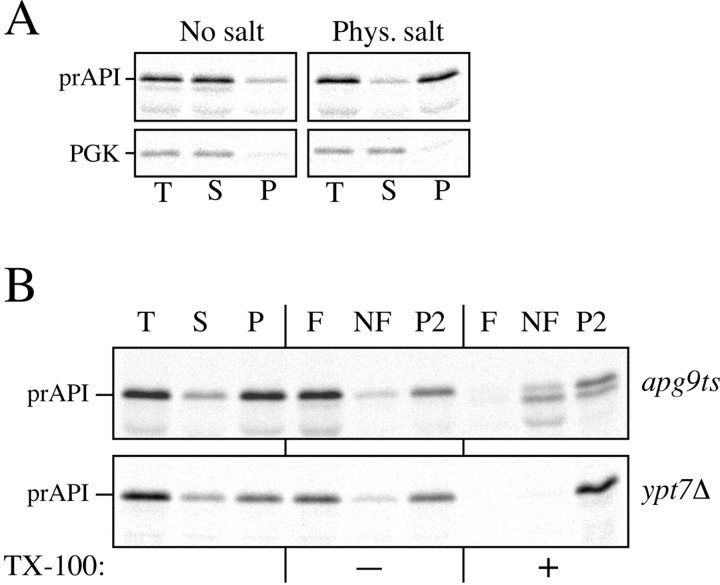
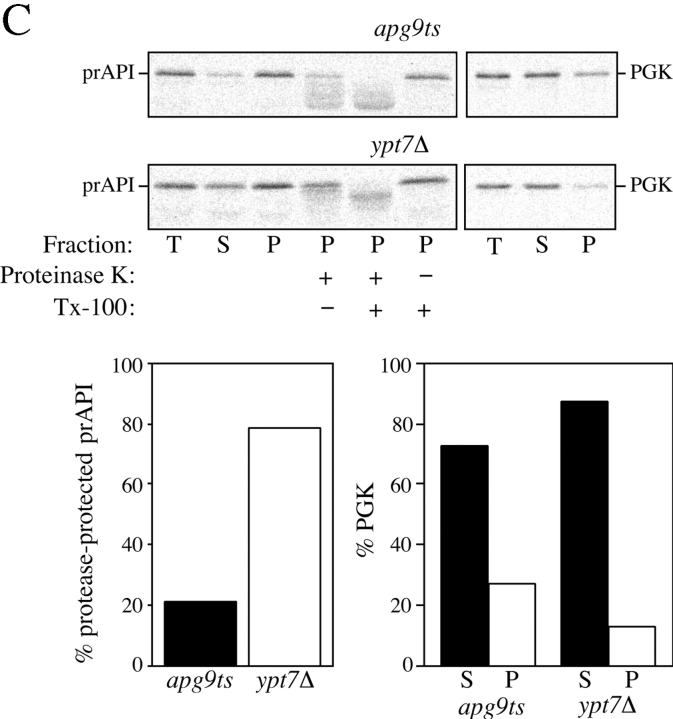
Figure 10.
Analysis of the apg9ts strain indicates Apg9p is directly required for the vesicle formation step. A, The nature of prAPI binding and pelleting in the apg9ts strain. Spheroplasts of apg9ts were labeled for 10 min and chased for 30 min at 38°C. The spheroplasts were osmotically lysed in a buffer containing no salt (20 mM Pipes, pH 6.8) or a physiological concentration of salt (100 mM KOAc, 50 mM KCl, 5 mM MgCl2, 20 mM Pipes, pH 6.8) and pelleted at 5,000 g. All samples were immunoprecipitated with antiserum to API and the cytosolic marker PGK. apg9ts retains the salt dependent binding and pelleting of prAPI. B, prAPI in the apg9ts strain accumulates in both a membrane-associated state and a large pelletable complex. Spheroplasts of apg9ts and ypt7Δ were labeled for 10 min and chased for 30 min at 38°C. The labeled spheroplasts were then osmotically lysed and separated into supernatant (S) and pellet (P) fractions by centrifugation at 5,000 g for 5 min. An aliquot was removed for the total lysate control (T). The pellet fraction (P) was resuspended in 15% Ficoll-400 in gradient buffer (20 mM Pipes, 5 mM MgCl2, complete EDTA-free protease inhibitor cocktail) in the presence or absence of Triton X-100 and overlaid with 13 and 2% Ficoll-400 in gradient buffer. The step gradients were centrifuged at 13,000 g for 10 min. Membrane-containing float (F), nonfloat (NF), and pellet (P2) fractions were immunoprecipitated with antiserum to API as described in Materials and Methods. The position of prAPI is indicated. C, prAPI in the apg9ts strain is protease accessible. Spheroplasts isolated from apg9ts and ypt7Δ cells were pulse-labeled for 10 min and chased for 30 min at 38°C. The labeled spheroplasts were then osmotically lysed and separated into low-speed supernatant (S) and pellet (P) fractions after a 5,000 g centrifugation step. The pellet fractions were subjected to protease treatment in the absence or presence of 0.2% Triton X-100 as described in Materials and Methods. The resulting samples were immunoprecipitated with antiserum to API and PGK. The immunoprecipitated bands were quantified by a Molecular Dynamics STORM PhosphorImager. The percent protease-protected prAPI was determined by dividing the value for prAPI in the protease-treated sample by the value for total prAPI before protease treatment. The percent PGK was determined by dividing the supernatant (S) or pellet (P) value over the sum of the S and P values.